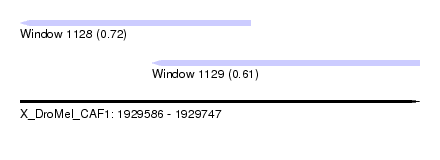
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,929,586 – 1,929,747 |
| Length | 161 |
| Max. P | 0.716792 |

| Location | 1,929,586 – 1,929,679 |
|---|---|
| Length | 93 |
| Sequences | 3 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 97.85 |
| Mean single sequence MFE | -25.59 |
| Consensus MFE | -24.39 |
| Energy contribution | -24.72 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.716792 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
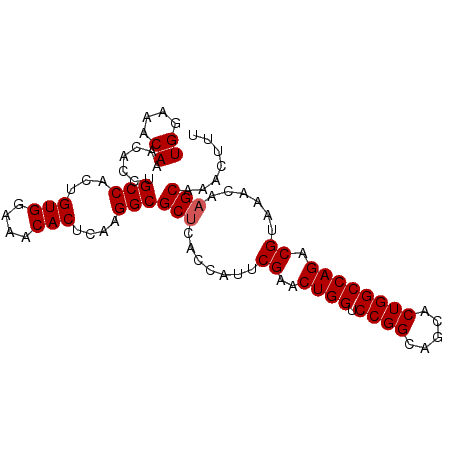
>X_DroMel_CAF1 1929586 93 - 22224390 UGGAAACAAACACCUGCCACUGUGGAACCACUCAAGGCGCGCACCAUUCGAACUGGUCCGGCAGCACUGGCCAGACGUAAACAAGCGAACUUU ((....)).......(((...(((....)))....))).(((......((..((((.((((.....)))))))).)).......)))...... ( -24.12) >DroSec_CAF1 989 93 - 1 UGGAAACAAACACCUGCCACUGUGGAAACACUCAAGGCGCUCACCAUUCGAACUGGUCCGGCAGCACUGGCCAGACGUAAACAAGCAAACUUU ((....)).......(((...(((....)))....)))(((.......((..((((.((((.....)))))))).))......)))....... ( -26.32) >DroSim_CAF1 1257 93 - 1 UGGAAACAAACACCUGCCACUGUGGAAACACUCAAGGCGCUCACCAUUCGAACUGGUCCGGCAGCACUGGCCAGACGUAAACAAGCAAACUUU ((....)).......(((...(((....)))....)))(((.......((..((((.((((.....)))))))).))......)))....... ( -26.32) >consensus UGGAAACAAACACCUGCCACUGUGGAAACACUCAAGGCGCUCACCAUUCGAACUGGUCCGGCAGCACUGGCCAGACGUAAACAAGCAAACUUU ((....)).......(((...(((....)))....)))(((.......((..((((.((((.....)))))))).))......)))....... (-24.39 = -24.72 + 0.33)

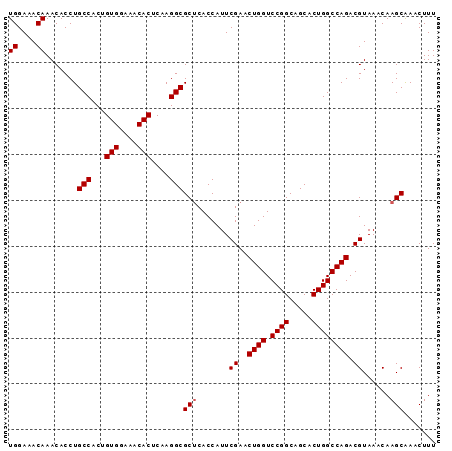
| Location | 1,929,639 – 1,929,747 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.29 |
| Mean single sequence MFE | -37.80 |
| Consensus MFE | -25.20 |
| Energy contribution | -25.82 |
| Covariance contribution | 0.62 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.605481 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
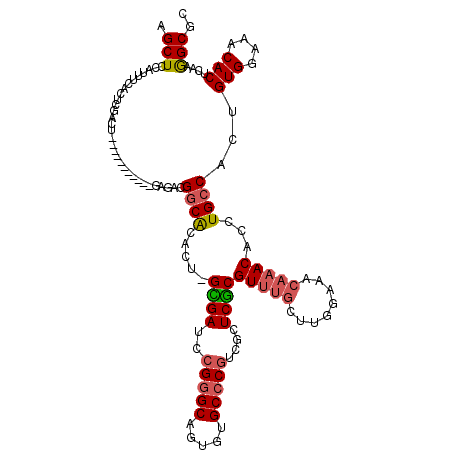
>X_DroMel_CAF1 1929639 108 - 22224390 AGCUCCAUUUCACUCGAUU-----------GGGACGGCACACU-GUGAUCCGGGCAGUGUGCCCGACGCUCGCGUUUGCUUGGAAACAAACACCUGCCACUGUGGAACCACUCAAGGCGC .(..(((..((....)).)-----------))..)((((((((-((.......)))))))))).((((....))))((.(((....))).))...(((...(((....)))....))).. ( -36.50) >DroSec_CAF1 1042 108 - 1 AGCUUCAUUUUACUCUACU-----------GAGACGGCACACU-GCGAUCCGGGCAGUGUGCCCGUCGCUCGCGUUUGCUUGGAAACAAACACCUGCCACUGUGGAAACACUCAAGGCGC (((..........((....-----------))(((((((((((-((.......)))))))).)))))))).((...((.(((....))).))...(((...(((....)))....))))) ( -40.50) >DroSim_CAF1 1310 108 - 1 AGCUUCAUUUUACUCGACU-----------GAGACGGCACACU-GCGAUCCGGGCAGUGUGCCCGUCGCUCGCGUUUGCUUGGAAACAAACACCUGCCACUGUGGAAACACUCAAGGCGC .((........((.(((..-----------..(((((((((((-((.......)))))))).)))))..))).)).((.(((....))).))...(((...(((....)))....))))) ( -40.60) >DroEre_CAF1 1142 114 - 1 AGCUCCAUUUCACUCUAUUGGAUUCCAAUUGGAACGCCGCACCAGGGAUACGAGCAGUGUGCCCAUC--UCCCGUUUGCCAGGGA----ACACCUGCCACUGUGGUCCCACUCAAAGCGC .(((...............((.((((....))))..))......(((((....((((((.((.....--((((........))))----......)))))))).)))))......))).. ( -33.60) >consensus AGCUCCAUUUCACUCGACU___________GAGACGGCACACU_GCGAUCCGGGCAGUGUGCCCGUCGCUCGCGUUUGCUUGGAAACAAACACCUGCCACUGUGGAAACACUCAAGGCGC .(((...............................((((.....((((..(((((.....)))))....))))(((((........)))))...))))...(((....)))....))).. (-25.20 = -25.82 + 0.62)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:51:34 2006