
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,955,112 – 19,955,226 |
| Length | 114 |
| Max. P | 0.663113 |

| Location | 19,955,112 – 19,955,226 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 64.84 |
| Mean single sequence MFE | -35.98 |
| Consensus MFE | -11.41 |
| Energy contribution | -12.69 |
| Covariance contribution | 1.28 |
| Combinations/Pair | 1.55 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.32 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.663113 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19955112 114 + 22224390 CAUUGACUUUUCCCAUGGGUGUGG--GUAAAAAAGGCUGCGAAAAUGGGAAAAUUCGGGG--UGGGGUUGCCUCUUGGCAGAUGUUCCC-AGUGUAUUACAUUU-GAGCGCCGUAAUUAU ....((.(((((((((...((..(--.(......).)..))...))))))))).))((.(--((((((((((....)))))....))))-((((.....)))).-..)).))........ ( -34.70) >DroSec_CAF1 67602 114 + 1 UAUUGACUUUUCCCAUGGGUGUGG--GUAA--AAGGCUGCGAAAAUGGGAAAAUCGGGGGGUAGGGGUUGCCUCUUGGCAGCUGUUCCC-UGUGUAUUACAUUU-GAGCGCCGUAAUUAU ......((((((((((....))))--).))--)))((.(((.(((((.......((.((((.((.(((((((....))))))).)))))-).)).....)))))-...))).))...... ( -41.30) >DroSim_CAF1 56984 114 + 1 UAUUGACUUUUCCCAUGGGUGUGG--GUAA--AAGGCUGCGAAAAUGGGAAAAUCGGUGGGUAGGGGUUGCCUCUUGGCAGCUGUUCCC-UGUGUAUUACAUUU-GAGCGCCGUAAUUAU ((((((.(((((((((...((..(--.(..--..).)..))...)))))))))))))))(((...(((((((....)))))))((((..-.(((.....)))..-)))))))........ ( -38.80) >DroYak_CAF1 56727 113 + 1 UAUUGACUUUUCCGAUGGGUGCGGCGGGAA--AAGGUUGCGGAAAUGGGAAAAUCGGGG---AGGGGCUGCUUCUUGGCAGCUGUUCCC-UGUGUAUUACAUUU-GAGCGGCGUAAUUAU ......(((((((..((....))...))))--)))(((((..(((((.......(((((---((.(((((((....))))))).)))))-)).......)))))-..)))))........ ( -44.54) >DroMoj_CAF1 116164 98 + 1 ----------------GCGUCCGG--GCCG--GAGUCUGUGAGAGCCGCUGUUUCGUGGCGCAGGCUUUGCUGCCUAGCCAAAGUUUGC-CCACUAUCAAAAUC-AAGCAACUAAUAUGU ----------------((.(((((--((..--..))))).))(((((((......)))))((((((((((((....)).))))))))))-......))......-..))........... ( -29.00) >DroAna_CAF1 108633 97 + 1 UAUUGACUUUUCCGAAAAG---------GA--AAGUGUCCGGAAAAGCCAG-AUGAGGG--AGAGGGUUACUCUUUGGCAGCUGUUCCCUUGUAUACUACAUUCCGAG---------UGU ....(((((((((.....)---------))--))).)))(((((.......-(..((((--((..((((.((....)).)))).))))))..)........)))))..---------... ( -27.56) >consensus UAUUGACUUUUCCCAUGGGUGUGG__GUAA__AAGGCUGCGAAAAUGGGAAAAUCGGGGG__AGGGGUUGCCUCUUGGCAGCUGUUCCC_UGUGUAUUACAUUU_GAGCGCCGUAAUUAU ..................................(((.((..(((((.........(((...((.(((((((....))))))).)))))..........)))))...)))))........ (-11.41 = -12.69 + 1.28)

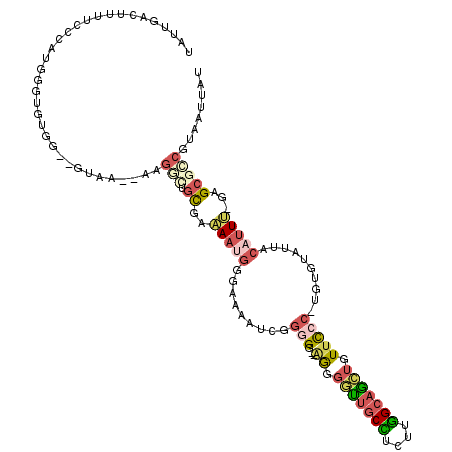
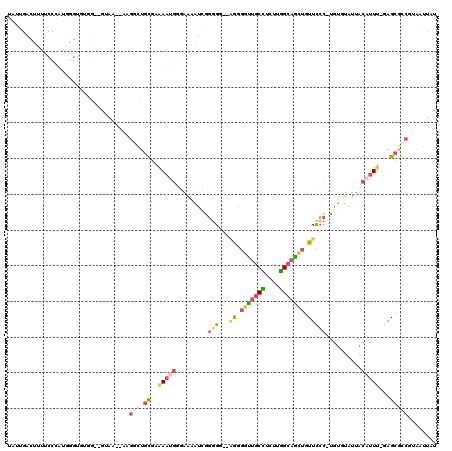
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:53:38 2006