
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 17,766,504 – 17,766,594 |
| Length | 90 |
| Max. P | 0.830965 |

| Location | 17,766,504 – 17,766,594 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 83.13 |
| Mean single sequence MFE | -19.20 |
| Consensus MFE | -13.00 |
| Energy contribution | -13.24 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.553980 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
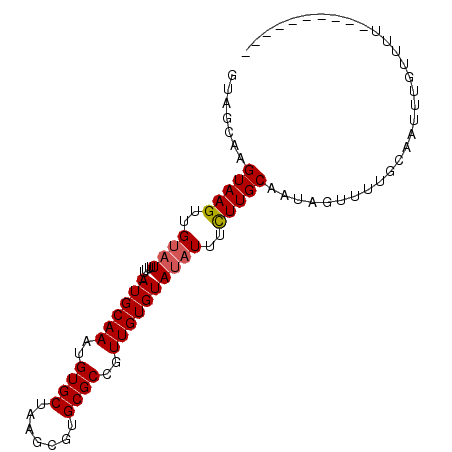
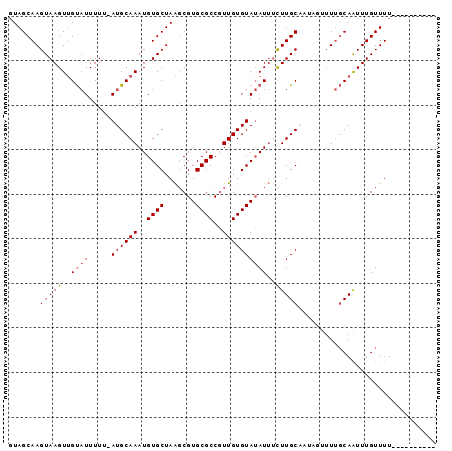
>X_DroMel_CAF1 17766504 90 + 22224390 GUAGCAAGUAAGUUGUAUUUUGAAUGCAAAUGUGCUAAGCGUGCGCCGUUGUGUAUAUUUCUUGCAAAAGUUUUGCAAUUUGUUUUGUUU------ ..((((((..(((((((.......(((((((((((..((((.....))))..))))))...))))).......)))))))...)))))).------ ( -18.74) >DroSec_CAF1 25444 84 + 1 UUAGCAAGUAAGUAGC-UUUUU-AUGCAAAUGUGCUAGCCGUGCGCCUUUGUGUAGAUACUUUGCAAUAGUAUCUCAAUUUGUUUU---------- ..(((((((.......-....(-(((((((.((((.......)))).))))))))((((((.......))))))...)))))))..---------- ( -20.80) >DroSim_CAF1 26399 85 + 1 GUAGCAAGUAAGUUGUAUUUUU-AUGCAAAUGUGCUAAGCGUGCGCCGUUGUGUAUAUUUCUUGCAAAAGUUUUGCAAUUUGUUUU---------- ...(((((....(((((....(-(((((...((((.......)))).....)))))).....)))))....)))))..........---------- ( -19.10) >DroEre_CAF1 23900 94 + 1 GUAGCAAGUAAGUUGUAUUUUU-AUGCAAAUGUGCUAAGCGUGCGCCGUUGUGUAUAUUUCUUGCUUU-UUUUUGCAAUUUGUUUUGUUUUUUUGG ...(((((.((((........(-((((((..((((.......))))..)))))))........)))).-..))))).................... ( -18.19) >DroYak_CAF1 24156 83 + 1 GUAGCAAGUAAGUUGUAUUUUU-AUGCAAAUGUGCUAAGCGUGCGCCGUUGUGUAUAUUUCUUGCUUU---UUUGCAGUUUGUUUUU--------- ...(((((.((((........(-((((((..((((.......))))..)))))))........)))).---)))))...........--------- ( -19.19) >consensus GUAGCAAGUAAGUUGUAUUUUU_AUGCAAAUGUGCUAAGCGUGCGCCGUUGUGUAUAUUUCUUGCAAUAGUUUUGCAAUUUGUUUU__________ .......(((((..((((.....((((((..((((.......))))..))))))))))..)))))............................... (-13.00 = -13.24 + 0.24)

| Location | 17,766,504 – 17,766,594 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 83.13 |
| Mean single sequence MFE | -11.42 |
| Consensus MFE | -8.70 |
| Energy contribution | -8.94 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.830965 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
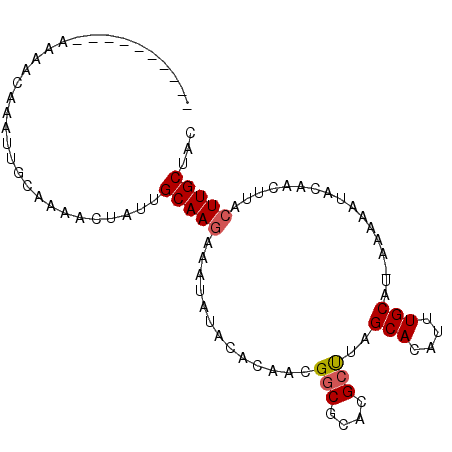
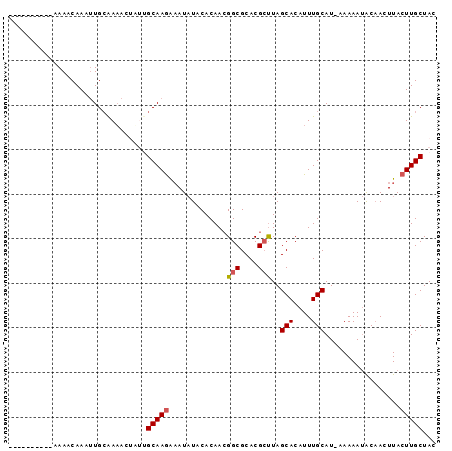
>X_DroMel_CAF1 17766504 90 - 22224390 ------AAACAAAACAAAUUGCAAAACUUUUGCAAGAAAUAUACACAACGGCGCACGCUUAGCACAUUUGCAUUCAAAAUACAACUUACUUGCUAC ------.........................(((((.............(((....)))..(((....))).................)))))... ( -10.50) >DroSec_CAF1 25444 84 - 1 ----------AAAACAAAUUGAGAUACUAUUGCAAAGUAUCUACACAAAGGCGCACGGCUAGCACAUUUGCAU-AAAAA-GCUACUUACUUGCUAA ----------.........(((((((((.......))))))).))....(((.....))).(((....)))..-....(-((.........))).. ( -12.30) >DroSim_CAF1 26399 85 - 1 ----------AAAACAAAUUGCAAAACUUUUGCAAGAAAUAUACACAACGGCGCACGCUUAGCACAUUUGCAU-AAAAAUACAACUUACUUGCUAC ----------.....................(((((.............(((....)))..(((....)))..-..............)))))... ( -10.50) >DroEre_CAF1 23900 94 - 1 CCAAAAAAACAAAACAAAUUGCAAAAA-AAAGCAAGAAAUAUACACAACGGCGCACGCUUAGCACAUUUGCAU-AAAAAUACAACUUACUUGCUAC ...........................-..((((((.............(((....)))..(((....)))..-..............)))))).. ( -11.90) >DroYak_CAF1 24156 83 - 1 ---------AAAAACAAACUGCAAA---AAAGCAAGAAAUAUACACAACGGCGCACGCUUAGCACAUUUGCAU-AAAAAUACAACUUACUUGCUAC ---------................---..((((((.............(((....)))..(((....)))..-..............)))))).. ( -11.90) >consensus __________AAAACAAAUUGCAAAACUAUUGCAAGAAAUAUACACAACGGCGCACGCUUAGCACAUUUGCAU_AAAAAUACAACUUACUUGCUAC ...............................(((((.............(((....)))..(((....))).................)))))... ( -8.70 = -8.94 + 0.24)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:29:42 2006