
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 21,307,536 – 21,307,630 |
| Length | 94 |
| Max. P | 0.913263 |

| Location | 21,307,536 – 21,307,630 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 86.17 |
| Mean single sequence MFE | -31.16 |
| Consensus MFE | -18.93 |
| Energy contribution | -19.33 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.22 |
| Mean z-score | -3.21 |
| Structure conservation index | 0.61 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.873602 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 21307536 94 + 22224390 ACAAUCUCCAUUGGCCCCCUGGAAGUGG-------GCAAGCGGCUAGAACGCUUUCGUCCUUGUUCUCCUUUUGCGG-CAAAACAGGACGCCCAAGGAAAUA ......(((.....((....))...(((-------(((((((.......)))))..(((((.(((..((......))-...))))))))))))).))).... ( -28.10) >DroSec_CAF1 49007 93 + 1 ACAAACUCCAUUGGCCCCCUGAAGGUGG-------CCACUCUUCUGG-CCGCCUUCGUCCUUGUUCUCCUUCUGCGG-CAAAACAGGACGCCCAAGGAAAUA ......(((...(((.....((((((((-------(((......)))-))))))))(((((.(((..((......))-...)))))))))))...))).... ( -38.20) >DroSim_CAF1 45422 93 + 1 ACAAACUCCAUUGGCCCCCUGAAGGUGG-------CCACUCUUCUGG-CCGCCUUCGUCCUUGUUCUCCUUCUGCGG-CAAAACAGGACGCCCAAGGAAAUA ......(((...(((.....((((((((-------(((......)))-))))))))(((((.(((..((......))-...)))))))))))...))).... ( -38.20) >DroEre_CAF1 48044 94 + 1 ACAAUCUCCAUUGGCCCCCUGAAAGCGG-------CCACACUUCUAG-CCGCCUUCGUCCUUGUUCUCCCCCUGCUGCCAAAACAGGACGCCCAAGGAAAUA ......(((...(((.....(((.((((-------(..........)-)))).)))(((((.(((................)))))))))))...))).... ( -24.79) >DroYak_CAF1 50603 101 + 1 ACAAUCUUCAUUGGCCCCCUAAAAGUGGCCACUGGCCACACUUCCAG-CCGUCUUCGUCCUUGUGCUCCUUCUGCUGCCAAAACAGGACGCCCAAGGAAAUA ............(((.........((((((...)))))).......)-)).......((((((.((((((..............)))).)).)))))).... ( -26.53) >consensus ACAAUCUCCAUUGGCCCCCUGAAAGUGG_______CCACACUUCUAG_CCGCCUUCGUCCUUGUUCUCCUUCUGCGG_CAAAACAGGACGCCCAAGGAAAUA ......(((...(((.....(((((((......................)))))))(((((.(((..((......))....)))))))))))...))).... (-18.93 = -19.33 + 0.40)

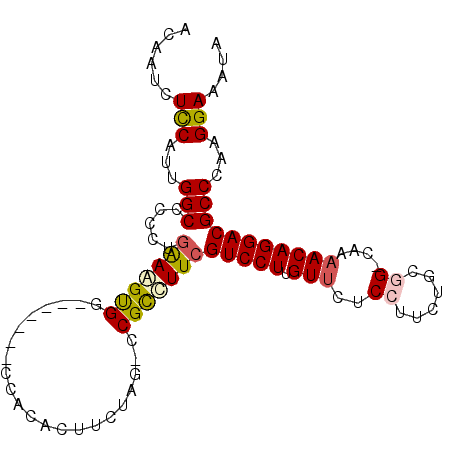
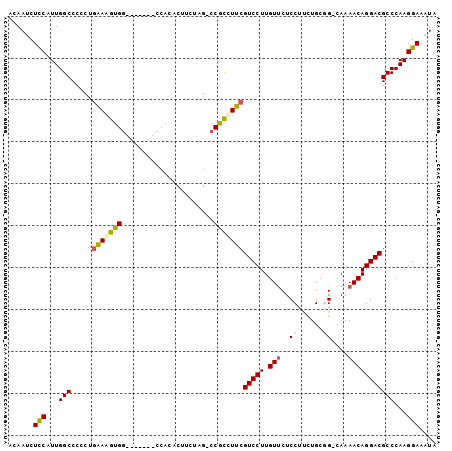
| Location | 21,307,536 – 21,307,630 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 86.17 |
| Mean single sequence MFE | -38.54 |
| Consensus MFE | -24.88 |
| Energy contribution | -26.28 |
| Covariance contribution | 1.40 |
| Combinations/Pair | 1.12 |
| Mean z-score | -3.02 |
| Structure conservation index | 0.65 |
| SVM decision value | 1.09 |
| SVM RNA-class probability | 0.913263 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 21307536 94 - 22224390 UAUUUCCUUGGGCGUCCUGUUUUG-CCGCAAAAGGAGAACAAGGACGAAAGCGUUCUAGCCGCUUGC-------CCACUUCCAGGGGGCCAAUGGAGAUUGU .((((((((((.(((((((((((.-((......))))))).)))))..(((((.......))))).(-------((.......)))).)))).))))))... ( -34.00) >DroSec_CAF1 49007 93 - 1 UAUUUCCUUGGGCGUCCUGUUUUG-CCGCAGAAGGAGAACAAGGACGAAGGCGG-CCAGAAGAGUGG-------CCACCUUCAGGGGGCCAAUGGAGUUUGU ...((((((((.(((((((((((.-((......))))))).)))))(((((.((-(((......)))-------)).)))))....).)))).))))..... ( -42.40) >DroSim_CAF1 45422 93 - 1 UAUUUCCUUGGGCGUCCUGUUUUG-CCGCAGAAGGAGAACAAGGACGAAGGCGG-CCAGAAGAGUGG-------CCACCUUCAGGGGGCCAAUGGAGUUUGU ...((((((((.(((((((((((.-((......))))))).)))))(((((.((-(((......)))-------)).)))))....).)))).))))..... ( -42.40) >DroEre_CAF1 48044 94 - 1 UAUUUCCUUGGGCGUCCUGUUUUGGCAGCAGGGGGAGAACAAGGACGAAGGCGG-CUAGAAGUGUGG-------CCGCUUUCAGGGGGCCAAUGGAGAUUGU .((((((((((.(((((((((((..(......)..))))).)))))((((((((-(((......)))-------))))))))....).)))).))))))... ( -40.90) >DroYak_CAF1 50603 101 - 1 UAUUUCCUUGGGCGUCCUGUUUUGGCAGCAGAAGGAGCACAAGGACGAAGACGG-CUGGAAGUGUGGCCAGUGGCCACUUUUAGGGGGCCAAUGAAGAUUGU ....((((((.((.((((..((((....)))))))))).)))))).......((-((.((((.((((((...))))))))))....))))............ ( -33.00) >consensus UAUUUCCUUGGGCGUCCUGUUUUG_CCGCAGAAGGAGAACAAGGACGAAGGCGG_CUAGAAGAGUGG_______CCACUUUCAGGGGGCCAAUGGAGAUUGU .((((((((((.(((((((((((..((......))))))).))))).........((.((((.(((((.....))))))))).)).).)))).))))))... (-24.88 = -26.28 + 1.40)

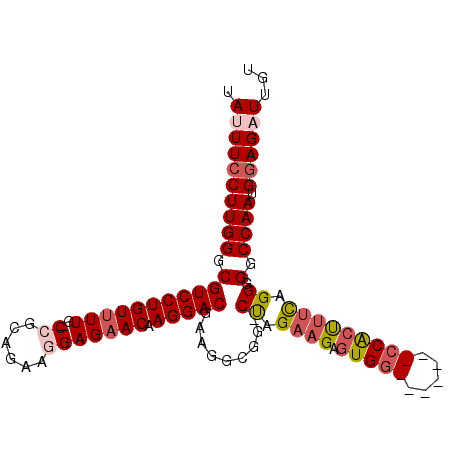
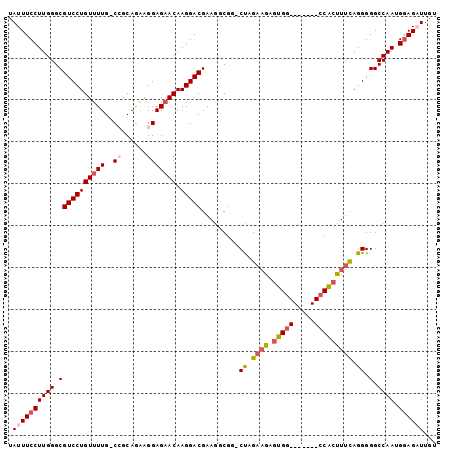
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:07:56 2006