
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 21,937,701 – 21,937,794 |
| Length | 93 |
| Max. P | 0.996795 |

| Location | 21,937,701 – 21,937,794 |
|---|---|
| Length | 93 |
| Sequences | 3 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 74.50 |
| Mean single sequence MFE | -18.23 |
| Consensus MFE | -12.73 |
| Energy contribution | -12.40 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.70 |
| SVM decision value | 2.75 |
| SVM RNA-class probability | 0.996795 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
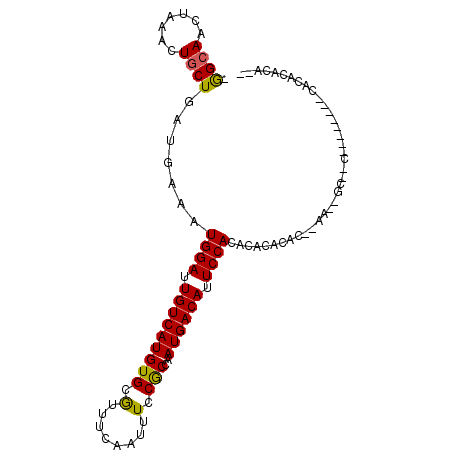
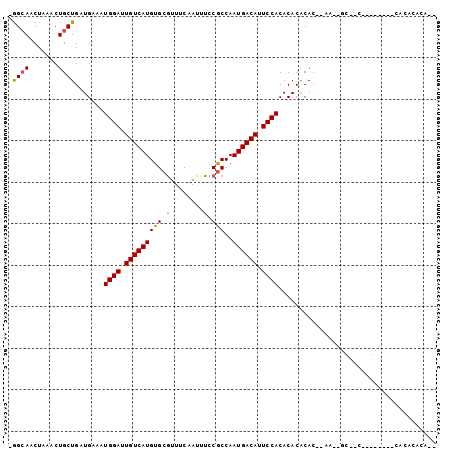
>X_DroMel_CAF1 21937701 93 + 22224390 UGGAAACUAAACUGCUGAUGAAAUGGAUUGUCAUGUGCAUUUCAAUUUCCACCAAUGACAUUCCACACACCCAC--AACUGCUUC--------CACACACAUC (((((.(.....((.((.((...((((.(((((((((.(........).)))..)))))).))))..))..)))--)...).)))--------))........ ( -15.40) >DroVir_CAF1 74773 75 + 1 -AGCAACUAAACUGCUGAUGAAAUGGAUUGUCAUGUGCGUUUCAAUUUCCGCCAAUGACAUUCCACACACACAC--AC------------------------- -((((.......)))).......((((.((((((..(((..........)))..)))))).)))).........--..------------------------- ( -17.50) >DroEre_CAF1 70165 102 + 1 -GGCAACUAAACUGCUGAUGAAAUGGAUUGUCAUGUGCGUUUCAAUUUCCGCCAAUGACACUCCACACACACACUGAAAGGCAACCAGCCAGACACACACACA -(....)....((((((......((((.((((((..(((..........)))..)))))).))))..............(....)))).)))........... ( -21.80) >consensus _GGCAACUAAACUGCUGAUGAAAUGGAUUGUCAUGUGCGUUUCAAUUUCCGCCAAUGACAUUCCACACACACAC__AA__GC__C________CACACACA__ .((((.......)))).......((((.(((((((((.(........).)))..)))))).))))...................................... (-12.73 = -12.40 + -0.33)

| Location | 21,937,701 – 21,937,794 |
|---|---|
| Length | 93 |
| Sequences | 3 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 74.50 |
| Mean single sequence MFE | -25.53 |
| Consensus MFE | -19.34 |
| Energy contribution | -18.90 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.952662 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
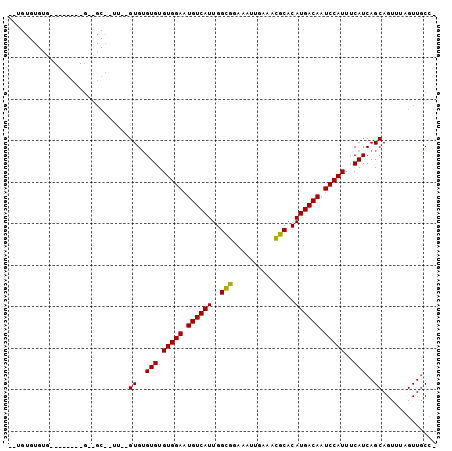
>X_DroMel_CAF1 21937701 93 - 22224390 GAUGUGUGUG--------GAAGCAGUU--GUGGGUGUGUGGAAUGUCAUUGGUGGAAAUUGAAAUGCACAUGACAAUCCAUUUCAUCAGCAGUUUAGUUUCCA ........((--------(((((..((--((.((((.(((((.((((((..(((.(........).))))))))).)))))..)))).))))....))))))) ( -29.00) >DroVir_CAF1 74773 75 - 1 -------------------------GU--GUGUGUGUGUGGAAUGUCAUUGGCGGAAAUUGAAACGCACAUGACAAUCCAUUUCAUCAGCAGUUUAGUUGCU- -------------------------.(--(.(((...(((((.((((((..(((..........)))..)))))).)))))..)))))((((.....)))).- ( -21.60) >DroEre_CAF1 70165 102 - 1 UGUGUGUGUGUCUGGCUGGUUGCCUUUCAGUGUGUGUGUGGAGUGUCAUUGGCGGAAAUUGAAACGCACAUGACAAUCCAUUUCAUCAGCAGUUUAGUUGCC- ........((.((((((((.......)))))(((...(((((.((((((..(((..........)))..)))))).)))))..))))))))...........- ( -26.00) >consensus __UGUGUGUG________G__GC__UU__GUGUGUGUGUGGAAUGUCAUUGGCGGAAAUUGAAACGCACAUGACAAUCCAUUUCAUCAGCAGUUUAGUUGCC_ .............................((..(((.(((((.((((((..(((..........)))..)))))).)))))..)))..))............. (-19.34 = -18.90 + -0.44)
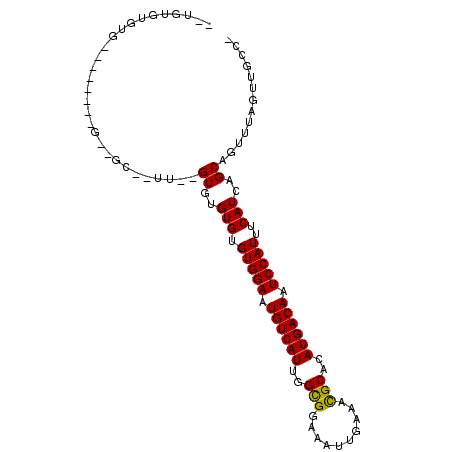


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:09:36 2006