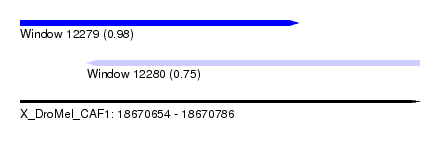

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,670,654 – 18,670,786 |
| Length | 132 |
| Max. P | 0.980473 |

| Location | 18,670,654 – 18,670,746 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 74.43 |
| Mean single sequence MFE | -19.52 |
| Consensus MFE | -11.90 |
| Energy contribution | -11.65 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.61 |
| SVM decision value | 1.86 |
| SVM RNA-class probability | 0.980473 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
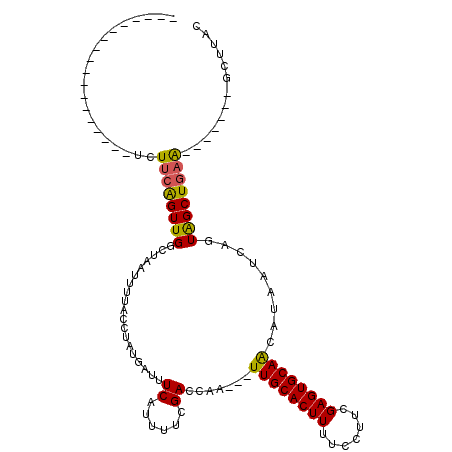
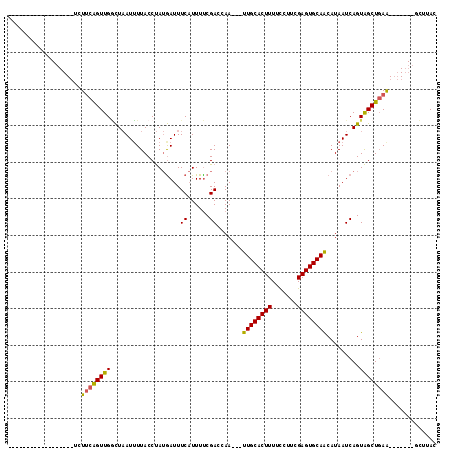
>X_DroMel_CAF1 18670654 92 + 22224390 ------------------UCUGCAGUUGAUUAAUUUUACCUAUAAUUUCAUUUUCGACCAAUUAUUGCACUUUUCCUUCGAGUGCAACAUAAUCAGUAGCUGAG-------UCUUAC ------------------((.((..(((((((..........(((((((......))..)))))((((((((.......))))))))..)))))))..)).)).-------...... ( -17.40) >DroSec_CAF1 15590 89 + 1 ------------------UCUUCGGUUGGCUAAUUUUACCUAUGAUUUCAUUUUCGACCAA---UUGCACUUUUCCUUCGAGUGCAACAUAAUCAGUAGCUGAA-------GCUUAC ------------------.(((((((((.((...........(((........))).....---((((((((.......)))))))).......))))))))))-------)..... ( -18.70) >DroSim_CAF1 13744 89 + 1 ------------------UCUUCAGUUGGCUAAUUUUACCUAUGAUUUCAUUUUCGAUGAA---UUGCACUUUUCCUUCGAGUGCAACAUAAUCAGUAGCUGAA-------GCUUAC ------------------.(((((((((.((...............((((((...))))))---((((((((.......)))))))).......))))))))))-------)..... ( -21.40) >DroYak_CAF1 15767 114 + 1 UACAAGCCAAACUCAAAUUCUUUAGUUGCCUCAUGUUACCCAAGAUUUCCUUUCCGAACAA---UUGCACUUUUUUUUCGAGUGCAGCAUAAUCAGUGGCUUAAUGCUCACUGCUAC ....(((...................(((....((((....(((.....)))....)))).---.(((((((.......)))))))))).....((((((.....)).))))))).. ( -20.60) >consensus __________________UCUUCAGUUGGCUAAUUUUACCUAUGAUUUCAUUUUCGACCAA___UUGCACUUUUCCUUCGAGUGCAACAUAAUCAGUAGCUGAA_______GCUUAC ....................((((((((...................((......)).......((((((((.......)))))))).........))))))))............. (-11.90 = -11.65 + -0.25)

| Location | 18,670,676 – 18,670,786 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 83.93 |
| Mean single sequence MFE | -28.20 |
| Consensus MFE | -17.55 |
| Energy contribution | -19.18 |
| Covariance contribution | 1.63 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.751793 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
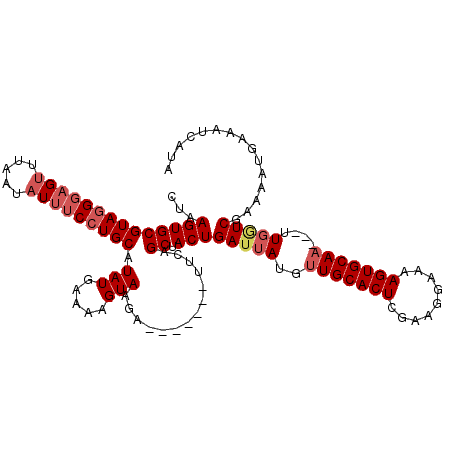
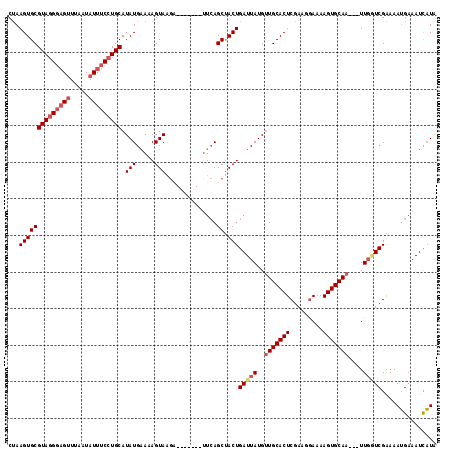
>X_DroMel_CAF1 18670676 110 - 22224390 CUAAGUGCGUAGGGAGUUUAAUAUUUCCUGCAUAUGAAAAGUAAGA-------CUCAGCUACUGAUUAUGUUGCACUCGAAGGAAAAGUGCAAUAAUUGGUCGAAAAUGAAAUUAUA ...((((((((((((((.....)))))))))...(((.........-------.))))).)))(((((((((((((((....))...))))))))..)))))............... ( -27.30) >DroSec_CAF1 15612 107 - 1 CUAAGUGCGUAGGGUGUUUAAUAUUUCCUGCAUAUGGAAAGUAAGC-------UUCAGCUACUGAUUAUGUUGCACUCGAAGGAAAAGUGCAA---UUGGUCGAAAAUGAAAUCAUA ...((((.((.(((.(((((...(((((.......))))).)))))-------))).))))))(((((.(((((((((....))...))))))---))))))....(((....))). ( -26.10) >DroSim_CAF1 13766 107 - 1 CUAAGUGCGUAGGGAGUUUAAUAUUUCCUGCAUAUGGAGAGUAAGC-------UUCAGCUACUGAUUAUGUUGCACUCGAAGGAAAAGUGCAA---UUCAUCGAAAAUGAAAUCAUA ...((((((((((((((.....)))))))))...(((((......)-------)))))).)))((((..(((((((((....))...))))))---)((((.....))))))))... ( -30.00) >DroYak_CAF1 15807 113 - 1 CCAAGUGCGUAUGCAGCUUAGUAUUUCCUGCAUAUGAA-AGUAGCAGUGAGCAUUAAGCCACUGAUUAUGCUGCACUCGAAAAAAAAGUGCAA---UUGUUCGGAAAGGAAAUCUUG ((.(((((((((((((...........))))))))...-((((.(((((.((.....)))))))....)))))))))((((....((......---)).))))....))........ ( -29.40) >consensus CUAAGUGCGUAGGGAGUUUAAUAUUUCCUGCAUAUGAAAAGUAAGA_______UUCAGCUACUGAUUAUGUUGCACUCGAAGGAAAAGUGCAA___UUGGUCGAAAAUGAAAUCAUA ...((((((((((((((.....))))))))).(((.....)))..............)).)))(((((..(((((((.........)))))))....)))))............... (-17.55 = -19.18 + 1.63)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:40:07 2006