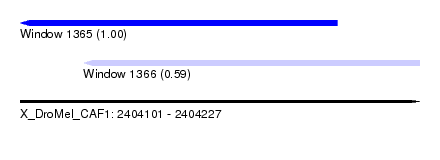

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 2,404,101 – 2,404,227 |
| Length | 126 |
| Max. P | 0.999072 |

| Location | 2,404,101 – 2,404,201 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 71.52 |
| Mean single sequence MFE | -27.12 |
| Consensus MFE | -14.68 |
| Energy contribution | -15.99 |
| Covariance contribution | 1.31 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.78 |
| Structure conservation index | 0.54 |
| SVM decision value | 3.36 |
| SVM RNA-class probability | 0.999072 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
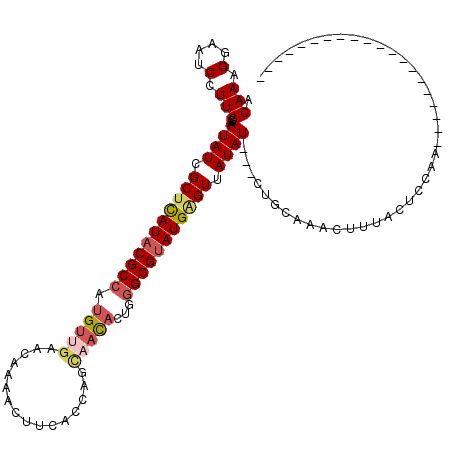
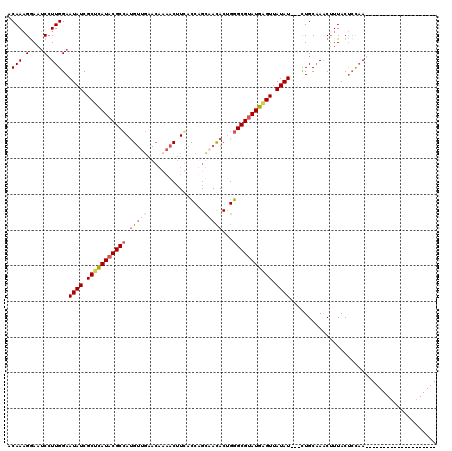
>X_DroMel_CAF1 2404101 100 - 22224390 ACAAUGGAAACCUUGGAAUAUCGCUCAUACGCCAUGUUGAACACAACUUGACCUGUUACACUGGGCGUAUGAGUUAUAUUUUUUGCAAACUUUACCCCAA-------------------- .(((.(....).)))((((((.(((((((((((......((((..........))))......))))))))))).))))))...................-------------------- ( -28.00) >DroSec_CAF1 46941 97 - 1 ACAAAGGAAUCGUUGGAAUAUCGCUCAUACGCCAUGUUGAACACAACUUGACCUGUAACACUGGGCGUAUGGGUAAUAU---CUGCCAACUUUACUACAA-------------------- ...........(((((.((((..((((((((((.(((((....(.....).....)))))...))))))))))..))))---...)))))..........-------------------- ( -29.70) >DroEre_CAF1 49942 91 - 1 ACAAAGAAAUCCUUGGAAUAUCGCAUAUACGCGAUG-------AUACAUCACCAGCCACACUGGGCGUAUGAGUUAUAU---UUGCAAACAUUACUCUUAG------------------- ...(((((((..(((((((((.((.(((((((((((-------...)))).((((.....))))))))))).)).))))---)).)))..)))..))))..------------------- ( -22.80) >DroYak_CAF1 49807 117 - 1 ACAAAGGAAUCCUUGGAAUAUCGCUUAUACGCCACGUUGAACAAAACUUCAUCAGCAAUACGGGGCGGAUGAGUUAUAU---CUGCAAGCUUUACUCCAAGAUAGCUGCACAAUAAGCUG ...........((((((.((..((((.....((.(((((((......))))........)))))(((((((.....)))---))))))))..)).)))))).(((((........))))) ( -28.00) >consensus ACAAAGGAAUCCUUGGAAUAUCGCUCAUACGCCAUGUUGAACAAAACUUCACCAGCAACACUGGGCGUAUGAGUUAUAU___CUGCAAACUUUACUCCAA____________________ .(((.(....).)))..((((.(((((((((((.(((((................)))))...))))))))))).))))......................................... (-14.68 = -15.99 + 1.31)

| Location | 2,404,121 – 2,404,227 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 77.73 |
| Mean single sequence MFE | -27.00 |
| Consensus MFE | -17.55 |
| Energy contribution | -18.86 |
| Covariance contribution | 1.31 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.594125 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
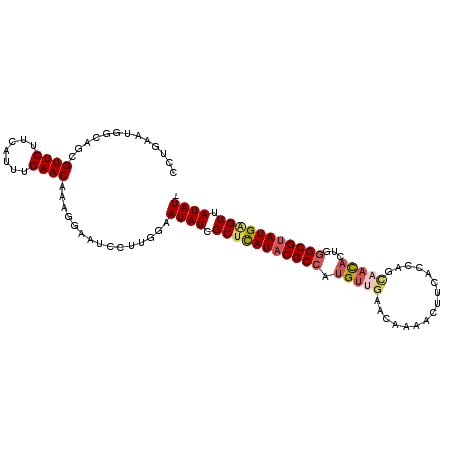
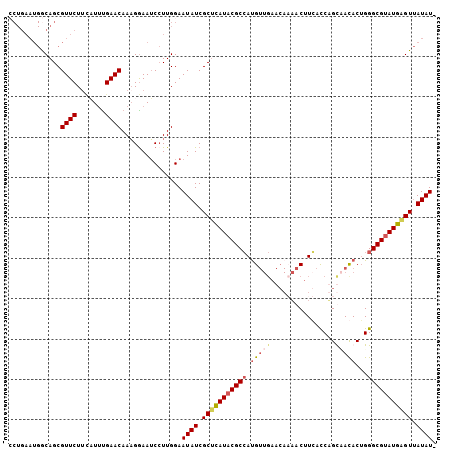
>X_DroMel_CAF1 2404121 106 - 22224390 CCUGAAUGGCAGCGUUCUUCUUUUGAACAAUGGAAACCUUGGAAUAUCGCUCAUACGCCAUGUUGAACACAACUUGACCUGUUACACUGGGCGUAUGAGUUAUAUU ...(((((....)))))..........(((.(....).))).(((((.(((((((((((......((((..........))))......))))))))))).))))) ( -29.70) >DroSec_CAF1 46959 97 - 1 --------GCAGCGUUCUUCUUUCGAACAAAGGAAUCGUUGGAAUAUCGCUCAUACGCCAUGUUGAACACAACUUGACCUGUAACACUGGGCGUAUGGGUAAUAU- --------.(((((...((((((......)))))).)))))..((((..((((((((((.(((((....(.....).....)))))...))))))))))..))))- ( -29.90) >DroEre_CAF1 49961 98 - 1 ACUGAAUGACAGCGUUCUCUAUGUGAACAAAGAAAUCCUUGGAAUAUCGCAUAUACGCGAUG-------AUACAUCACCAGCCACACUGGGCGUAUGAGUUAUAU- .....(((((.(((...(((.((....)).)))..((....))....))).(((((((((((-------...)))).((((.....))))))))))).)))))..- ( -26.40) >DroYak_CAF1 49845 105 - 1 CCUGAAUGGCAGCGUUCUUGAUUUGAACAAAGGAAUCCUUGGAAUAUCGCUUAUACGCCACGUUGAACAAAACUUCAUCAGCAAUACGGGGCGGAUGAGUUAUAU- ((...........((((.......)))).((((...)))))).((((.((((((.((((.(((((((......))))........))).)))).)))))).))))- ( -22.00) >consensus CCUGAAUGGCAGCGUUCUUCAUUUGAACAAAGGAAUCCUUGGAAUAUCGCUCAUACGCCAUGUUGAACAAAACUUCACCAGCAACACUGGGCGUAUGAGUUAUAU_ .............((((.......))))...............((((.(((((((((((.(((((................)))))...))))))))))).)))). (-17.55 = -18.86 + 1.31)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:55:11 2006