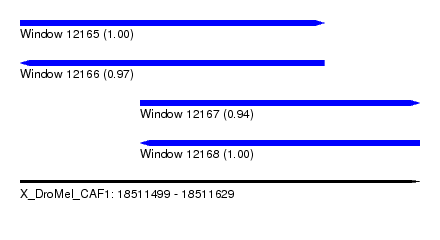

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,511,499 – 18,511,629 |
| Length | 130 |
| Max. P | 0.998714 |

| Location | 18,511,499 – 18,511,598 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.63 |
| Mean single sequence MFE | -33.40 |
| Consensus MFE | -26.61 |
| Energy contribution | -26.61 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.40 |
| Structure conservation index | 0.80 |
| SVM decision value | 3.20 |
| SVM RNA-class probability | 0.998714 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18511499 99 + 22224390 UCAGCUGGCAUAAUACCAAGCCAAAGCCAUAUAGCCAUA--GCUAAAGCCAUA----UAGCC---------------AUAUAGCCAUAGCCUACGGUUGUAGGCCAGCGGUCAACGACCG ...((((((..........((....))..(((((((.((--((((..((.(((----(....---------------)))).))..))).))).))))))).))))))((((...)))). ( -31.90) >DroSec_CAF1 19876 91 + 1 UCAGCUGGCAUAUUACCAAGCCAAAGCCAUAUAGCCAUA--GCUAAAGCCAUA---------------------------UAGCCAUAGCCUACGGUUGUAGGCCAGCGGUCAACGACCG ...((((((..........((....))..(((((((.((--((((..((....---------------------------..))..))).))).))))))).))))))((((...)))). ( -30.40) >DroEre_CAF1 19535 114 + 1 UCAGCUGGCAUAAUACGCAGCCAAAGCCAUAUAGCCAUA--GCUAAAGCCAUA----UAGCCAUAUAGCCAUAUAGCAUAAAGCCAUAGCCUACGGUUGUAGGCCAGCGGUCAACGACCG ...((((((.......((.......))..(((((((.((--((((..((..((----(.((.((((....)))).)))))..))..))).))).))))))).))))))((((...)))). ( -34.40) >DroYak_CAF1 20599 119 + 1 UCAGCUGGCAUAAUACCAAGCCAAAGCCAUAUAGCCAUAUAGCUAAAGCCAUAGCUAAAGCCAUAUAGCUAAA-GCCAUAUAGCCAUAGCCUACGGUUGUAGGCCAGCGGUCAACGACCG ...((((((..........((....))..(((((((.(((.((....)).)))((((..((.((((.((....-)).)))).))..))))....))))))).))))))((((...)))). ( -36.90) >consensus UCAGCUGGCAUAAUACCAAGCCAAAGCCAUAUAGCCAUA__GCUAAAGCCAUA____UAGCC_______________AUAUAGCCAUAGCCUACGGUUGUAGGCCAGCGGUCAACGACCG ...((((((..........((....))..(((((((.....((((..((.................................))..))))....))))))).))))))((((...)))). (-26.61 = -26.61 + -0.00)

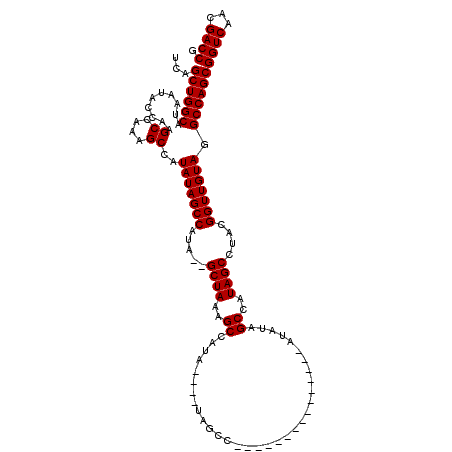
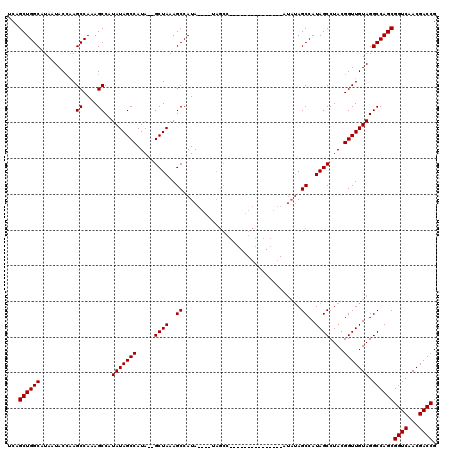
| Location | 18,511,499 – 18,511,598 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.63 |
| Mean single sequence MFE | -49.55 |
| Consensus MFE | -32.00 |
| Energy contribution | -32.74 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -6.00 |
| Structure conservation index | 0.65 |
| SVM decision value | 1.67 |
| SVM RNA-class probability | 0.971307 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18511499 99 - 22224390 CGGUCGUUGACCGCUGGCCUACAACCGUAGGCUAUGGCUAUAU---------------GGCUA----UAUGGCUUUAGC--UAUGGCUAUAUGGCUUUGGCUUGGUAUUAUGCCAGCUGA .((((...))))((((((.((..((((..(((((.((((((((---------------(((((----(..(((....))--))))))))))))))).)))))))))..)).))))))... ( -47.70) >DroSec_CAF1 19876 91 - 1 CGGUCGUUGACCGCUGGCCUACAACCGUAGGCUAUGGCUA---------------------------UAUGGCUUUAGC--UAUGGCUAUAUGGCUUUGGCUUGGUAAUAUGCCAGCUGA .((((...))))((((((.((..((((..(((((.(((((---------------------------(((((((.....--...)))))))))))).)))))))))..)).))))))... ( -43.60) >DroEre_CAF1 19535 114 - 1 CGGUCGUUGACCGCUGGCCUACAACCGUAGGCUAUGGCUUUAUGCUAUAUGGCUAUAUGGCUA----UAUGGCUUUAGC--UAUGGCUAUAUGGCUUUGGCUGCGUAUUAUGCCAGCUGA .((((...))))((((((((((....))))(((((((((....(((((((((((....)))))----))))))...)))--))))))(((..(((....)))..)))....))))))... ( -51.20) >DroYak_CAF1 20599 119 - 1 CGGUCGUUGACCGCUGGCCUACAACCGUAGGCUAUGGCUAUAUGGC-UUUAGCUAUAUGGCUUUAGCUAUGGCUUUAGCUAUAUGGCUAUAUGGCUUUGGCUUGGUAUUAUGCCAGCUGA .((((...))))((((((((((....))))((((.(((((...(((-(.(((((((((((((..(((....)))..)))))))))))))...)))).))))))))).....))))))... ( -55.70) >consensus CGGUCGUUGACCGCUGGCCUACAACCGUAGGCUAUGGCUAUAU_______________GGCUA____UAUGGCUUUAGC__UAUGGCUAUAUGGCUUUGGCUUGGUAUUAUGCCAGCUGA .((((...))))((((((.((..((((..(((((.(((((...........................(((((((..........)))))))))))).)))))))))..)).))))))... (-32.00 = -32.74 + 0.75)
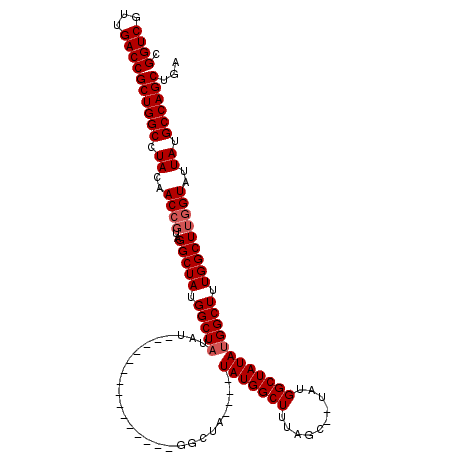


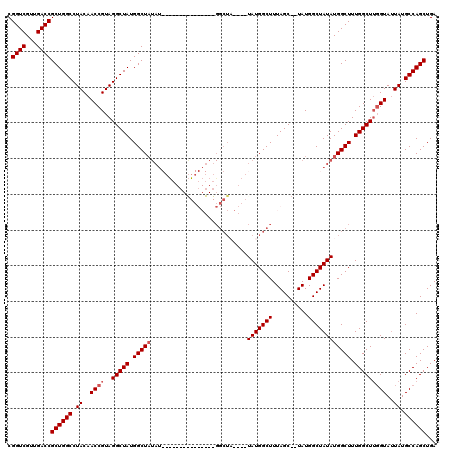
| Location | 18,511,538 – 18,511,629 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 71.57 |
| Mean single sequence MFE | -27.38 |
| Consensus MFE | -16.87 |
| Energy contribution | -17.37 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.62 |
| SVM decision value | 1.35 |
| SVM RNA-class probability | 0.944987 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18511538 91 + 22224390 -GCUAAAGCCAUA----UAGCC---------------AUAUAGCCAUAGCCUACGGUUGUAGGCCAGCGGUCAACGACCGCCAGCAGACCGACGACAAAAAGCAGCCGCAG -((((..((.(((----(....---------------)))).))..))))...(((((....((..((((((...))))))..)).)))))..........((....)).. ( -26.70) >DroPse_CAF1 22313 70 + 1 --------------------------------------UGCAGCCAUAGCCAACGGUUGUAGGCCAGCGGUCAACGACCGGCAGUCG--CAGCAACAGACA-CAGCCGCAG --------------------------------------(((.((....(((...((((((.((((...)))).))))))))).(((.--........))).-..)).))). ( -23.80) >DroSec_CAF1 19915 83 + 1 -GCUAAAGCCAUA---------------------------UAGCCAUAGCCUACGGUUGUAGGCCAGCGGUCAACGACCGCCAGCAGACCGACGACAAAAAGCAGCCGCAG -((((..((....---------------------------..))..))))...(((((....((..((((((...))))))..)).)))))..........((....)).. ( -25.20) >DroEre_CAF1 19574 106 + 1 -GCUAAAGCCAUA----UAGCCAUAUAGCCAUAUAGCAUAAAGCCAUAGCCUACGGUUGUAGGCCAGCGGUCAACGACCGCCAGCAGACCGACGACAAAAAGCAGCCGCAG -((((..((..((----(.((.((((....)))).)))))..))..))))...(((((....((..((((((...))))))..)).)))))..........((....)).. ( -28.90) >DroYak_CAF1 20639 110 + 1 AGCUAAAGCCAUAGCUAAAGCCAUAUAGCUAAA-GCCAUAUAGCCAUAGCCUACGGUUGUAGGCCAGCGGUCAACGACCGCCAGCAGACCGACGACAAAAAGCAGCCGCAG .(((.........((((..((.((((.((....-)).)))).))..))))...(((((....((..((((((...))))))..)).))))).........)))........ ( -32.70) >DroPer_CAF1 22864 70 + 1 --------------------------------------UGCUGCCAUAGCCUACGGUUGUAGGCCAGCGGUCAACGACCGGCAGUCG--CAGCAACAGACA-CAGCCGCAG --------------------------------------((((((.((.(((...((((((.((((...)))).))))))))).)).)--))))).......-......... ( -27.00) >consensus _GCUAAAGCCAUA_________________________UAUAGCCAUAGCCUACGGUUGUAGGCCAGCGGUCAACGACCGCCAGCAGACCGACGACAAAAAGCAGCCGCAG ..........................................((....((((((....))))))..((((((...))))))..........................)).. (-16.87 = -17.37 + 0.50)
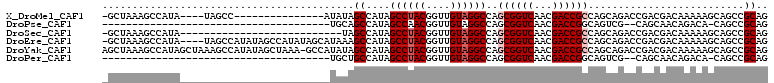
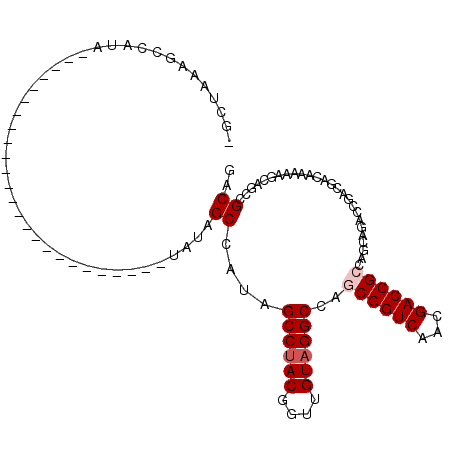
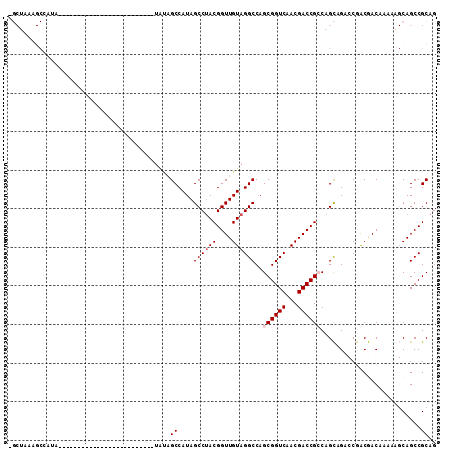
| Location | 18,511,538 – 18,511,629 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 71.57 |
| Mean single sequence MFE | -37.52 |
| Consensus MFE | -24.41 |
| Energy contribution | -24.80 |
| Covariance contribution | 0.39 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.96 |
| Structure conservation index | 0.65 |
| SVM decision value | 2.77 |
| SVM RNA-class probability | 0.996918 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 18511538 91 - 22224390 CUGCGGCUGCUUUUUGUCGUCGGUCUGCUGGCGGUCGUUGACCGCUGGCCUACAACCGUAGGCUAUGGCUAUAU---------------GGCUA----UAUGGCUUUAGC- (((((((.((.....)).)))(((..((..((((((...))))))..)).....)))))))((((.((((((((---------------....)----))))))).))))- ( -38.60) >DroPse_CAF1 22313 70 - 1 CUGCGGCUG-UGUCUGUUGCUG--CGACUGCCGGUCGUUGACCGCUGGCCUACAACCGUUGGCUAUGGCUGCA-------------------------------------- ..(((((((-((...(((...(--((((.....))))).))).((..((........))..))))))))))).-------------------------------------- ( -24.60) >DroSec_CAF1 19915 83 - 1 CUGCGGCUGCUUUUUGUCGUCGGUCUGCUGGCGGUCGUUGACCGCUGGCCUACAACCGUAGGCUAUGGCUA---------------------------UAUGGCUUUAGC- (((((((.((.....)).)))(((..((..((((((...))))))..)).....)))))))((((.((((.---------------------------...)))).))))- ( -34.40) >DroEre_CAF1 19574 106 - 1 CUGCGGCUGCUUUUUGUCGUCGGUCUGCUGGCGGUCGUUGACCGCUGGCCUACAACCGUAGGCUAUGGCUUUAUGCUAUAUGGCUAUAUGGCUA----UAUGGCUUUAGC- (((((((........)))).)))...((((((((((...)))))))(((((((....)))))))..))).....(((((((((((....)))))----))))))......- ( -44.90) >DroYak_CAF1 20639 110 - 1 CUGCGGCUGCUUUUUGUCGUCGGUCUGCUGGCGGUCGUUGACCGCUGGCCUACAACCGUAGGCUAUGGCUAUAUGGC-UUUAGCUAUAUGGCUUUAGCUAUGGCUUUAGCU ...((((.((.....)).))))((..((..((((((...))))))..))..))..((((((.(((.(((((((((((-....))))))))))).)))))))))........ ( -49.80) >DroPer_CAF1 22864 70 - 1 CUGCGGCUG-UGUCUGUUGCUG--CGACUGCCGGUCGUUGACCGCUGGCCUACAACCGUAGGCUAUGGCAGCA-------------------------------------- ..(((((..-........))))--)(.(((((((((...))))..((((((((....)))))))).)))))).-------------------------------------- ( -32.80) >consensus CUGCGGCUGCUUUUUGUCGUCGGUCUGCUGGCGGUCGUUGACCGCUGGCCUACAACCGUAGGCUAUGGCUAUA_________________________UAUGGCUUUAGC_ ..(((((........)))))......((((((((((...)))))))(((((((....)))))))..))).......................................... (-24.41 = -24.80 + 0.39)

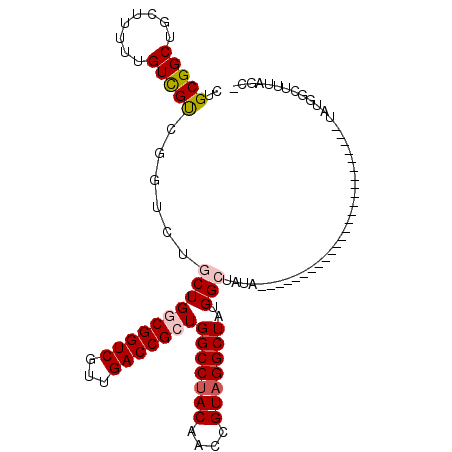
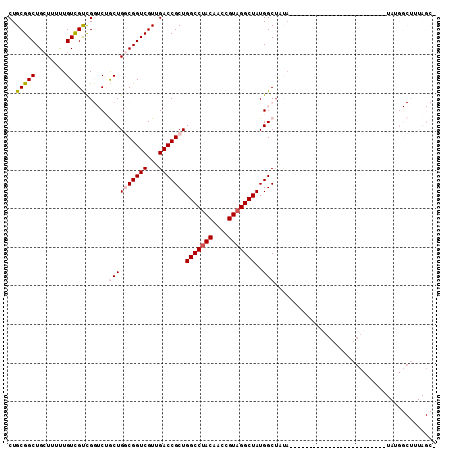
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:38:29 2006