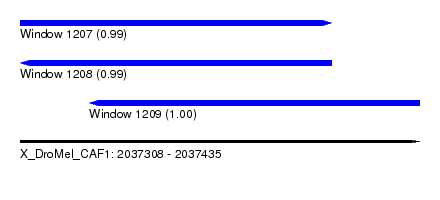

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 2,037,308 – 2,037,435 |
| Length | 127 |
| Max. P | 0.997359 |

| Location | 2,037,308 – 2,037,407 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 88.92 |
| Mean single sequence MFE | -26.64 |
| Consensus MFE | -22.40 |
| Energy contribution | -24.00 |
| Covariance contribution | 1.60 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.45 |
| Structure conservation index | 0.84 |
| SVM decision value | 2.53 |
| SVM RNA-class probability | 0.994951 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
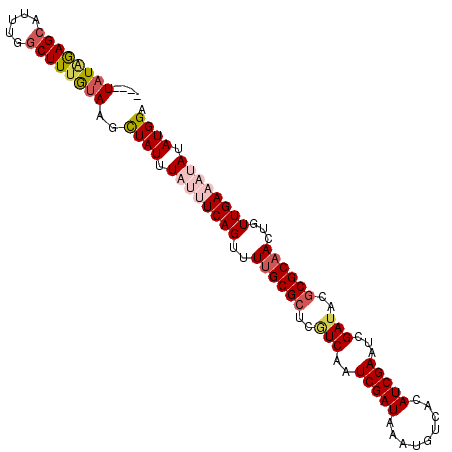
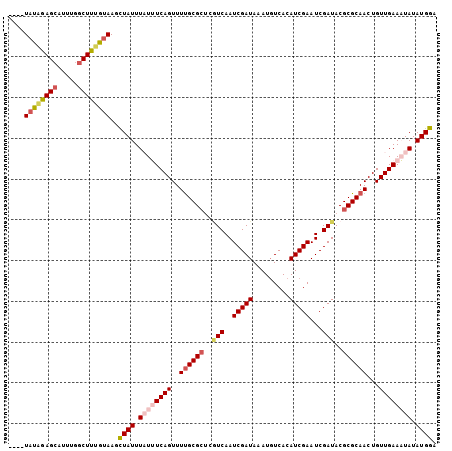
>X_DroMel_CAF1 2037308 99 + 22224390 ----U-UAAAGCAUU-AUCUUUGUAAUUUAUUUAUUUCAGUUUCGCGCUGAUCAAUCGAUAAAUGUCACAUCGAAUCGAUACGCGCAACUGUUGAAAUAUAUGGA ----.-(((((....-..)))))...(((((.((((((((....((((..(((..(((((.........)))))...)))..)))).....)))))))).))))) ( -18.10) >DroSec_CAF1 3999 101 + 1 ----UAUAGAGCAUUUGGCUUUGUAAGCUAUUUAUUUCAGUUUUGCGCUGGUCAAUCGAUAAAUGUCACAUCGAAUCGAUACGCGCAACUGUUGAAAUAUAUGGA ----((((((((.....))))))))..((((.((((((((..((((((..(((..(((((.........)))))...)))..))))))...)))))))).)))). ( -30.00) >DroSim_CAF1 3990 97 + 1 ----UAUAGAGCAUUUGGCUUUGUAAGCUAUUUAUUUCAGUUUUGCG----UCAAUCGAUAAAUGUCACAUCGAAUCGAUACGCGCAACUGUUGAAAUAUAUGGA ----((((((((.....))))))))..((((.((((((((..(((((----(..((((((..(((...)))...))))))..))))))...)))))))).)))). ( -27.90) >DroEre_CAF1 4020 105 + 1 UGCUUAUAGAGCAUUUGGCUUCAUAAGCUAUUUUUGUCAGUUUUGCGCUCGUCAAUCGAUAAAUUUCACAUCGAAUCGAUACGCGCAACUGUUGAAUUAUAUGGA .((((((..(((.....)))..))))))....((((.((((..(((((..(((..(((((.........)))))...)))..))))))))).))))......... ( -27.20) >DroYak_CAF1 3989 105 + 1 UGCUUAUGGAGCAUUUGGCUUUAUAAGCUAUUUUUGUCAGUUUUGCGCUCGUCAAUCGAUAAAUGUCACAUCGAAUCGAUACGCGCAACUGUUGAAUUAUAUGGA .(((((((((((.....)))))))))))....((((.((((..(((((..(((..(((((.........)))))...)))..))))))))).))))......... ( -30.00) >consensus ____UAUAGAGCAUUUGGCUUUGUAAGCUAUUUAUUUCAGUUUUGCGCUCGUCAAUCGAUAAAUGUCACAUCGAAUCGAUACGCGCAACUGUUGAAAUAUAUGGA ....((((((((.....))))))))..((((.((((((((..((((((..(((..(((((.........)))))...)))..))))))...)))))))).)))). (-22.40 = -24.00 + 1.60)

| Location | 2,037,308 – 2,037,407 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 88.92 |
| Mean single sequence MFE | -26.20 |
| Consensus MFE | -19.88 |
| Energy contribution | -21.12 |
| Covariance contribution | 1.24 |
| Combinations/Pair | 1.04 |
| Mean z-score | -3.42 |
| Structure conservation index | 0.76 |
| SVM decision value | 2.45 |
| SVM RNA-class probability | 0.994094 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
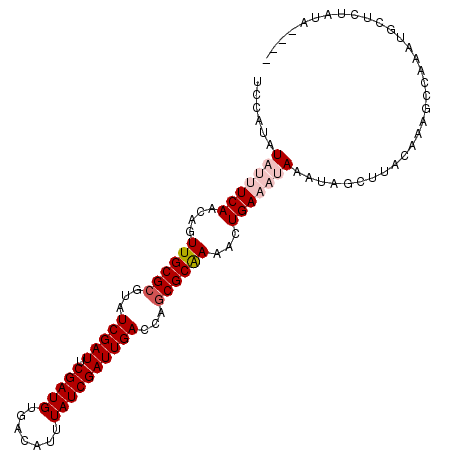
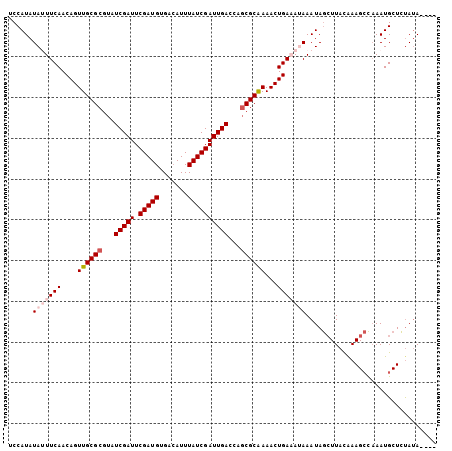
>X_DroMel_CAF1 2037308 99 - 22224390 UCCAUAUAUUUCAACAGUUGCGCGUAUCGAUUCGAUGUGACAUUUAUCGAUUGAUCAGCGCGAAACUGAAAUAAAUAAAUUACAAAGAU-AAUGCUUUA-A---- ......(((((((....((((((..((((((.(((((.......)))))))))))..))))))...)))))))..........((((..-....)))).-.---- ( -24.80) >DroSec_CAF1 3999 101 - 1 UCCAUAUAUUUCAACAGUUGCGCGUAUCGAUUCGAUGUGACAUUUAUCGAUUGACCAGCGCAAAACUGAAAUAAAUAGCUUACAAAGCCAAAUGCUCUAUA---- ......(((((((....((((((...(((((.(((((.......))))))))))...))))))...)))))))....(((.....))).............---- ( -25.00) >DroSim_CAF1 3990 97 - 1 UCCAUAUAUUUCAACAGUUGCGCGUAUCGAUUCGAUGUGACAUUUAUCGAUUGA----CGCAAAACUGAAAUAAAUAGCUUACAAAGCCAAAUGCUCUAUA---- ......(((((((....(((((((.((((((..((((...)))).)))))))).----)))))...)))))))....(((.....))).............---- ( -19.80) >DroEre_CAF1 4020 105 - 1 UCCAUAUAAUUCAACAGUUGCGCGUAUCGAUUCGAUGUGAAAUUUAUCGAUUGACGAGCGCAAAACUGACAAAAAUAGCUUAUGAAGCCAAAUGCUCUAUAAGCA ..............(((((((((...(((((.(((((.......))))))))))...)))))..)))).........(((((((.(((.....))).))))))). ( -31.50) >DroYak_CAF1 3989 105 - 1 UCCAUAUAAUUCAACAGUUGCGCGUAUCGAUUCGAUGUGACAUUUAUCGAUUGACGAGCGCAAAACUGACAAAAAUAGCUUAUAAAGCCAAAUGCUCCAUAAGCA ..............(((((((((...(((((.(((((.......))))))))))...)))))..)))).........((((((..(((.....)))..)))))). ( -29.90) >consensus UCCAUAUAUUUCAACAGUUGCGCGUAUCGAUUCGAUGUGACAUUUAUCGAUUGACCAGCGCAAAACUGAAAUAAAUAGCUUACAAAGCCAAAUGCUCUAUA____ ......(((((((....((((((...(((((.(((((.......))))))))))...))))))...)))))))................................ (-19.88 = -21.12 + 1.24)

| Location | 2,037,330 – 2,037,435 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 90.54 |
| Mean single sequence MFE | -25.68 |
| Consensus MFE | -21.80 |
| Energy contribution | -22.84 |
| Covariance contribution | 1.04 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.85 |
| SVM decision value | 2.84 |
| SVM RNA-class probability | 0.997359 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 2037330 105 - 22224390 AAGCUGGGAAAACAAACGGCUGAUAGCUUCCAUAUAUUUCAACAGUUGCGCGUAUCGAUUCGAUGUGACAUUUAUCGAUUGAUCAGCGCGAAACUGAAAUAAAUA .(((((..........))))).............(((((((....((((((..((((((.(((((.......)))))))))))..))))))...))))))).... ( -29.90) >DroSec_CAF1 4023 105 - 1 AAGCUGGGAAAACAAACGGCGUAUAGCUUCCAUAUAUUUCAACAGUUGCGCGUAUCGAUUCGAUGUGACAUUUAUCGAUUGACCAGCGCAAAACUGAAAUAAAUA ..((((..........))))..............(((((((....((((((...(((((.(((((.......))))))))))...))))))...))))))).... ( -28.50) >DroSim_CAF1 4014 101 - 1 AAGCUGGGAAAACAAACGUCGUAUAGCUUCCAUAUAUUUCAACAGUUGCGCGUAUCGAUUCGAUGUGACAUUUAUCGAUUGA----CGCAAAACUGAAAUAAAUA ((((((..(..((....))..).)))))).....(((((((....(((((((.((((((..((((...)))).)))))))).----)))))...))))))).... ( -22.80) >DroEre_CAF1 4048 102 - 1 AAACUGAUAAAACAAACGGU---UAGCUUCCAUAUAAUUCAACAGUUGCGCGUAUCGAUUCGAUGUGAAAUUUAUCGAUUGACGAGCGCAAAACUGACAAAAAUA .(((((..........))))---)..................(((((((((...(((((.(((((.......))))))))))...)))))..))))......... ( -22.50) >DroYak_CAF1 4017 102 - 1 AAGCUGAGAAAAUAAACGGU---UAGCUUCCAUAUAAUUCAACAGUUGCGCGUAUCGAUUCGAUGUGACAUUUAUCGAUUGACGAGCGCAAAACUGACAAAAAUA (((((((............)---)))))).............(((((((((...(((((.(((((.......))))))))))...)))))..))))......... ( -24.70) >consensus AAGCUGGGAAAACAAACGGC__AUAGCUUCCAUAUAUUUCAACAGUUGCGCGUAUCGAUUCGAUGUGACAUUUAUCGAUUGACCAGCGCAAAACUGAAAUAAAUA ..((((..........))))..............(((((((....((((((...(((((.(((((.......))))))))))...))))))...))))))).... (-21.80 = -22.84 + 1.04)

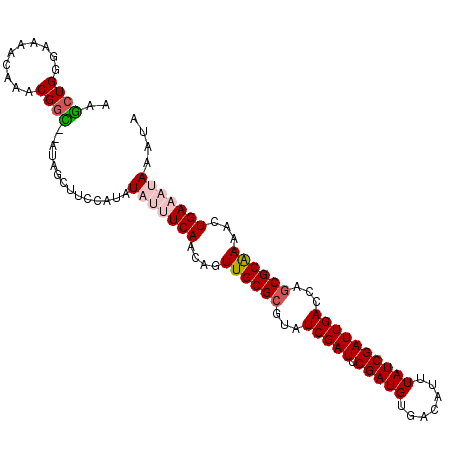
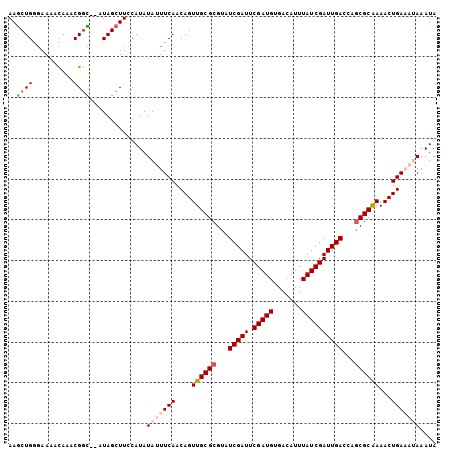
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:52:48 2006