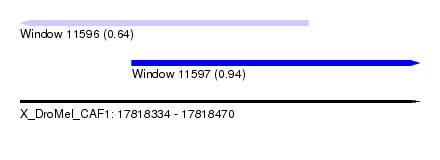
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 17,818,334 – 17,818,470 |
| Length | 136 |
| Max. P | 0.938126 |

| Location | 17,818,334 – 17,818,432 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 86.86 |
| Mean single sequence MFE | -28.45 |
| Consensus MFE | -17.81 |
| Energy contribution | -19.00 |
| Covariance contribution | 1.19 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.22 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.636410 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17818334 98 - 22224390 UAGCUUCCGCUAUAGCUAUGG--------CUUUAGCUAACCCUAAAGGCCCUGCGAUAUACCCACAACGCUUCAAAAGCGGCAGGUGCAACACACCGGUCUGUCCA ((((....))))..((...((--------((((((......))))).)))..))((((.(((.....((((.....))))...((((.....))))))).)))).. ( -27.50) >DroSec_CAF1 76194 97 - 1 UAGCUUUCGCUAUAGCCAUGU--------CUUUAGCUAACCCUAAAGGCCCUGCGAUAUCCCCACUUCGCUUCAAAAGCGGCAGGUGCAACACAC-GCUCUGUCCA ..(((((.((....))...((--------((((((......))))))))...((((..........))))....)))))((((((.((.......-)))))))).. ( -24.20) >DroEre_CAF1 78425 97 - 1 UAGCUUCCGCUAUAGCUUUGG--------CUUUAACUAACCCUAAAGGCCCUGCGAUAUCCCCACUUUGCCACAAAAGCGGCAGGUGCAACACAC-GCUCUGGCCA .((((...((....))...))--------))...............((((..(((.......(((((.(((.(....).)))))))).......)-))...)))). ( -26.44) >DroYak_CAF1 76171 105 - 1 UAGCUUCCGCCAUGGCUUUGGCUAUAUGGCUUUAGCUAACCCCAAAGGCCCAGCGAUAUCCCCACUUCGCUUCAAAAGCGGCAGGUGCAACACAC-GCUCUGGCCA (((((...((((((((....)))..)))))...)))))........((((.((((.......(((((((((.....)))))..)))).......)-)))..)))). ( -35.64) >consensus UAGCUUCCGCUAUAGCUAUGG________CUUUAGCUAACCCUAAAGGCCCUGCGAUAUCCCCACUUCGCUUCAAAAGCGGCAGGUGCAACACAC_GCUCUGGCCA ........((((.(((.............((((((......))))))((((((((.......)....((((.....))))))))).))........))).)))).. (-17.81 = -19.00 + 1.19)

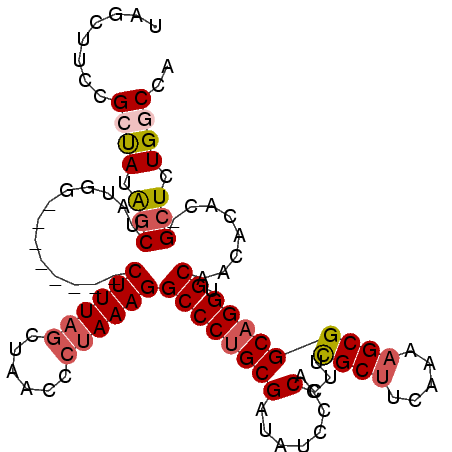
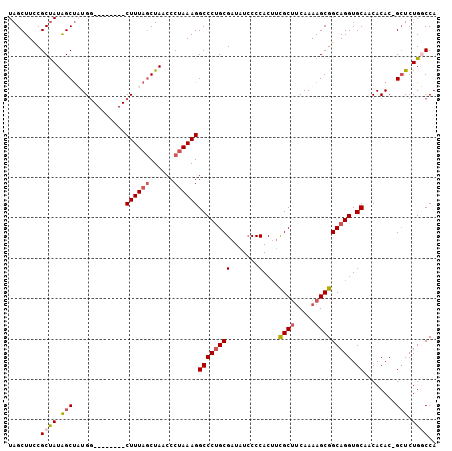
| Location | 17,818,372 – 17,818,470 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 87.15 |
| Mean single sequence MFE | -31.05 |
| Consensus MFE | -22.55 |
| Energy contribution | -22.43 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.61 |
| Structure conservation index | 0.73 |
| SVM decision value | 1.27 |
| SVM RNA-class probability | 0.938126 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 17818372 98 + 22224390 GUUGUGGGUAUAUCGCAGGGCCUUUAGGGUUAGCUAAAG--------CCAUAGCUAUAGCGGAAGCUAAAGCACAACACGAUUUUUAUAGAC-CCACAUAGCGAAGU (((((((((..((((...(((.(((((......))))))--------))...(((.((((....)))).)))......))))........))-)))))..))..... ( -32.70) >DroSec_CAF1 76231 98 + 1 GAAGUGGGGAUAUCGCAGGGCCUUUAGGGUUAGCUAAAG--------ACAUGGCUAUAGCGAAAGCUAAAGCACAACACGAUUUUUAUAGAC-CCACACAGCGAAGU ...(((((..(((.((..(.((....)).)..)).....--------...(((((.((((....)))).))).))...........)))..)-)))).......... ( -29.00) >DroEre_CAF1 78462 99 + 1 AAAGUGGGGAUAUCGCAGGGCCUUUAGGGUUAGUUAAAG--------CCAAAGCUAUAGCGGAAGCUAAAGCACAACUCGAUUUUUAUAGACCCCACACAGCGAAAU ...((((((..((((.(((((.(((((......))))))--------))...(((.((((....)))).)))....)))))).........)))))).......... ( -31.70) >DroYak_CAF1 76208 106 + 1 GAAGUGGGGAUAUCGCUGGGCCUUUGGGGUUAGCUAAAGCCAUAUAGCCAAAGCCAUGGCGGAAGCUAAAGCACAACACGAUUUUUAUAGAC-CCACACAGCAAAGU ...(((((..(((.(((((((.(((((......))))))))...))))....((..((((....))))..))..............)))..)-)))).......... ( -30.80) >consensus GAAGUGGGGAUAUCGCAGGGCCUUUAGGGUUAGCUAAAG________CCAAAGCUAUAGCGGAAGCUAAAGCACAACACGAUUUUUAUAGAC_CCACACAGCGAAGU ...(((((..(((.((.((.((....)).)).))..................(((.((((....)))).)))..............)))..).)))).......... (-22.55 = -22.43 + -0.12)

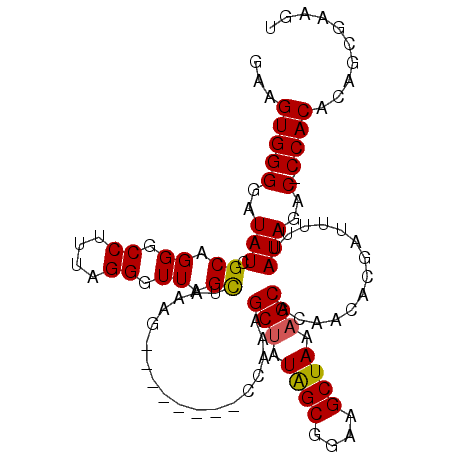
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:30:11 2006