
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,709,567 – 19,709,657 |
| Length | 90 |
| Max. P | 0.575356 |

| Location | 19,709,567 – 19,709,657 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 90 |
| Reading direction | forward |
| Mean pairwise identity | 89.70 |
| Mean single sequence MFE | -28.18 |
| Consensus MFE | -24.49 |
| Energy contribution | -23.80 |
| Covariance contribution | -0.69 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.547445 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19709567 90 + 22224390 UUUGGCUCCUCGGACGGUUCUCCUUGCUUCUCCUUGUGCCUUUUUUUAUUGCGCCACACGGACGGAAUGGACCUCACCGGACAGCCGCCG ...(((..((.(..((((..(((...((..(((.((((((..........).).)))).))).))...)))....))))..)))..))). ( -23.50) >DroSec_CAF1 5641 88 + 1 UUUGGCUCCUCGGACGGUUCUCCUUGCUUCUCCUUGUGCCUUU--UUAUUCCGCCACCCGGACGGAAUGGACCGCACCGGACAGCCGCCG ...(((.....(((......)))..(((..(((..((((...(--((((((((((....)).)))))))))..)))).))).))).))). ( -30.10) >DroEre_CAF1 3067 88 + 1 UUUGGCUUCUCGGACGGUUUUCCUUGCUUCUCCUUGCGCUUUU--UUAUUCCGCCACCCGGACGGAGUGGACCACACCGGACAGCCACCG ..(((((((.(((..((((......((........))......--.(((((((((....)).)))))))))))...))))).)))))... ( -27.60) >DroYak_CAF1 2819 88 + 1 UUUGGCUUCUCGGAUGGUUCUCCUUGGUACUCGUUGCUCCUUU--UUAUUCCGCCACCCGGACGGAAUGGACCGUACCGGACAGCCGCCG ...((((((.((((((((((.....((..(.....)..))...--..((((((((....)).))))))))))))).))))).)))).... ( -31.50) >consensus UUUGGCUCCUCGGACGGUUCUCCUUGCUUCUCCUUGCGCCUUU__UUAUUCCGCCACCCGGACGGAAUGGACCGCACCGGACAGCCGCCG ..(((((..((((..(((((.....((..........))........((((((((....)).)))))))))))...))))..)))))... (-24.49 = -23.80 + -0.69)

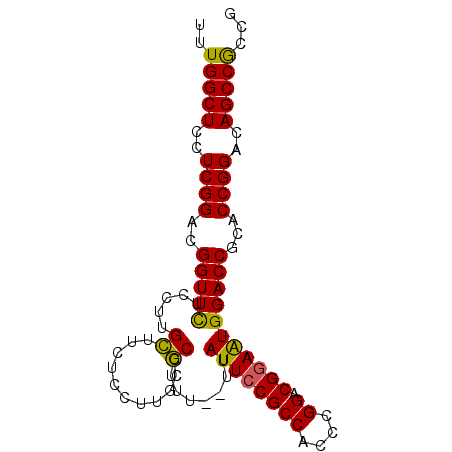
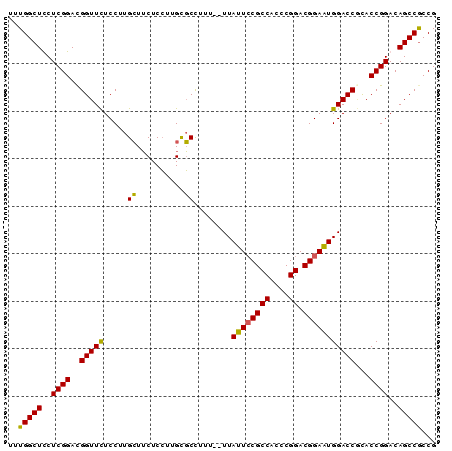
| Location | 19,709,567 – 19,709,657 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 90 |
| Reading direction | reverse |
| Mean pairwise identity | 89.70 |
| Mean single sequence MFE | -29.95 |
| Consensus MFE | -25.30 |
| Energy contribution | -25.30 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.42 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.575356 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19709567 90 - 22224390 CGGCGGCUGUCCGGUGAGGUCCAUUCCGUCCGUGUGGCGCAAUAAAAAAAGGCACAAGGAGAAGCAAGGAGAACCGUCCGAGGAGCCAAA ....((((.(.(((...(((...((((.(((.((((.(............).)))).))).......)))).)))..))).).))))... ( -26.70) >DroSec_CAF1 5641 88 - 1 CGGCGGCUGUCCGGUGCGGUCCAUUCCGUCCGGGUGGCGGAAUAA--AAAGGCACAAGGAGAAGCAAGGAGAACCGUCCGAGGAGCCAAA .(((.((..(((((.((((......)))))))))..))(((....--....((..........))..((....)).))).....)))... ( -31.10) >DroEre_CAF1 3067 88 - 1 CGGUGGCUGUCCGGUGUGGUCCACUCCGUCCGGGUGGCGGAAUAA--AAAAGCGCAAGGAGAAGCAAGGAAAACCGUCCGAGAAGCCAAA ...(((((.(((((.((..(((..((((((.....))))))....--......((........))..)))..))...))).))))))).. ( -27.80) >DroYak_CAF1 2819 88 - 1 CGGCGGCUGUCCGGUACGGUCCAUUCCGUCCGGGUGGCGGAAUAA--AAAGGAGCAACGAGUACCAAGGAGAACCAUCCGAGAAGCCAAA .(((.((..(((((.((((......)))))))))..))(((....--...((.((.....)).))..((....)).))).....)))... ( -34.20) >consensus CGGCGGCUGUCCGGUGCGGUCCAUUCCGUCCGGGUGGCGGAAUAA__AAAGGCACAAGGAGAAGCAAGGAGAACCGUCCGAGAAGCCAAA .(((.((..(((((.((((......)))))))))..)).............................(((......))).....)))... (-25.30 = -25.30 + 0.00)

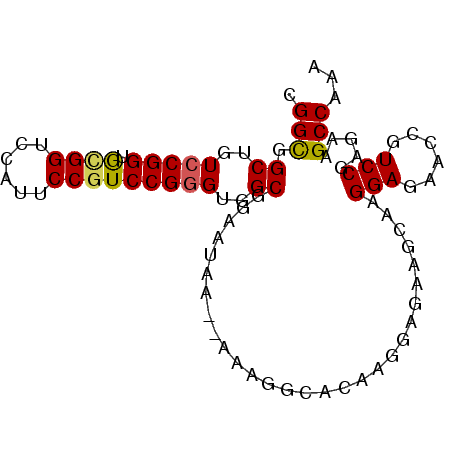
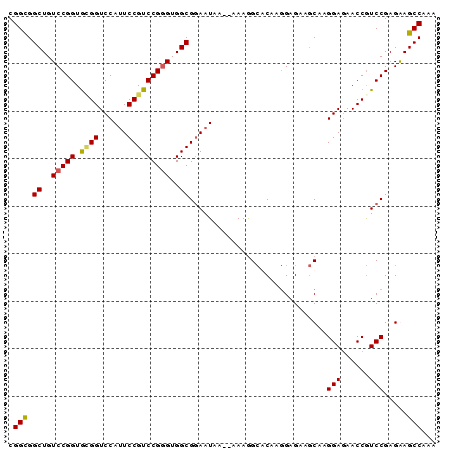
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:50:49 2006