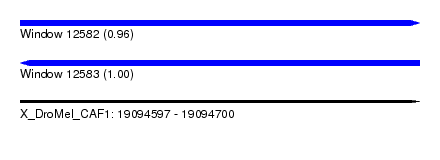
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,094,597 – 19,094,700 |
| Length | 103 |
| Max. P | 0.999941 |

| Location | 19,094,597 – 19,094,700 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 70.36 |
| Mean single sequence MFE | -26.00 |
| Consensus MFE | -13.64 |
| Energy contribution | -15.67 |
| Covariance contribution | 2.03 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.52 |
| SVM decision value | 1.45 |
| SVM RNA-class probability | 0.955013 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
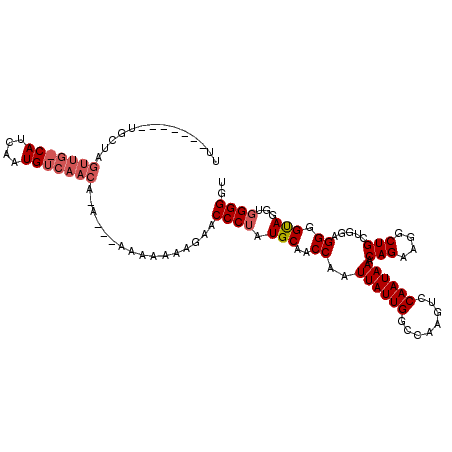
>X_DroMel_CAF1 19094597 103 + 22224390 UUGU-----UGCUAGUUGCCAUCAAUGUCAACA-----AAAAAAAGAACCCUUUGCUACCAAUUAUUGGCCAAGUCCAAUAAACAGAAGGCUGCUGGAGGGGUAGGUGGGGUU ((((-----((...((((....))))..)))))-----).......((((((...(((((..(((((((......))))))).(((.......)))....)))))..)))))) ( -28.00) >DroSim_CAF1 4372 111 + 1 NNNN--NNNNNNNNNNNNNNNNNNNNNNCAACAAAAAAAAAAAAGGAACCCUAUGCAACCAAUUAUUGGCAAAGUCCAAUAAACAGAAGGCUGCUGGAGGGGUAGUUCGGGGU ....--.........................................(((((..((.(((..(((((((......))))))).(((.......)))....))).))..))))) ( -19.50) >DroYak_CAF1 4651 107 + 1 UU--GCUGCUGCUAGUUGACAUCAAUGUCAACACA---AAAAA-AAAACCCUAUGCAACCAAUUAUUGACCAAGUCCAAUAAACAGAAGGCUGCGGAAGGGGCAGGUGGGGGU ..--(((.(..((.(((((((....)))))))...---.....-....((((.((((.((..((((((........))))))......)).))))..))))...))..).))) ( -30.50) >consensus UU_______UGCUAGUUG_CAUCAAUGUCAACA_A___AAAAAAAGAACCCUAUGCAACCAAUUAUUGGCCAAGUCCAAUAAACAGAAGGCUGCUGGAGGGGUAGGUGGGGGU ..............(((((((....)))))))................((((.(((..((..((((((........)))))).(((....))).....)).)))...)))).. (-13.64 = -15.67 + 2.03)

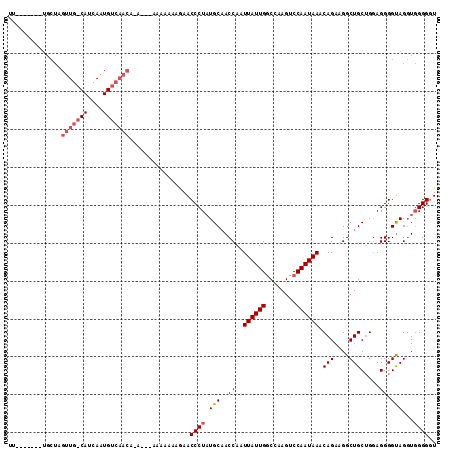
| Location | 19,094,597 – 19,094,700 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 70.36 |
| Mean single sequence MFE | -25.03 |
| Consensus MFE | -18.69 |
| Energy contribution | -18.80 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.75 |
| SVM decision value | 4.71 |
| SVM RNA-class probability | 0.999941 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
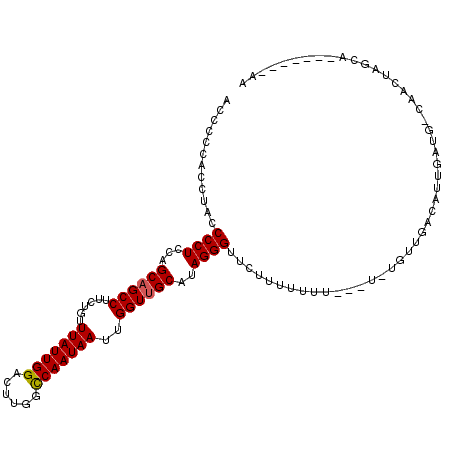
>X_DroMel_CAF1 19094597 103 - 22224390 AACCCCACCUACCCCUCCAGCAGCCUUCUGUUUAUUGGACUUGGCCAAUAAUUGGUAGCAAAGGGUUCUUUUUUU-----UGUUGACAUUGAUGGCAACUAGCA-----ACAA (((((...(((((......((((....))))(((((((......)))))))..)))))....))))).......(-----(((((...(((....)))....))-----)))) ( -23.30) >DroSim_CAF1 4372 111 - 1 ACCCCGAACUACCCCUCCAGCAGCCUUCUGUUUAUUGGACUUUGCCAAUAAUUGGUUGCAUAGGGUUCCUUUUUUUUUUUUGUUGNNNNNNNNNNNNNNNNNNNNNN--NNNN ............((((...((((((......(((((((......)))))))..))))))..))))..........................................--.... ( -20.40) >DroYak_CAF1 4651 107 - 1 ACCCCCACCUGCCCCUUCCGCAGCCUUCUGUUUAUUGGACUUGGUCAAUAAUUGGUUGCAUAGGGUUUU-UUUUU---UGUGUUGACAUUGAUGUCAACUAGCAGCAGC--AA ........((((((((...((((((......(((((((......)))))))..))))))..))))....-.....---((((((((((....)))))))..))))))).--.. ( -31.40) >consensus ACCCCCACCUACCCCUCCAGCAGCCUUCUGUUUAUUGGACUUGGCCAAUAAUUGGUUGCAUAGGGUUCUUUUUUU___U_UGUUGACAUUGAUG_CAACUAGCA_______AA ............((((...((((((......(((((((......)))))))..))))))..))))................................................ (-18.69 = -18.80 + 0.11)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:44:40 2006