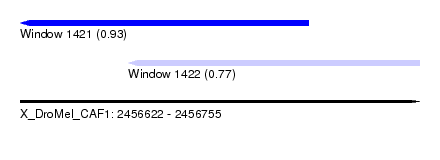
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 2,456,622 – 2,456,755 |
| Length | 133 |
| Max. P | 0.932194 |

| Location | 2,456,622 – 2,456,718 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 55.07 |
| Mean single sequence MFE | -25.71 |
| Consensus MFE | -13.30 |
| Energy contribution | -13.43 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.52 |
| SVM decision value | 1.22 |
| SVM RNA-class probability | 0.932194 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
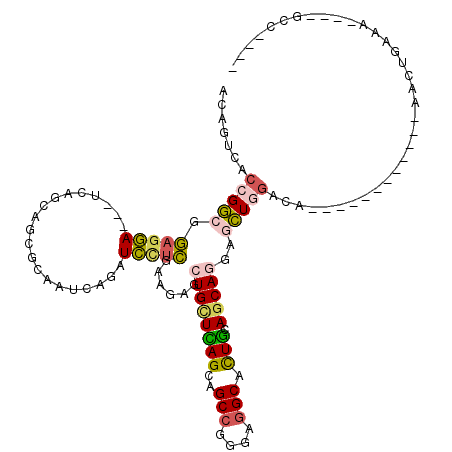
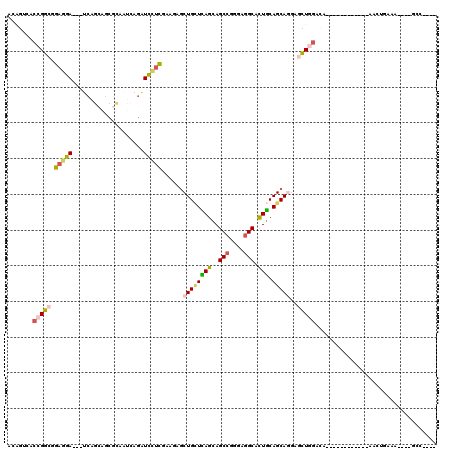
>X_DroMel_CAF1 2456622 96 - 22224390 CUCGAAGAGCUGCUCAGCAGCCGAGAGGCACUGCAGCAGGAGCUGGACA------------AGCUUAGAGCCAAGAACAAGUCAAAAAGCAAGGCAGUUAAGUCCGAG (((((..(((((((..((.(((....))).((...((..(((((.....------------)))))...))..)).............))..)))))))...).)))) ( -31.10) >DroSim_CAF1 1520 96 - 1 CUCGAAGAGCUGCUCAGCAGCCGGGAGGCACUGCAGCAGGAGCUGGACA------------NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN .((..((..((((((((..(((....))).))).)))))...)).))..------------............................................... ( -17.00) >DroEre_CAF1 16177 96 - 1 CUCGAAGAACUGCUCAGCAGCCGAGAGGCACUGCAGCAUGAGCUGGACG------------AACUAAAAGCCAAGACCAAGUCAAAAGCCAAGCCAGUUCCGGCUACU ..........(((((((..(((....))).))).))))((.(((.(((.------------...................)))...)))))((((......))))... ( -25.25) >DroAna_CAF1 15443 105 - 1 UUGGACGAGCUGAUUAGCAGCCGGGAGGCAUUGCAGCAGGAACUGGAAGCCCUGCGCUCGAAAAUGACCGCCAAGAAGAAGCCAAAGGA---GCCGGAAGUGAUUUUG .....(((((......((((((....)))..))).(((((..........))))))))))(((((.((..((.....(....)...(..---..)))..)).))))). ( -29.50) >consensus CUCGAAGAGCUGCUCAGCAGCCGAGAGGCACUGCAGCAGGAGCUGGACA____________AACU_A_AGCCAAGAACAAGUCAAAAG____GCCAGUU__G_CU__G .........((((((((..(((....))).))).)))))..................................................................... (-13.30 = -13.43 + 0.12)

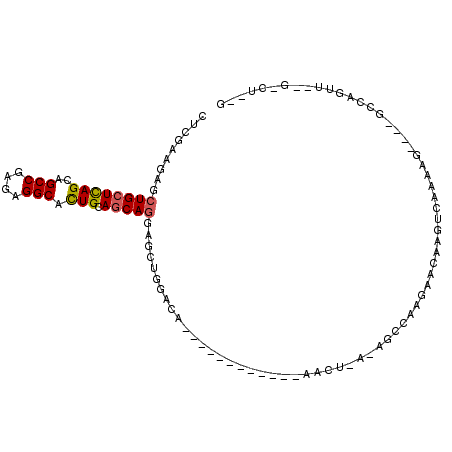
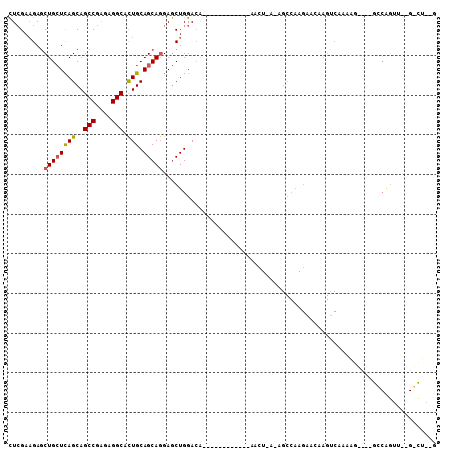
| Location | 2,456,658 – 2,456,755 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 66.24 |
| Mean single sequence MFE | -34.14 |
| Consensus MFE | -19.66 |
| Energy contribution | -19.92 |
| Covariance contribution | 0.26 |
| Combinations/Pair | 1.38 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.771361 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 2456658 97 - 22224390 ACGGUCCCCGGCGGAGGA---UCAGCAGCGCAAUCAGAUCCUCGAAGAGCUGCUCAGCAGCCGAGAGGCACUGCAGCAGGAGCUGGACA------------AGCUUAGA----GCC---- ...((((..(((.(((((---((.............)))))))......((((((((..(((....))).))).)))))..))))))).------------.((.....----)).---- ( -36.92) >DroSec_CAF1 728 97 - 1 UCAGUCACCGGCGGAGGA---UCAGCAGCGCAAUCAGAUCCUCGAAGAGCUGCUCAGCAGCCGGGAGGCACUGCAGCAGGAGCUGGACA------------AGCUUAGA----GCC---- .......(((((.(((((---((.............)))))))......((((((((..(((....))).))).)))))..)))))...------------.((.....----)).---- ( -36.42) >DroSim_CAF1 1556 97 - 1 UCAGACACCGGCGGAGGA---UCAGCAGCGCAAUCAGAUCCUCGAAGAGCUGCUCAGCAGCCGGGAGGCACUGCAGCAGGAGCUGGACA------------NNNNNNNN----NNN---- .......(((((.(((((---((.............)))))))......((((((((..(((....))).))).)))))..)))))...------------........----...---- ( -35.92) >DroEre_CAF1 16213 100 - 1 ACAGUCACCGGCGGAGGAGCUGAAGCAUCACAAUGAAAUCCUCGAAGAACUGCUCAGCAGCCGAGAGGCACUGCAGCAUGAGCUGGACG------------AACUAAAA----GCC---- .......(((((.(((((((....)).((.....))..))))).......(((((((..(((....))).))).))))...)))))...------------........----...---- ( -31.40) >DroAna_CAF1 15476 109 - 1 GCAGGCACAGGAGCAGGA---ACAGCAGCGUCAACAGUUCUUGGACGAGCUGAUUAGCAGCCGGGAGGCAUUGCAGCAGGAACUGGAAGCCCUGCGCUCGAAAAUGACC----GCC---- ...(((.(((((((.(..---((......))...).)))))))..(((((......((((((....)))..))).(((((..........)))))))))).........----)))---- ( -37.20) >DroPer_CAF1 8284 92 - 1 ---------GCAGGCGAA---UC----GCACCAAGAGAUUAUCGAGGAGCUGUUGAGCAGCCGCGAAGCCCUUCAGCAACAGGUCGAUC------------GUCUGAAGUCGAGCAAGGA ---------.(((((((.---((----(.(((..((.....))((((.(((.....((....))..))).)))).......))))))))------------))))).............. ( -27.00) >consensus ACAGUCACCGGCGGAGGA___UCAGCAGCGCAAUCAGAUCCUCGAAGAGCUGCUCAGCAGCCGGGAGGCACUGCAGCAGGAGCUGGACA____________AACUGAAA____GCC____ .......(((((.(((((....................)))))......((((((((..(((....))).))).)))))..))))).................................. (-19.66 = -19.92 + 0.26)
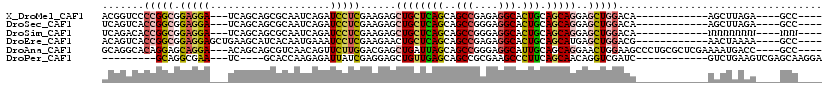
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:56:01 2006