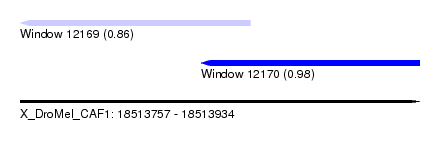
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,513,757 – 18,513,934 |
| Length | 177 |
| Max. P | 0.981870 |

| Location | 18,513,757 – 18,513,859 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 71.93 |
| Mean single sequence MFE | -39.13 |
| Consensus MFE | -20.09 |
| Energy contribution | -22.87 |
| Covariance contribution | 2.78 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.51 |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.855145 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18513757 102 - 22224390 UGCGUUGGAUAGUUGGCUUC--U----------------GUGGACUUUUGUGGGUAUAUUCCGCAAGGGCAAGGGCUGCAGCUUCCCAGCUGCUACGAGAUCUGACCAAGCUCUAGCCCA ..........((((.((...--.----------------)).)))).(((((((.....)))))))((((.((((((((((((....))))))...(....)......)))))).)))). ( -37.10) >DroSec_CAF1 22065 102 - 1 UGCGUUGGAUAGUUGGCUUC--U----------------CUGGGCUUUUGUGGGUAUAUUCCGCAAGGGCAAGGGCUGCAGCUUCUCAGCUGCUGCGAGAUCUGGCCAAGAUAUUGCCCA .......((((.((((((((--(----------------(...(((((((((((.....)))))))))))....((.((((((....)))))).))))))...))))))..))))..... ( -41.20) >DroEre_CAF1 21821 111 - 1 UGUGUUGGCU-GUUAUCUUUUGUGGGCUUUUUGGGCUUUUUGGGCUUUUGUGGGCAUAUUCCGCAAGGGCAAGGGCUGCAGCUUCCU--------CAAGAUUCGAGCAAGAUACAGCCGA ....((((((-((.((((..(((..(..(((((((........(((((((((((.....)))))))))))..((((....)))).))--------)))))..)..))))))))))))))) ( -39.10) >consensus UGCGUUGGAUAGUUGGCUUC__U________________CUGGGCUUUUGUGGGUAUAUUCCGCAAGGGCAAGGGCUGCAGCUUCCCAGCUGCU_CGAGAUCUGACCAAGAUAUAGCCCA ...........................................(((((((((((.....)))))))))))..((((.(((((......)))))......((((.....))))...)))). (-20.09 = -22.87 + 2.78)

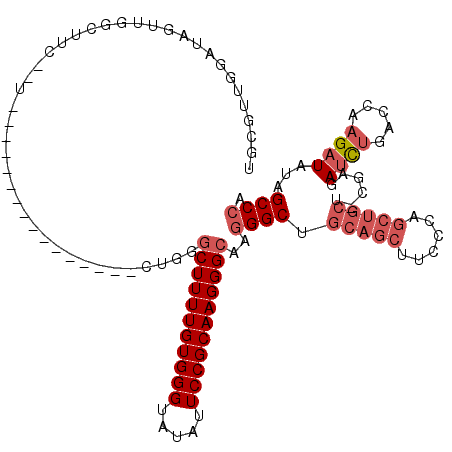
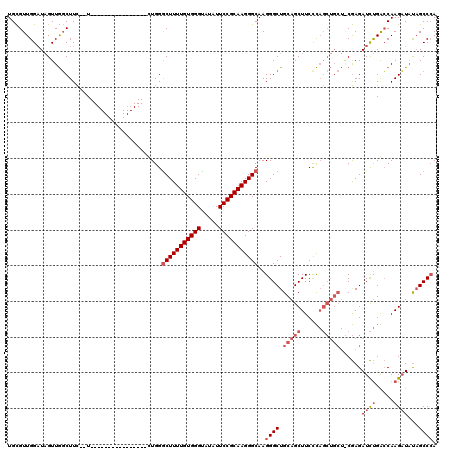
| Location | 18,513,837 – 18,513,934 |
|---|---|
| Length | 97 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 69.45 |
| Mean single sequence MFE | -30.45 |
| Consensus MFE | -21.80 |
| Energy contribution | -23.05 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.72 |
| SVM decision value | 1.90 |
| SVM RNA-class probability | 0.981870 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18513837 97 - 22224390 -GUCGGCUAACCAGAUCUUAAAGUUAAGAUCUCAGAUCGAGACCCUUCGUCUCAGUCGUGGGCAAUCAGCCAAGGUUGCGUUGGAUAGUUGGCUUC--U----------------G -((((((((.((((((((((....)))))))((((((.(((((.....))))).))).)))((((((......))))))..))).))))))))...--.----------------. ( -35.60) >DroSec_CAF1 22145 83 - 1 -GUCGGCUAACCAGAUCUUAAAGUUAAGAUCUCAGAUCGAGACCCUUCGUCUC--------------GCCCAAGGUUGCGUUGGAUAGUUGGCUUC--U----------------C -((((((((.((((((((((....)))))))...(..((((((.....)))))--------------)..)..........))).))))))))...--.----------------. ( -27.20) >DroEre_CAF1 21893 114 - 1 AGAUGCUUAACCAGAUCUUAGAGCUAAGACCUCAGAUCGAAACCCUUGGUCUCACUCGUGGACAAUCGC-CAAGGUUGUGUUGGCU-GUUAUCUUUUGUGGGCUUUUUGGGCUUUU .............(((((..(((.......))))))))(((((((..((((.(((....(((.(((.((-(((.......))))).-))).)))...)))))))....))).)))) ( -27.40) >DroYak_CAF1 22887 113 - 1 AGUCGGCUAACCAGAUCUUAAAGUUAAGAUCUCGGAACGAAACCCUUGGUCUUACUCAUAGAUAAUCACACAAGGUUGUCUUGGAUAGUUGGCUUC--UUGGCUUCUUGG-CUUCU (((((((((.((((.......(((.((((((..((........))..)))))))))....(((((((......))))))))))).)))))))))..--..((((....))-))... ( -31.60) >consensus _GUCGGCUAACCAGAUCUUAAAGUUAAGAUCUCAGAUCGAAACCCUUCGUCUCACUCGUGGACAAUCACCCAAGGUUGCGUUGGAUAGUUGGCUUC__U________________U .((((((((.((((((((((....))))))).......(((((.....)))))........((((((......))))))..))).))))))))....................... (-21.80 = -23.05 + 1.25)

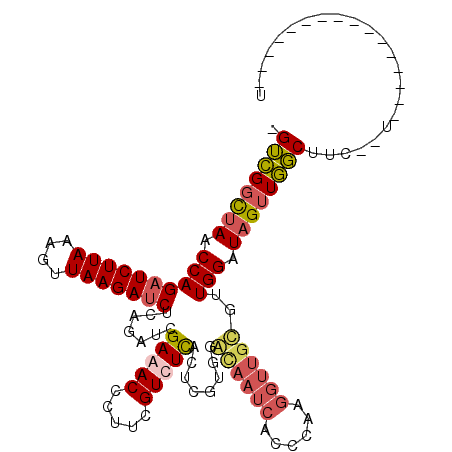
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:38:31 2006