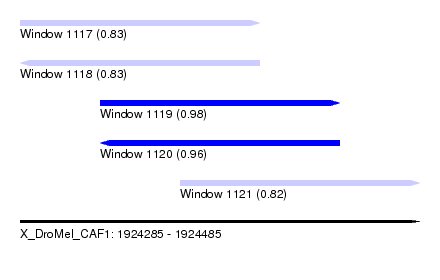

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,924,285 – 1,924,485 |
| Length | 200 |
| Max. P | 0.978091 |

| Location | 1,924,285 – 1,924,405 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.33 |
| Mean single sequence MFE | -45.50 |
| Consensus MFE | -37.26 |
| Energy contribution | -38.08 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.830149 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1924285 120 + 22224390 CGUGCCAAGGACAGCGGACACGACCCAAACCCAAAGCUCCAGUCUGGAGCGGAGCUCCGUAAUGGGUGCACUUAGCACCCACUGCUAGCAGACGAUCGUUCUGCUUCACGAGCAGCACGC (((((...((....((....))..)).........((((.....(((((((((((..(((..(((((((.....)))))))(((....))))))...))))))))))).)))).))))). ( -47.40) >DroSec_CAF1 8255 120 + 1 CGUGCCAAGGACAGCGGACACGACCCAAGCCCAAAGCUCCAGUCUUGAGCGGAGCUCCGUAAUGGGUGCACUUGGCACCCACUGCUAGCAGACGAUCGUUCUGCUUCACAAGCAGAACGC .((((((((..(..((....))(((((.......((((((.((.....))))))))......))))))..))))))))...(((....))).....(((((((((.....))))))))). ( -46.82) >DroEre_CAF1 8323 120 + 1 CGUGCCAAGGACAGCGGACACGACCCAAACCCAAGGCUCCAGUCUGGAGCGGAGCUCCGUAAUGGGUGCACUUCGCACCCACGGCAAGCAGACGAUCGUUCUGCUUUACAAGCAGCACGC (((((.......((((((.((((............(((((.....)))))...(((((((...((((((.....))))))))))..)))......)))))))))).........))))). ( -44.59) >DroYak_CAF1 8359 120 + 1 CGUGCCACGGACAGCGAACACGACCCAAACCCAAGGCUCCAGUCUGAAGCGGAGCUCCGUAGUGGGUGCACUUGGCACCCACGGCCAGCAGACGAUCAUUCUGCUUCACAAGCAGCACGC (((((.(((((...((....)).............(((((.((.....)))))))))))).((((((((.....))))))))(...((((((.......))))))...).....))))). ( -43.20) >consensus CGUGCCAAGGACAGCGGACACGACCCAAACCCAAAGCUCCAGUCUGGAGCGGAGCUCCGUAAUGGGUGCACUUGGCACCCACGGCUAGCAGACGAUCGUUCUGCUUCACAAGCAGCACGC (((((....((.((((((.((((...........((((((.((.....))))))))..(((.(((((((.....))))))).)))..........)))))))))))).......))))). (-37.26 = -38.08 + 0.81)

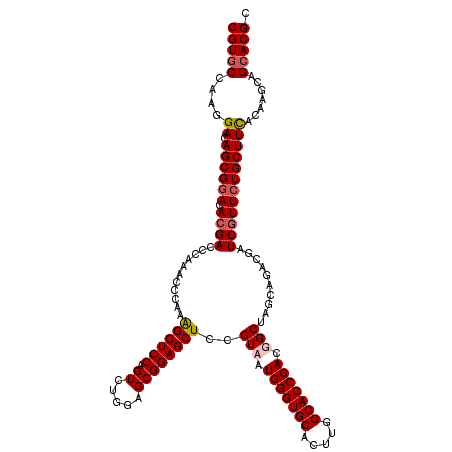
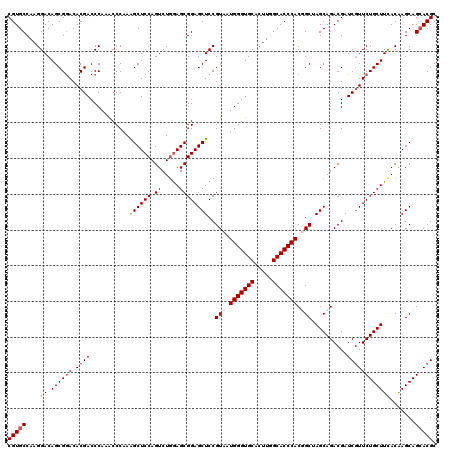
| Location | 1,924,285 – 1,924,405 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.33 |
| Mean single sequence MFE | -52.27 |
| Consensus MFE | -46.97 |
| Energy contribution | -46.60 |
| Covariance contribution | -0.37 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.70 |
| SVM RNA-class probability | 0.827484 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1924285 120 - 22224390 GCGUGCUGCUCGUGAAGCAGAACGAUCGUCUGCUAGCAGUGGGUGCUAAGUGCACCCAUUACGGAGCUCCGCUCCAGACUGGAGCUUUGGGUUUGGGUCGUGUCCGCUGUCCUUGGCACG .((((((........((((((.(....)))))))((((((((((((.....)))))))))(((((((...))))).((((.((((.....)))).))))))....)))......)))))) ( -51.30) >DroSec_CAF1 8255 120 - 1 GCGUUCUGCUUGUGAAGCAGAACGAUCGUCUGCUAGCAGUGGGUGCCAAGUGCACCCAUUACGGAGCUCCGCUCAAGACUGGAGCUUUGGGCUUGGGUCGUGUCCGCUGUCCUUGGCACG .(((((((((.....))))))))).(((.(((....))))))((((((((..((((((...((((((((((........))))))))))....))))).((....)).)..)))))))). ( -54.90) >DroEre_CAF1 8323 120 - 1 GCGUGCUGCUUGUAAAGCAGAACGAUCGUCUGCUUGCCGUGGGUGCGAAGUGCACCCAUUACGGAGCUCCGCUCCAGACUGGAGCCUUGGGUUUGGGUCGUGUCCGCUGUCCUUGGCACG .((((((.......(((((((.(....))))))))((.((((((((.....))))))))((((((((...))))).((((.(((((...))))).)))))))...)).......)))))) ( -50.20) >DroYak_CAF1 8359 120 - 1 GCGUGCUGCUUGUGAAGCAGAAUGAUCGUCUGCUGGCCGUGGGUGCCAAGUGCACCCACUACGGAGCUCCGCUUCAGACUGGAGCCUUGGGUUUGGGUCGUGUUCGCUGUCCGUGGCACG .(((((..(..((((((((((.......))))))((((((((((((.....))))))))....((((((.(((((.....)))))...)))))).))))....)))).....)..))))) ( -52.70) >consensus GCGUGCUGCUUGUGAAGCAGAACGAUCGUCUGCUAGCAGUGGGUGCCAAGUGCACCCAUUACGGAGCUCCGCUCCAGACUGGAGCCUUGGGUUUGGGUCGUGUCCGCUGUCCUUGGCACG .((((((........((((((.(....)))))))(((.((((((((.....))))))))((((((((...))))).((((.(((((...))))).)))))))...)))......)))))) (-46.97 = -46.60 + -0.37)

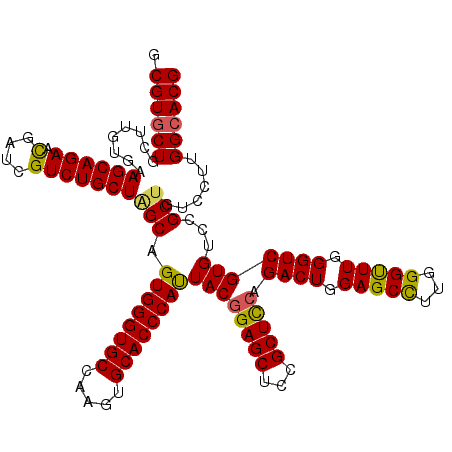
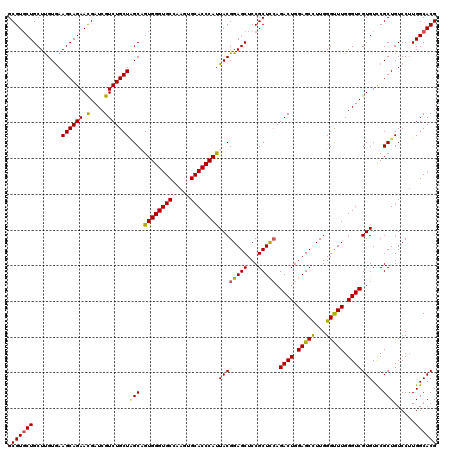
| Location | 1,924,325 – 1,924,445 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.75 |
| Mean single sequence MFE | -48.78 |
| Consensus MFE | -42.09 |
| Energy contribution | -42.40 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.06 |
| Mean z-score | -3.34 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.81 |
| SVM RNA-class probability | 0.978091 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1924325 120 + 22224390 AGUCUGGAGCGGAGCUCCGUAAUGGGUGCACUUAGCACCCACUGCUAGCAGACGAUCGUUCUGCUUCACGAGCAGCACGCGGGUGUCCUCAUCGAAGUCCACCUGCUUCAUUUCGUUCUC .....((((((((((((.(((.(((((((.....))))))).))).(((((((....).))))))....)))).....(((((((..((......))..))))))).....)))))))). ( -51.00) >DroSec_CAF1 8295 120 + 1 AGUCUUGAGCGGAGCUCCGUAAUGGGUGCACUUGGCACCCACUGCUAGCAGACGAUCGUUCUGCUUCACAAGCAGAACGCGGGUGUCCUCAUCGAAGUCCACCUGCUUCAUCUCGUUCUC ......(((((..(((..(((.(((((((.....))))))).))).)))(((.((..((((((((.....))))))))(((((((..((......))..))))))).)).)))))))).. ( -51.50) >DroEre_CAF1 8363 120 + 1 AGUCUGGAGCGGAGCUCCGUAAUGGGUGCACUUCGCACCCACGGCAAGCAGACGAUCGUUCUGCUUUACAAGCAGCACGCGGUUGUCCUCAUCGAAGUCCACUUGCUUCAUCUCGUUCUC .....(((((((((((((((...((((((.....))))))))))..))).(((((((((.(((((.....)))))...)))))))))......(((((......))))).)).)))))). ( -47.40) >DroYak_CAF1 8399 120 + 1 AGUCUGAAGCGGAGCUCCGUAGUGGGUGCACUUGGCACCCACGGCCAGCAGACGAUCAUUCUGCUUCACAAGCAGCACGCGGGUGUCCUCAUCGAAGUCCACUUGCUUCAUCUCGUUCUC .(((((..((((....)))).((((((((.....))))))))......)))))(((....(((((.....)))))...(((((((..((......))..)))))))...)))........ ( -45.20) >consensus AGUCUGGAGCGGAGCUCCGUAAUGGGUGCACUUGGCACCCACGGCUAGCAGACGAUCGUUCUGCUUCACAAGCAGCACGCGGGUGUCCUCAUCGAAGUCCACCUGCUUCAUCUCGUUCUC .(((((..((((....))))..(((((((.....))))))).......)))))((.(((.((((((...)))))).))(((((((..((......))..)))))))........).)).. (-42.09 = -42.40 + 0.31)

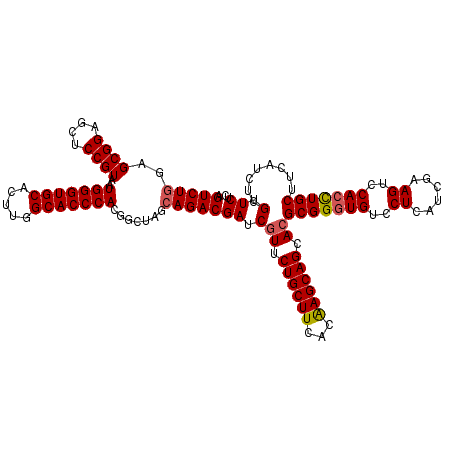
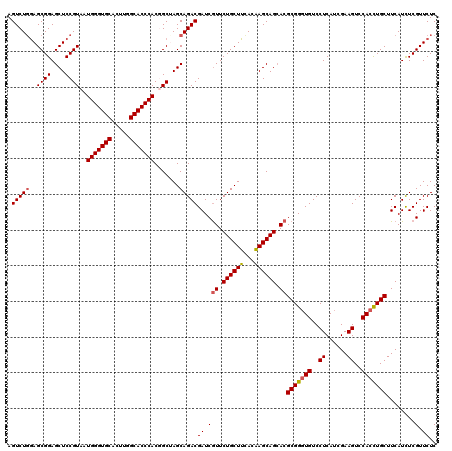
| Location | 1,924,325 – 1,924,445 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.75 |
| Mean single sequence MFE | -48.25 |
| Consensus MFE | -41.83 |
| Energy contribution | -41.70 |
| Covariance contribution | -0.13 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.45 |
| SVM RNA-class probability | 0.955328 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1924325 120 - 22224390 GAGAACGAAAUGAAGCAGGUGGACUUCGAUGAGGACACCCGCGUGCUGCUCGUGAAGCAGAACGAUCGUCUGCUAGCAGUGGGUGCUAAGUGCACCCAUUACGGAGCUCCGCUCCAGACU ..((.((.......((.((((..(((....)))..)))).))...(((((.....)))))..)).))(((((..((((((((((((.....)))))))))..((....))))).))))). ( -48.40) >DroSec_CAF1 8295 120 - 1 GAGAACGAGAUGAAGCAGGUGGACUUCGAUGAGGACACCCGCGUUCUGCUUGUGAAGCAGAACGAUCGUCUGCUAGCAGUGGGUGCCAAGUGCACCCAUUACGGAGCUCCGCUCAAGACU (((....((((((.((.((((..(((....)))..)))).))((((((((.....))))))))..))))))(((.(.(((((((((.....))))))))).)..)))....)))...... ( -52.00) >DroEre_CAF1 8363 120 - 1 GAGAACGAGAUGAAGCAAGUGGACUUCGAUGAGGACAACCGCGUGCUGCUUGUAAAGCAGAACGAUCGUCUGCUUGCCGUGGGUGCGAAGUGCACCCAUUACGGAGCUCCGCUCCAGACU ..............(((((..(((.((((((.((....)).))).(((((.....)))))..)))..)))..))))).((((((((.....))))))))...(((((...)))))..... ( -46.50) >DroYak_CAF1 8399 120 - 1 GAGAACGAGAUGAAGCAAGUGGACUUCGAUGAGGACACCCGCGUGCUGCUUGUGAAGCAGAAUGAUCGUCUGCUGGCCGUGGGUGCCAAGUGCACCCACUACGGAGCUCCGCUUCAGACU ..........((((((....((((((((....((.((((((((.(((........((((((.......))))))))))))))))))).((((....)))).))))).))))))))).... ( -46.10) >consensus GAGAACGAGAUGAAGCAAGUGGACUUCGAUGAGGACACCCGCGUGCUGCUUGUGAAGCAGAACGAUCGUCUGCUAGCAGUGGGUGCCAAGUGCACCCAUUACGGAGCUCCGCUCCAGACU .................(((((((((((((((((....)).(((.(((((.....))))).))).))))).....(.(((((((((.....))))))))).))))).))))))....... (-41.83 = -41.70 + -0.13)

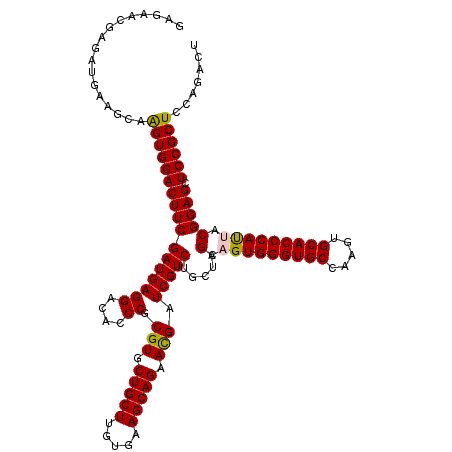
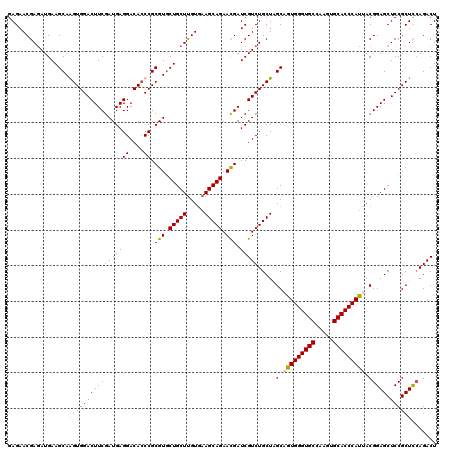
| Location | 1,924,365 – 1,924,485 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.06 |
| Mean single sequence MFE | -36.35 |
| Consensus MFE | -31.98 |
| Energy contribution | -32.10 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.824435 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1924365 120 + 22224390 ACUGCUAGCAGACGAUCGUUCUGCUUCACGAGCAGCACGCGGGUGUCCUCAUCGAAGUCCACCUGCUUCAUUUCGUUCUCCUUCAGAUCCGUAACUCUGCAGUCCACGGGGACCGCACUU .(((..((.((((((..((.((((((...)))))).))(((((((..((......))..)))))))......))).))).)).)))...........(((.((((....)))).)))... ( -38.80) >DroSec_CAF1 8335 120 + 1 ACUGCUAGCAGACGAUCGUUCUGCUUCACAAGCAGAACGCGGGUGUCCUCAUCGAAGUCCACCUGCUUCAUCUCGUUCUCCUUCAGAUCCGUAACCUUGCAGUCCACAGGGACCGCACUG .(((..((.((((((..((((((((.....))))))))(((((((..((......))..)))))))......))).))).)).)))...........(((.((((....)))).)))... ( -42.80) >DroEre_CAF1 8403 120 + 1 ACGGCAAGCAGACGAUCGUUCUGCUUUACAAGCAGCACGCGGUUGUCCUCAUCGAAGUCCACUUGCUUCAUCUCGUUCUCCUUCAGAUCCGUAACUCUGCAGUCCACGAGGACCGCACUG ((((......(((((((((.(((((.....)))))...)))))))))......(((((......)))))...................)))).....(((.((((....)))).)))... ( -34.60) >DroYak_CAF1 8439 120 + 1 ACGGCCAGCAGACGAUCAUUCUGCUUCACAAGCAGCACGCGGGUGUCCUCAUCGAAGUCCACUUGCUUCAUCUCGUUCUCCUUCAGAUCCGUAACUCUGCAGUCCACGAGCACCGCACUG .(((..((((((.......)))))).........((..(((((((..((......))..)))))))......((((....((.((((........)))).))...)))))).)))..... ( -29.20) >consensus ACGGCUAGCAGACGAUCGUUCUGCUUCACAAGCAGCACGCGGGUGUCCUCAUCGAAGUCCACCUGCUUCAUCUCGUUCUCCUUCAGAUCCGUAACUCUGCAGUCCACGAGGACCGCACUG ...((..((((((((((((.((((((...)))))).))(((((((..((......))..)))))))...................))).)))....)))).((((....)))).)).... (-31.98 = -32.10 + 0.12)

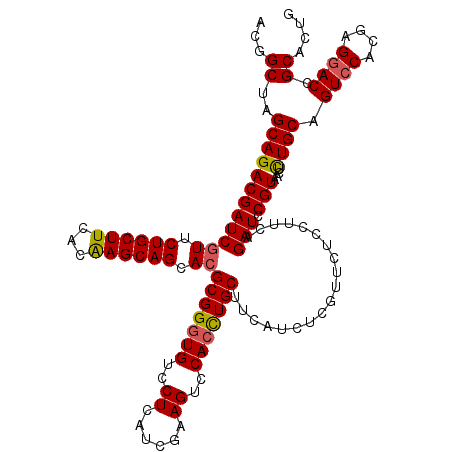
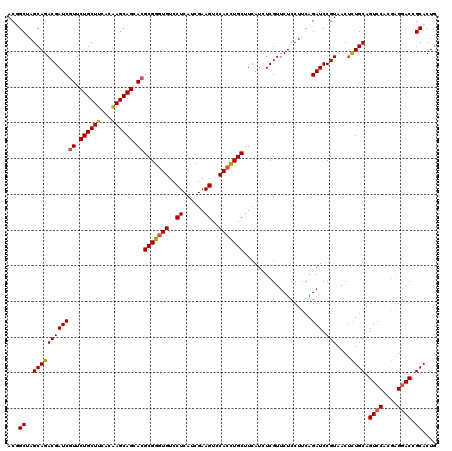
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:51:27 2006