
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 2,514,233 – 2,514,353 |
| Length | 120 |
| Max. P | 0.882091 |

| Location | 2,514,233 – 2,514,353 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.68 |
| Mean single sequence MFE | -23.10 |
| Consensus MFE | -16.58 |
| Energy contribution | -17.38 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.59 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.92 |
| SVM RNA-class probability | 0.882091 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 2514233 120 + 22224390 UUUUUUUUUCUUUUGCACAUCUUUCCAGAUGCCGUUAAGCUGUUGCUUAAUUUAUAAUUCAUUCCUGUUUGAUUUAACACCUCCGCACUUUAGGUCUCUUGAUAAAUAUGUAUGUGCAGU ............(((((((((..((.(((.((((((((((....)))))))...(((.(((........))).)))................))).))).)).......).)))))))). ( -23.40) >DroSec_CAF1 12532 113 + 1 UUUUUUU---UUUAGCACAUCUCUCCAGAUUCCGUUAAGCUGUUGCUUAAUUUAUAAUUCAUUCCUGUUUUAUUUAACACCUCCGCACUUUAGGUCUCUUGAUAAAUA----UGUGCAGU .......---....((((((...((.(((..(((((((((....)))))))..............((((......)))).............))..))).)).....)----)))))... ( -20.70) >DroSim_CAF1 12490 113 + 1 UUUUUUU---UUUAGCACAUCUUUCCAGAUGCCGUUAAGCUGUUGCUUAAUUUAUAAUUCAUUCCUGUUUGAUUUAACACCUCCGCACUUUAGGUCUCUUGAUAAAUA----UGUGCAGU .......---....((((((...((.(((.((((((((((....)))))))...(((.(((........))).)))................))).))).)).....)----)))))... ( -23.90) >DroEre_CAF1 13173 108 + 1 -----UU---UUUAGCUCAUAUUUGCAGAUGCCGUUAAGCUGUUGCUUAAUUUAUAAUUCAUUCUUGUUUGAUUUAACACCUCCGCACUUUAGGUCUCUUGAUAAGUA----UGUGCAGU -----..---....((.((((((((((((.((((((((((....)))))))...(((.(((........))).)))................))).)).)).))))))----)).))... ( -24.80) >DroYak_CAF1 14398 108 + 1 -----UU---GCUAGCACAUCGUUCCAGAUGCCGUUAAGCUGUUGCUUAAUUUAUAAUUCAUUCUUGUUUAAUUUAACACCUCCGCACUUCAGGUCUCCUGAUAAAUA----UGUGCAGU -----..---(((.((((((......((.(((.(((((((....)))))))..............((((......)))).....)))))(((((...))))).....)----)))))))) ( -22.70) >consensus UUUUUUU___UUUAGCACAUCUUUCCAGAUGCCGUUAAGCUGUUGCUUAAUUUAUAAUUCAUUCCUGUUUGAUUUAACACCUCCGCACUUUAGGUCUCUUGAUAAAUA____UGUGCAGU ..............(((((....((.(((.((((((((((....)))))))..............((((......)))).............))).))).))..........)))))... (-16.58 = -17.38 + 0.80)

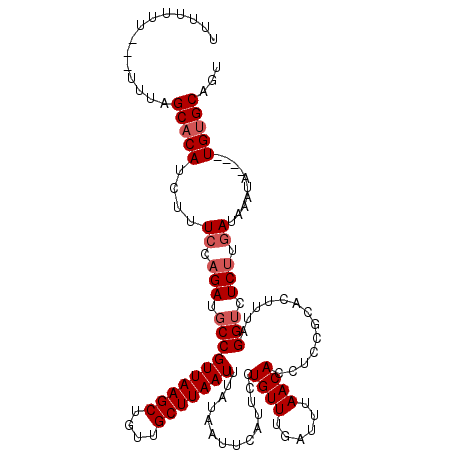
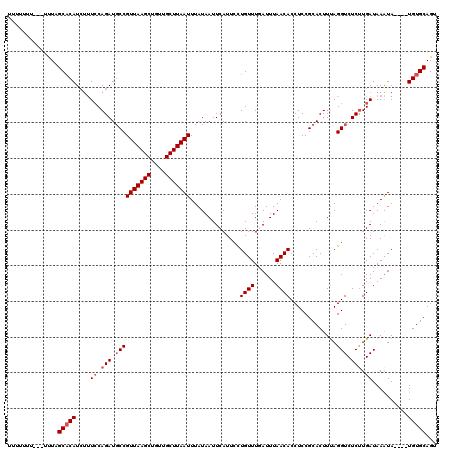
| Location | 2,514,233 – 2,514,353 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.68 |
| Mean single sequence MFE | -21.03 |
| Consensus MFE | -15.97 |
| Energy contribution | -16.17 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.686607 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 2514233 120 - 22224390 ACUGCACAUACAUAUUUAUCAAGAGACCUAAAGUGCGGAGGUGUUAAAUCAAACAGGAAUGAAUUAUAAAUUAAGCAACAGCUUAACGGCAUCUGGAAAGAUGUGCAAAAGAAAAAAAAA ..(((((((.................((........))(((((((...(((........)))........((((((....)))))).)))))))......)))))))............. ( -21.30) >DroSec_CAF1 12532 113 - 1 ACUGCACA----UAUUUAUCAAGAGACCUAAAGUGCGGAGGUGUUAAAUAAAACAGGAAUGAAUUAUAAAUUAAGCAACAGCUUAACGGAAUCUGGAGAGAUGUGCUAAA---AAAAAAA ...(((((----(.....((.(((..((.............((((......))))...............((((((....)))))).))..))).))...))))))....---....... ( -20.30) >DroSim_CAF1 12490 113 - 1 ACUGCACA----UAUUUAUCAAGAGACCUAAAGUGCGGAGGUGUUAAAUCAAACAGGAAUGAAUUAUAAAUUAAGCAACAGCUUAACGGCAUCUGGAAAGAUGUGCUAAA---AAAAAAA .((((((.----......((....))......)))))).(((............................((((((....))))))..((((((....)))))))))...---....... ( -20.92) >DroEre_CAF1 13173 108 - 1 ACUGCACA----UACUUAUCAAGAGACCUAAAGUGCGGAGGUGUUAAAUCAAACAAGAAUGAAUUAUAAAUUAAGCAACAGCUUAACGGCAUCUGCAAAUAUGAGCUAAA---AA----- .((((((.----......((....))......))))))(((((((...(((........)))........((((((....)))))).)))))))((........))....---..----- ( -18.82) >DroYak_CAF1 14398 108 - 1 ACUGCACA----UAUUUAUCAGGAGACCUGAAGUGCGGAGGUGUUAAAUUAAACAAGAAUGAAUUAUAAAUUAAGCAACAGCUUAACGGCAUCUGGAACGAUGUGCUAGC---AA----- .(((((((----(.....(((((...))))).((((.(...((((......))))...............((((((....))))))).))))........)))))).)).---..----- ( -23.80) >consensus ACUGCACA____UAUUUAUCAAGAGACCUAAAGUGCGGAGGUGUUAAAUCAAACAGGAAUGAAUUAUAAAUUAAGCAACAGCUUAACGGCAUCUGGAAAGAUGUGCUAAA___AAAAAAA .((((((.........................))))))(((((((........((....)).........((((((....)))))).))))))).......................... (-15.97 = -16.17 + 0.20)

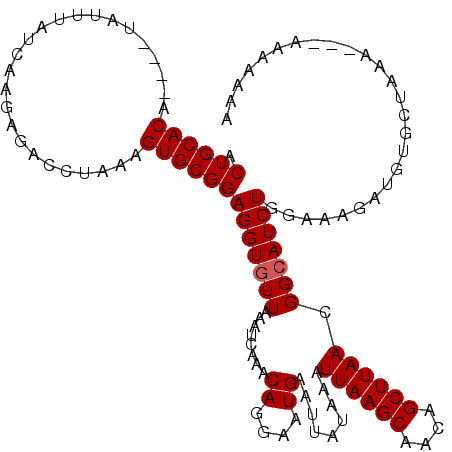
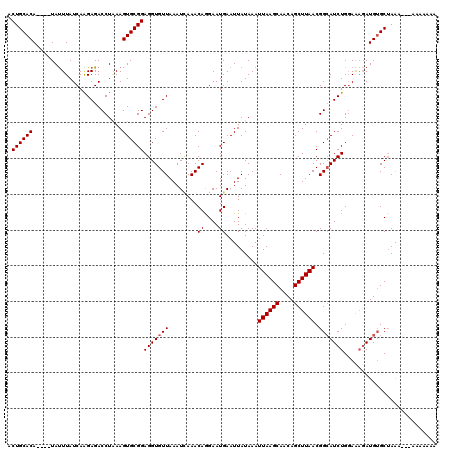
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:56:38 2006