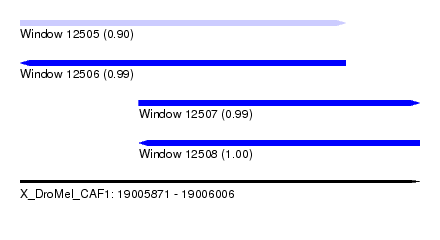

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 19,005,871 – 19,006,006 |
| Length | 135 |
| Max. P | 0.998063 |

| Location | 19,005,871 – 19,005,981 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 89.81 |
| Mean single sequence MFE | -23.13 |
| Consensus MFE | -18.01 |
| Energy contribution | -17.64 |
| Covariance contribution | -0.37 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.28 |
| Structure conservation index | 0.78 |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.899939 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19005871 110 + 22224390 UAUAGAAAAUCAUCACUUCCUGGCACAUUGAAAAUCACACAAUUCAAACAGCUCGCGUCAAGCGCCAUUUGUCAAAGUAUCUAAAUACUUGAGCAAUCAGAUACUUUGAC ....................((((...(((((..........))))).......((.....))))))...((((((((((((........((....)))))))))))))) ( -23.00) >DroSec_CAF1 28327 106 + 1 UAUAGAAUAUCAUCACUUCCUGGCACUUUGGAAAUCGCA---UUCAAACAACUCGCGUCAAGCGCCAU-UGUCAAAGUAUCUAAAUACUUGAGCAAUCAGAUACUUUGGC ....................((((..((((((.......---))))))......((.....)))))).-.((((((((((((........((....)))))))))))))) ( -22.90) >DroSim_CAF1 22923 106 + 1 UAUAGAAAAUCAUCACUUCCUGGCACUUUGGAAAUCGCA---UUCAAACAGCUCGCGUCAAGCGCCAU-UGUCAAAGUAUCUAAAUACUUGAGCAAUCAGAUACUUUGGC ....................((((..((((((.......---))))))......((.....)))))).-.((((((((((((........((....)))))))))))))) ( -22.90) >DroYak_CAF1 28535 102 + 1 UACAGAAAAUCACCAUUUCCUGGC----CGCCAAUCGCA---UUCAAACAACUCGCGUCAAGCGCCAU-UGUCAAUGUAUCUAAAUGCUUGAGCAAUCAGAUACUUUGAC ....................((((----.((....(((.---............)))....)))))).-.(((((.((((((.(.(((....))).).)))))).))))) ( -23.72) >consensus UAUAGAAAAUCAUCACUUCCUGGCACUUUGGAAAUCGCA___UUCAAACAACUCGCGUCAAGCGCCAU_UGUCAAAGUAUCUAAAUACUUGAGCAAUCAGAUACUUUGAC ....................((((....((.....)).................((.....))))))...((((((((((((................)))))))))))) (-18.01 = -17.64 + -0.37)

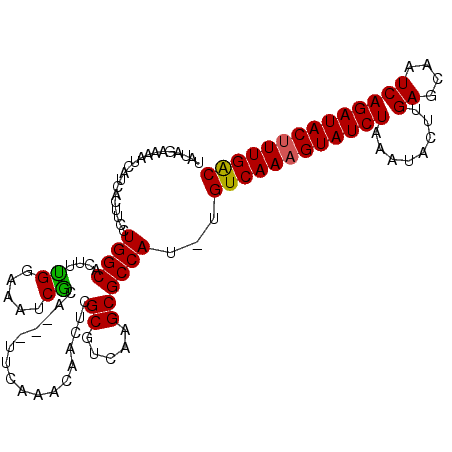
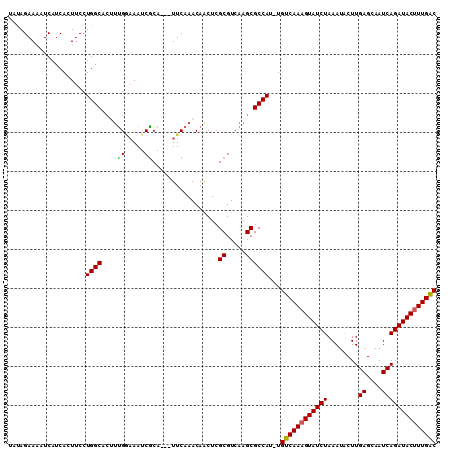
| Location | 19,005,871 – 19,005,981 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 89.81 |
| Mean single sequence MFE | -35.95 |
| Consensus MFE | -30.20 |
| Energy contribution | -31.20 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.11 |
| Mean z-score | -3.80 |
| Structure conservation index | 0.84 |
| SVM decision value | 2.23 |
| SVM RNA-class probability | 0.990829 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19005871 110 - 22224390 GUCAAAGUAUCUGAUUGCUCAAGUAUUUAGAUACUUUGACAAAUGGCGCUUGACGCGAGCUGUUUGAAUUGUGUGAUUUUCAAUGUGCCAGGAAGUGAUGAUUUUCUAUA ((((((((((((((.(((....))).))))))))))))))....((((((((((((((..........)))))......)))).)))))(((((((....)))))))... ( -37.80) >DroSec_CAF1 28327 106 - 1 GCCAAAGUAUCUGAUUGCUCAAGUAUUUAGAUACUUUGACA-AUGGCGCUUGACGCGAGUUGUUUGAA---UGCGAUUUCCAAAGUGCCAGGAAGUGAUGAUAUUCUAUA ..((((((((((((.(((....))).))))))))))))...-.((((((((...(.(((((((.....---.))))))).).))))))))((((.........))))... ( -34.60) >DroSim_CAF1 22923 106 - 1 GCCAAAGUAUCUGAUUGCUCAAGUAUUUAGAUACUUUGACA-AUGGCGCUUGACGCGAGCUGUUUGAA---UGCGAUUUCCAAAGUGCCAGGAAGUGAUGAUUUUCUAUA ..((((((((((((.(((....))).))))))))))))...-..(((((((..((((..(.....)..---)))).......)))))))(((((((....)))))))... ( -34.30) >DroYak_CAF1 28535 102 - 1 GUCAAAGUAUCUGAUUGCUCAAGCAUUUAGAUACAUUGACA-AUGGCGCUUGACGCGAGUUGUUUGAA---UGCGAUUGGCG----GCCAGGAAAUGGUGAUUUUCUGUA (((((.((((((((.(((....))).)))))))).))))).-...........(((.((((((.....---.)))))).)))----..((((((((....)))))))).. ( -37.10) >consensus GCCAAAGUAUCUGAUUGCUCAAGUAUUUAGAUACUUUGACA_AUGGCGCUUGACGCGAGCUGUUUGAA___UGCGAUUUCCAAAGUGCCAGGAAGUGAUGAUUUUCUAUA ((((((((((((((.(((....))).))))))))))))))....((((((((.((((...(......)...)))).....))).)))))(((((((....)))))))... (-30.20 = -31.20 + 1.00)

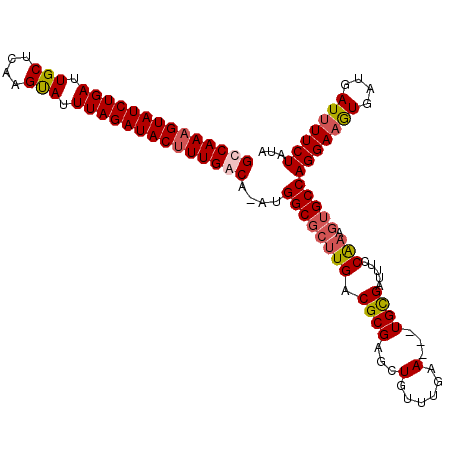
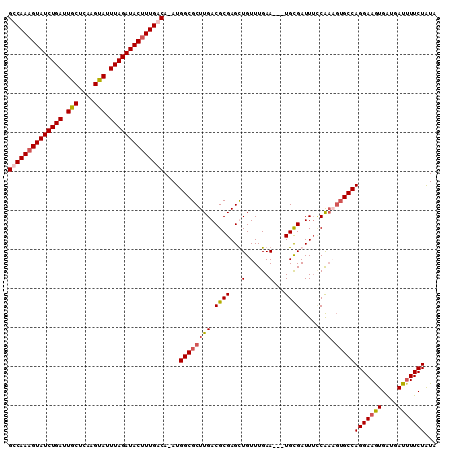
| Location | 19,005,911 – 19,006,006 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 88.41 |
| Mean single sequence MFE | -23.04 |
| Consensus MFE | -18.59 |
| Energy contribution | -18.67 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.81 |
| SVM decision value | 2.09 |
| SVM RNA-class probability | 0.987589 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19005911 95 + 22224390 AAUUCAAACAGCUCGCGUCAAGCGCCAUUUGUCAAAGUAUCUAAAUACUUGAGCAAUCAGAUACUUUGACACCUGCGCCUGCAAUCCGAAUCGUU ............(((.((((.((((....(((((((((((((........((....)))))))))))))))...)))).))..)).)))...... ( -25.00) >DroSec_CAF1 28366 92 + 1 --UUCAAACAACUCGCGUCAAGCGCCAU-UGUCAAAGUAUCUAAAUACUUGAGCAAUCAGAUACUUUGGCACCUCCGCCUGCAAUCCGAAUCGUU --..........(((.((((.(((....-(((((((((((((........((....)))))))))))))))....))).))..)).)))...... ( -21.40) >DroSim_CAF1 22962 92 + 1 --UUCAAACAGCUCGCGUCAAGCGCCAU-UGUCAAAGUAUCUAAAUACUUGAGCAAUCAGAUACUUUGGCACCUCCGCCUGCAAUCCGAAUCGUU --..........(((.((((.(((....-(((((((((((((........((....)))))))))))))))....))).))..)).)))...... ( -21.40) >DroEre_CAF1 27380 82 + 1 ------------UCGGCGCAAGCGGCAU-UGCCAAAGUAUCUAAAUACUUGAGCCAUCAGAUACUUUGACACCUCCGUCUGCAAUCCGAAUCGUG ------------((((.(((..(((...-((.((((((((((................)))))))))).))...)))..)))...))))...... ( -25.49) >DroYak_CAF1 28570 92 + 1 --UUCAAACAACUCGCGUCAAGCGCCAU-UGUCAAUGUAUCUAAAUGCUUGAGCAAUCAGAUACUUUGACACCUCCGCCUGCAAUCCGAAUCGUG --..........(((.((((.(((....-((((((.((((((.(.(((....))).).)))))).))))))....))).))..)).)))...... ( -21.90) >consensus __UUCAAACAACUCGCGUCAAGCGCCAU_UGUCAAAGUAUCUAAAUACUUGAGCAAUCAGAUACUUUGACACCUCCGCCUGCAAUCCGAAUCGUU ............(((...((.(((.....(((((((((((((................)))))))))))))....))).)).....)))...... (-18.59 = -18.67 + 0.08)
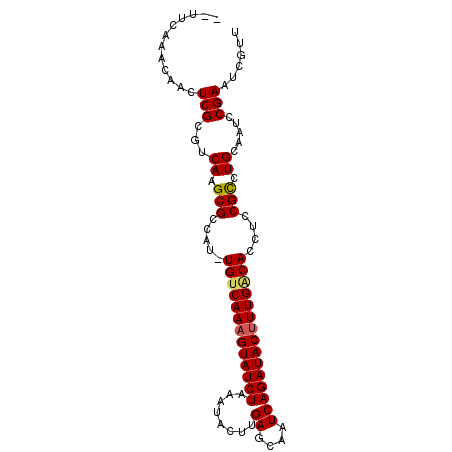


| Location | 19,005,911 – 19,006,006 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 88.41 |
| Mean single sequence MFE | -37.76 |
| Consensus MFE | -29.02 |
| Energy contribution | -30.58 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.06 |
| Mean z-score | -5.03 |
| Structure conservation index | 0.77 |
| SVM decision value | 3.00 |
| SVM RNA-class probability | 0.998063 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 19005911 95 - 22224390 AACGAUUCGGAUUGCAGGCGCAGGUGUCAAAGUAUCUGAUUGCUCAAGUAUUUAGAUACUUUGACAAAUGGCGCUUGACGCGAGCUGUUUGAAUU ...(((((((((..(((((((...(((((((((((((((.(((....))).)))))))))))))))....))))))).(....)..))))))))) ( -41.90) >DroSec_CAF1 28366 92 - 1 AACGAUUCGGAUUGCAGGCGGAGGUGCCAAAGUAUCUGAUUGCUCAAGUAUUUAGAUACUUUGACA-AUGGCGCUUGACGCGAGUUGUUUGAA-- (((((((((.....((((((....((.((((((((((((.(((....))).)))))))))))).))-....))))))...)))))))))....-- ( -36.20) >DroSim_CAF1 22962 92 - 1 AACGAUUCGGAUUGCAGGCGGAGGUGCCAAAGUAUCUGAUUGCUCAAGUAUUUAGAUACUUUGACA-AUGGCGCUUGACGCGAGCUGUUUGAA-- .....(((((((.((..(((.((((((((((((((((((.(((....))).)))))))))).....-.))))))))..)))..)).)))))))-- ( -36.00) >DroEre_CAF1 27380 82 - 1 CACGAUUCGGAUUGCAGACGGAGGUGUCAAAGUAUCUGAUGGCUCAAGUAUUUAGAUACUUUGGCA-AUGCCGCUUGCGCCGA------------ ......((((...(((..(((...(((((((((((((((..((....))..)))))))))))))))-...)))..))).))))------------ ( -36.20) >DroYak_CAF1 28570 92 - 1 CACGAUUCGGAUUGCAGGCGGAGGUGUCAAAGUAUCUGAUUGCUCAAGCAUUUAGAUACAUUGACA-AUGGCGCUUGACGCGAGUUGUUUGAA-- .((((((((.....((((((....((((((.((((((((.(((....))).)))))))).))))))-....))))))...)))))))).....-- ( -38.50) >consensus AACGAUUCGGAUUGCAGGCGGAGGUGUCAAAGUAUCUGAUUGCUCAAGUAUUUAGAUACUUUGACA_AUGGCGCUUGACGCGAGCUGUUUGAA__ .(((.((((.....((((((....(((((((((((((((.(((....))).))))))))))))))).....))))))...)))).)))....... (-29.02 = -30.58 + 1.56)
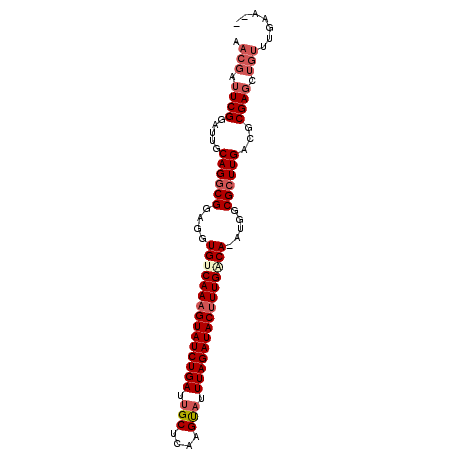


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:43:34 2006