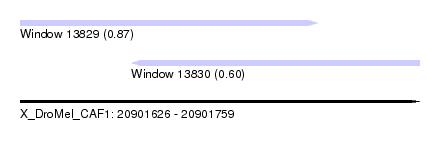

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 20,901,626 – 20,901,759 |
| Length | 133 |
| Max. P | 0.872494 |

| Location | 20,901,626 – 20,901,725 |
|---|---|
| Length | 99 |
| Sequences | 3 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 83.50 |
| Mean single sequence MFE | -30.40 |
| Consensus MFE | -19.75 |
| Energy contribution | -20.20 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.872494 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
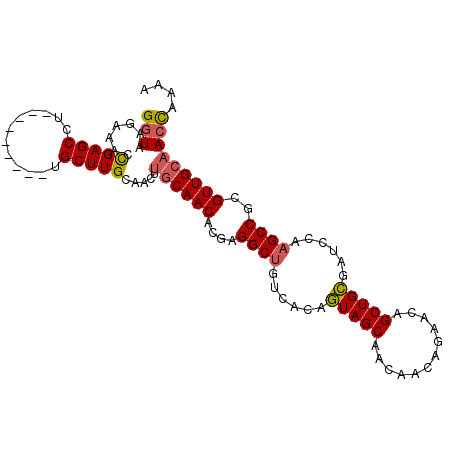
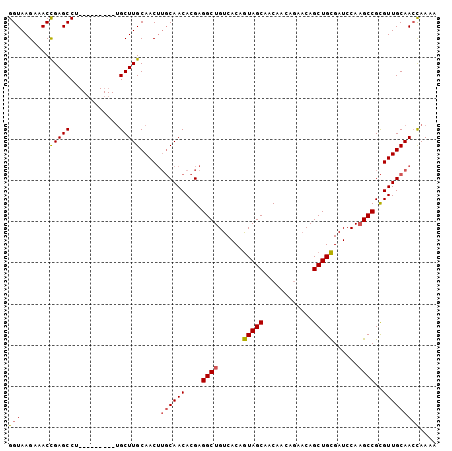
>X_DroMel_CAF1 20901626 99 + 22224390 GGUAAGAAACCGAGCCUUGCUCGCCUUGCUUGCAACGGCCAACACGAGGCUGUCACAAUAGCAACAA-----CAGCUGUGAUCCAAGCCGUGUUGCAAGUAAAA (((.....)))((((...))))...((((((((((((.....)....((((......(((((.....-----..)))))......))))..))))))))))).. ( -33.00) >DroSec_CAF1 20896 95 + 1 GGUAAGAAACUGAGCCU---------CGCUUGCAACUUGCAACACGAGGCUGUCACAGUAGCAACAACAGAACAGCUGCGAUCCACGCCGCGUUGCAACCAAAA (((.....(((((((((---------((.((((.....))))..)))))))....)))).(((((.........((.(((.....))).))))))).))).... ( -31.20) >DroSim_CAF1 22784 95 + 1 GGUAAGAAACCGAGCCU---------UGCUUGCAACUUGCAACACGAGGCUGUCACAGUAGCAACAACAGAACAGCUGCGAUCCAAGCCGCGUUGCAACCAAAA (((.......(((((..---------.))))).....((((((....((((......(((((............)))))......))))..))))))))).... ( -27.00) >consensus GGUAAGAAACCGAGCCU_________UGCUUGCAACUUGCAACACGAGGCUGUCACAGUAGCAACAACAGAACAGCUGCGAUCCAAGCCGCGUUGCAACCAAAA (((.......(((((............))))).....((((((....((((......(((((............)))))......))))..))))))))).... (-19.75 = -20.20 + 0.45)

| Location | 20,901,663 – 20,901,759 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 68.76 |
| Mean single sequence MFE | -26.50 |
| Consensus MFE | -9.67 |
| Energy contribution | -12.79 |
| Covariance contribution | 3.12 |
| Combinations/Pair | 1.28 |
| Mean z-score | -2.44 |
| Structure conservation index | 0.36 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.595314 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
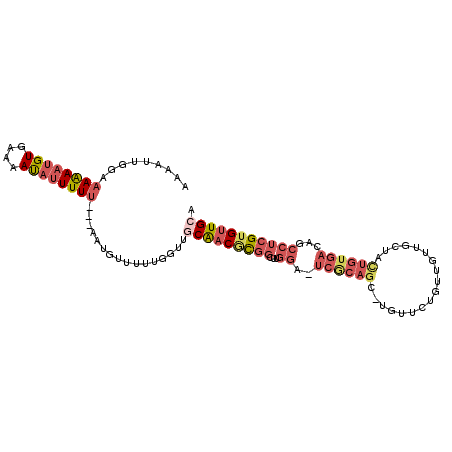
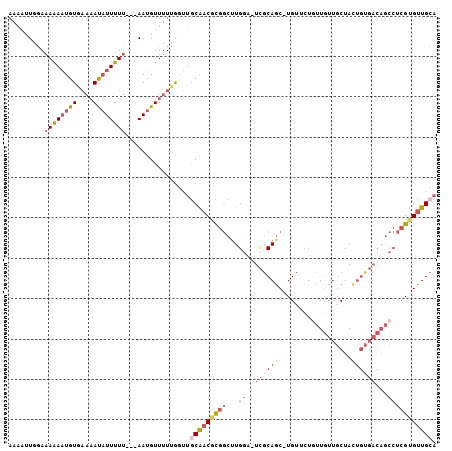
>X_DroMel_CAF1 20901663 96 - 22224390 AACUCUAAAAAAAAUGUGAAAACAUUUUU---AAUGUUUUUACUUGCAACACGGCUUGGA-UCACAGC-UG-----UUGUUGCUAUUGUGACAGCCUCGUGUUGGC ...............((((((((((....---.))))))))))...((((((((((.(..-...))))-.(-----((((..(....)..)))))..))))))).. ( -29.00) >DroSec_CAF1 20924 101 - 1 AAAAUUGGAAAAAAUGUGAAAAUAUUUUU---AAUGUUUUUGGUUGCAACGCGGCGUGGA-UCGCAGC-UGUUCUGUUGUUGCUACUGUGACAGCCUCGUGUUGCA ..(((..((((((((((....))))))..---....))))..)))(((((((((((....-.)))...-......(((((..(....)..)))))..)))))))). ( -31.30) >DroSim_CAF1 22812 101 - 1 AAAAAUGGAAAAAAUGUAAAAAUAUUUUU---AAUGUUUUUGGUUGCAACGCGGCUUGGA-UCGCAGC-UGUUCUGUUGUUGCUACUGUGACAGCCUCGUGUUGCA (((((((..((((((((....))))))))---..)))))))...((((((((((((....-....)))-......(((((..(....)..)))))..))))))))) ( -28.60) >DroAna_CAF1 11826 92 - 1 -UGUUUUGAUAGACUGUUUAUAUUUUUUUCGCAAUGUUAUUGGAUCCGUCCUCAGUUGCACUUGCAACAUGUCCUAUUAUUUCUA-------------GUGUUGCA -.((((....))))................((((..(((..(((....)))...(((((....)))))...............))-------------)..)))). ( -17.10) >consensus AAAAUUGGAAAAAAUGUGAAAAUAUUUUU___AAUGUUUUUGGUUGCAACGCGGCUUGGA_UCGCAGC_UGUUCUGUUGUUGCUACUGUGACAGCCUCGUGUUGCA .........((((((((....))))))))................(((((((((...((..((((((..................))))))...))))))))))). ( -9.67 = -12.79 + 3.12)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:03:31 2006