
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 2,106,088 – 2,106,192 |
| Length | 104 |
| Max. P | 0.970925 |

| Location | 2,106,088 – 2,106,192 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 79.94 |
| Mean single sequence MFE | -25.77 |
| Consensus MFE | -17.55 |
| Energy contribution | -19.55 |
| Covariance contribution | 2.00 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.68 |
| SVM decision value | 1.67 |
| SVM RNA-class probability | 0.970925 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
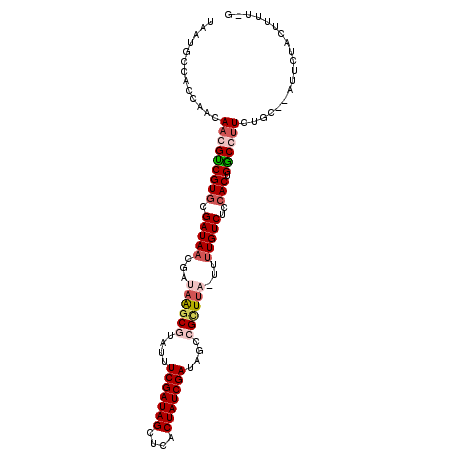
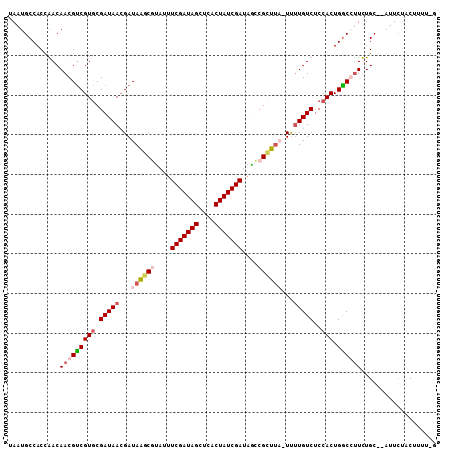
>X_DroMel_CAF1 2106088 104 + 22224390 UAAUGCCACUAACAACGUCGUGCGAUAACGAUAGGCAUAUUUCGAUAGUUCGCUAUCGAUAAUCGCU---UUUUGUCUCCACUGGCCUUUUGCGCAUUCUACUUUU-G .(((((((........((((((.((..((((.((((.....(((((((....))))))).....)))---).)))).))))).)))....)).)))))........-. ( -23.60) >DroSec_CAF1 6718 99 + 1 UAAUGCCACUAACAACGCCGUUCGAUAAAGAUAGGCUCA--UCGAUAGCUCACUAUCGAUAGUCGCUUAUUUUUGUCUCCACUGGCCUUUUGC--UUUCAACU----- ................(((((..(((((((((((((.((--(((((((....)))))))).)..)))))))))))))...)).))).......--........----- ( -27.50) >DroEre_CAF1 7297 105 + 1 UAAUGCCACCGACAACGUCGUGCGAUAACGAUAAUCGUAUUUCGAUAGCUCACUAUCGAUAGCCGCUUU-UCUUGUCUCCACUGACGCUCUGC--AUUUUACUUUUCA .(((((....((...(((((((.(((((.((.((.((..(.(((((((....))))))).)..)).)).-)))))))..))).)))).)).))--))).......... ( -23.10) >DroYak_CAF1 6619 105 + 1 UAAUGCCACCGACAACGUCGUGCGAUAACGAUAAGCGUACUUCGAUAGCUCUCUAUCGAUAGCCGUUUA-UUCUGUCUCCACUGGCGUUCUGC--AUUCUGCUUCUUG .(((((((..(((...)))(((.((((..((((((((....(((((((....)))))))....))))))-)).))))..))))))))))..((--.....))...... ( -28.90) >consensus UAAUGCCACCAACAACGUCGUGCGAUAACGAUAAGCGUAUUUCGAUAGCUCACUAUCGAUAGCCGCUUA_UUUUGUCUCCACUGGCCUUCUGC__AUUCUACUUUU_G .............(((((((((.(((((...((((((....(((((((....)))))))....))))))...)))))..))).))))))................... (-17.55 = -19.55 + 2.00)

| Location | 2,106,088 – 2,106,192 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 79.94 |
| Mean single sequence MFE | -27.82 |
| Consensus MFE | -21.27 |
| Energy contribution | -20.65 |
| Covariance contribution | -0.62 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.807865 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
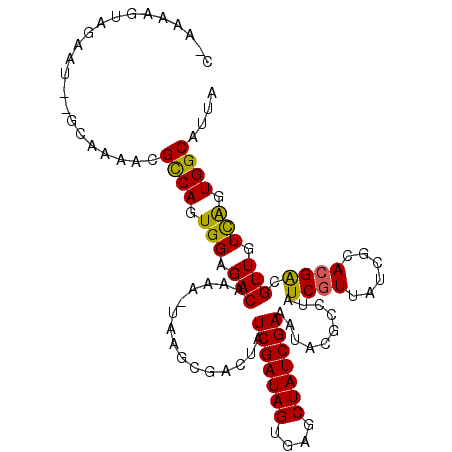
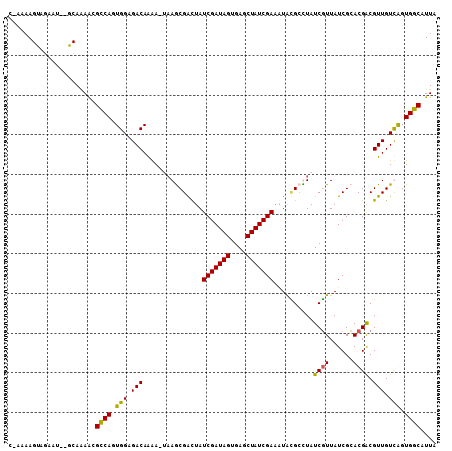
>X_DroMel_CAF1 2106088 104 - 22224390 C-AAAAGUAGAAUGCGCAAAAGGCCAGUGGAGACAAAA---AGCGAUUAUCGAUAGCGAACUAUCGAAAUAUGCCUAUCGUUAUCGCACGACGUUGUUAGUGGCAUUA .-....................((((.(((.(((....---.(((((...(((((((((....)))........))))))..))))).....))).))).)))).... ( -23.50) >DroSec_CAF1 6718 99 - 1 -----AGUUGAAA--GCAAAAGGCCAGUGGAGACAAAAAUAAGCGACUAUCGAUAGUGAGCUAUCGA--UGAGCCUAUCUUUAUCGAACGGCGUUGUUAGUGGCAUUA -----........--.......((((.((....))......(((((((((((((((....)))))))--)).(((..((......))..)))))))))..)))).... ( -29.30) >DroEre_CAF1 7297 105 - 1 UGAAAAGUAAAAU--GCAGAGCGUCAGUGGAGACAAGA-AAAGCGGCUAUCGAUAGUGAGCUAUCGAAAUACGAUUAUCGUUAUCGCACGACGUUGUCGGUGGCAUUA ..........(((--((.(((((((..((....))...-...((((...(((((((....)))))))...(((.....)))..))))..)))))..))....))))). ( -29.50) >DroYak_CAF1 6619 105 - 1 CAAGAAGCAGAAU--GCAGAACGCCAGUGGAGACAGAA-UAAACGGCUAUCGAUAGAGAGCUAUCGAAGUACGCUUAUCGUUAUCGCACGACGUUGUCGGUGGCAUUA ......((.....--)).....((((((((.(((.((.-(((.(((((.(((((((....)))))))))).)).)))))))).)))).((((...)))).)))).... ( -29.00) >consensus C_AAAAGUAGAAU__GCAAAACGCCAGUGGAGACAAAA_UAAGCGACUAUCGAUAGUGAGCUAUCGAAAUACGCCUAUCGUUAUCGCACGACGUUGUCAGUGGCAUUA ......................((((.(((.(((...............(((((((....)))))))..........((((......)))).))).))).)))).... (-21.27 = -20.65 + -0.62)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:53:18 2006