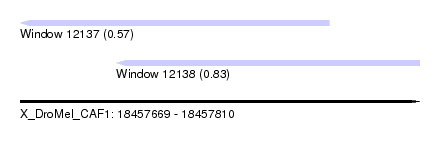

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 18,457,669 – 18,457,810 |
| Length | 141 |
| Max. P | 0.827371 |

| Location | 18,457,669 – 18,457,778 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 63.08 |
| Mean single sequence MFE | -29.64 |
| Consensus MFE | -9.34 |
| Energy contribution | -10.42 |
| Covariance contribution | 1.08 |
| Combinations/Pair | 1.32 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.32 |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.570896 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
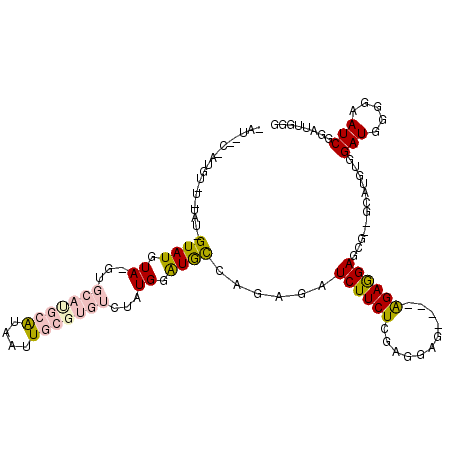
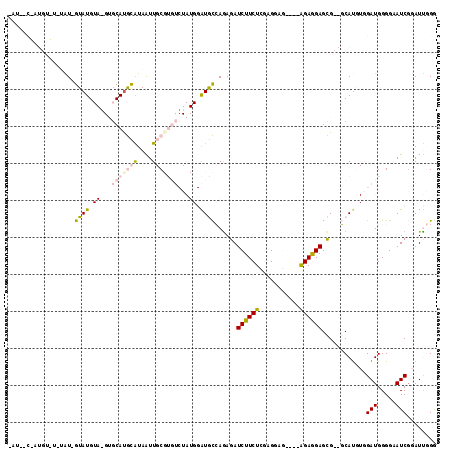
>X_DroMel_CAF1 18457669 109 - 22224390 CAUACCUAUGUAUGUAU-GUAUGUAAGUGCACGCAUAAUUGCGUGUCUAUGGAUGCCAAAGAUCUUCUCGAGGAG----AGAGGAGCG--GCAUUUGGAUGGGGAAUCGGAUUGGG ....((((..((((..(-((((....)))))..))))(((.(.((((((..((((((.....(((((((.....)----))))))..)--))))))))))).).))).....)))) ( -34.30) >DroPse_CAF1 5483 94 - 1 -----CGAUGUCUUUGUAGUAUG-----GCAUAUGUAGUU-C----------GUAUAAGAGAUCUUCUAGGGGAAAUACAGAAGAGUGCAUCAUC-CGAUGGGGCAUCCCAUCCGA -----.(((((((((...(((((-----((.......).)-)----------)))).)))).((((((.(........)))))))..))))).((-.((((((....)))))).)) ( -24.00) >DroSec_CAF1 1434 97 - 1 AAU------------AU-GUAUGUAGGUGCAUGCAUAAUUGCGUGUCUAUGGAUGCCAGAGAUCUUCUCGAGAAG----GGAGGAGCG--GCACGUGGAUGGGGAAUCGGAUUGGG ..(------------((-((((((....)))))))))(((.(.((((((((..((((.(...(((((((.....)----)))))).))--))))))))))).).)))......... ( -33.10) >DroSim_CAF1 1730 97 - 1 AAU------------AU-GUAUGUAGGUGCAUGCAUAAUUGCGUGUCUAUGGAUGCCAGAGAUCUUCUCGAGAAG----GGAGGAGCG--GCAUGUGGAUGGGGAAUCGGAUUGGG ..(------------((-((((((....)))))))))(((.(.(((((((..(((((.(...(((((((.....)----)))))).))--))))))))))).).)))......... ( -32.30) >DroPer_CAF1 6563 94 - 1 -----CGAUGUCUUUGUAGUAUG-----UCAUAUGUAGUU-C----------GUAUAAGAGAUCUUCUAGGGGAAAUACAGAAGAGUGCAUCAUG-CGAUGGGGCAUCGCAUCCGA -----(((((((((....(((((-----....(((((.((-(----------......))).((((((.(........))))))).)))))))))-)...)))))))))....... ( -24.50) >consensus _AU__C_AUGU_U_UAU_GUAUGUA_GUGCAUGCAUAAUUGCGUGUCUAUGGAUGCCAGAGAUCUUCUCGAGGAG____AGAGGAGCG__GCAUGUGGAUGGGGAAUCGGAUUGGG ..................((((.((...(((((((....)))))))...)).))))......((((((...........))))))............(((.....)))........ ( -9.34 = -10.42 + 1.08)

| Location | 18,457,703 – 18,457,810 |
|---|---|
| Length | 107 |
| Sequences | 3 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 85.05 |
| Mean single sequence MFE | -31.80 |
| Consensus MFE | -24.24 |
| Energy contribution | -24.13 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.29 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.70 |
| SVM RNA-class probability | 0.827371 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
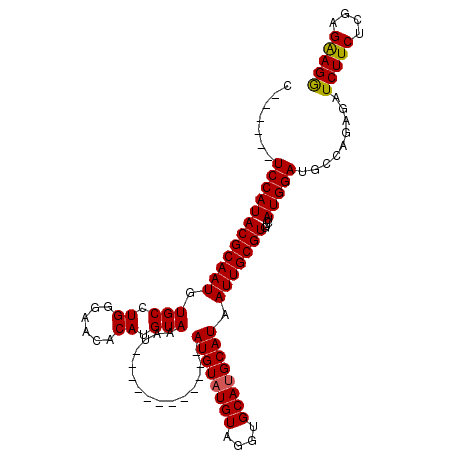
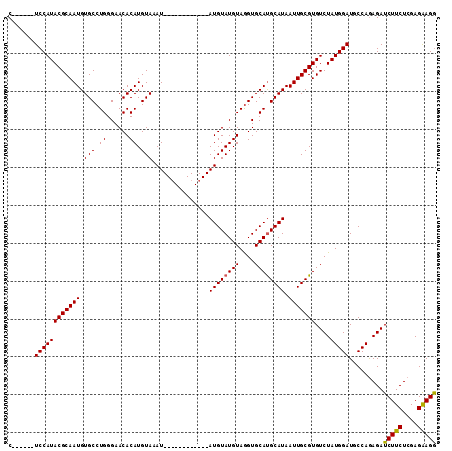
>X_DroMel_CAF1 18457703 107 - 22224390 C------UCCAUACGCAAUGUGCCUGGGAACACAUGUACAUACCUAUGUAUGUAUGUAUGUAAGUGCACGCAUAAUUGCGUGUCUAUGGAUGCCAAAGAUCUUCUCGAGGAGA (------(((.((((((((((((.((.(.(((.((((((((((....)))))))))).)))...).)).)))).))))))))(((.(((...))).))).........)))). ( -36.10) >DroSec_CAF1 1468 95 - 1 C------UCCAUACGCAAUGUGCCUGGGAACACAUGUAAAU------------AUGUAUGUAGGUGCAUGCAUAAUUGCGUGUCUAUGGAUGCCAGAGAUCUUCUCGAGAAGG .------((((((.((((((((........)))))((((.(------------((((((((....))))))))).)))).))).)))))).....(((.....)))....... ( -28.60) >DroSim_CAF1 1764 101 - 1 CUCCAUCUCCAUACGCAAUGUGCCUGGGAACACAUGUAAAU------------AUGUAUGUAGGUGCAUGCAUAAUUGCGUGUCUAUGGAUGCCAGAGAUCUUCUCGAGAAGG (((....((((((.((((((((........)))))((((.(------------((((((((....))))))))).)))).))).)))))).....(((.....)))))).... ( -30.70) >consensus C______UCCAUACGCAAUGUGCCUGGGAACACAUGUAAAU____________AUGUAUGUAGGUGCAUGCAUAAUUGCGUGUCUAUGGAUGCCAGAGAUCUUCUCGAGAAGG .......((((((((((((.(((.((......)).)))...............((((((((....)))))))).)))))))....))))).........(((((....))))) (-24.24 = -24.13 + -0.11)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:38:03 2006