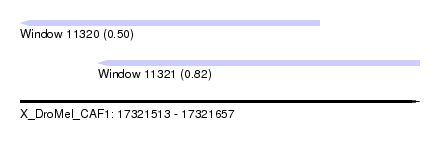

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 17,321,513 – 17,321,657 |
| Length | 144 |
| Max. P | 0.823717 |

| Location | 17,321,513 – 17,321,621 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 84.92 |
| Mean single sequence MFE | -13.65 |
| Consensus MFE | -11.22 |
| Energy contribution | -11.00 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.82 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
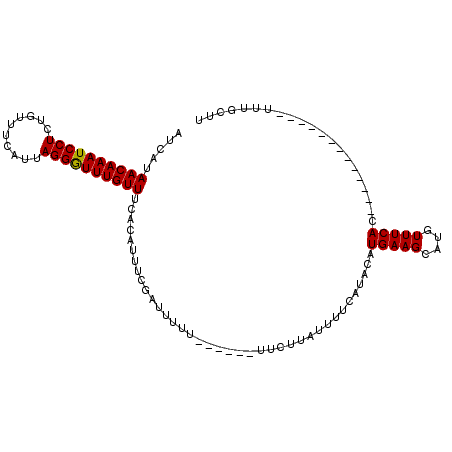
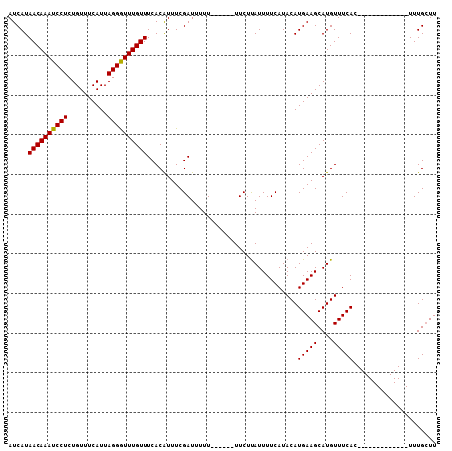
>X_DroMel_CAF1 17321513 108 - 22224390 AUCAUAACAAAUCCUCUGUUUCAUUAGGAUUUGUUUCACGUUUCGAUUUUUUUUUGUUUCUCAUUUUCAUACAUGAAGCAUGUUUCACUCACUGGCUGGCCUUUGGUU .....((((((((((..........))))))))))...................((((((..............))))))........(((..((....))..))).. ( -16.14) >DroSec_CAF1 38581 89 - 1 AUCAUAACAAAUCCUCUGUUUCAUUAGGGUUUGUUUCACAUUUCGAUUUUU------UUCUUAUUUUCAUACAUGAAGCAUGUUUCAC-------------UUUGCUU .((((((((((((((..........)))))))))).........((.....------.))............))))((((........-------------..)))). ( -12.40) >DroSim_CAF1 28473 89 - 1 AUCAUAACAAAUCCUCUGUUUCAUUAGGGUUUGUUUCACAUUUCGAUUUUU------UUCUUAUUUUCAUACAUGAAGCAUGUUUCAC-------------UUUGCUU .((((((((((((((..........)))))))))).........((.....------.))............))))((((........-------------..)))). ( -12.40) >consensus AUCAUAACAAAUCCUCUGUUUCAUUAGGGUUUGUUUCACAUUUCGAUUUUU______UUCUUAUUUUCAUACAUGAAGCAUGUUUCAC_____________UUUGCUU .....((((((((((..........))))))))))......................................(((((....)))))..................... (-11.22 = -11.00 + -0.22)

| Location | 17,321,541 – 17,321,657 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 90.52 |
| Mean single sequence MFE | -20.70 |
| Consensus MFE | -16.69 |
| Energy contribution | -16.80 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.823717 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
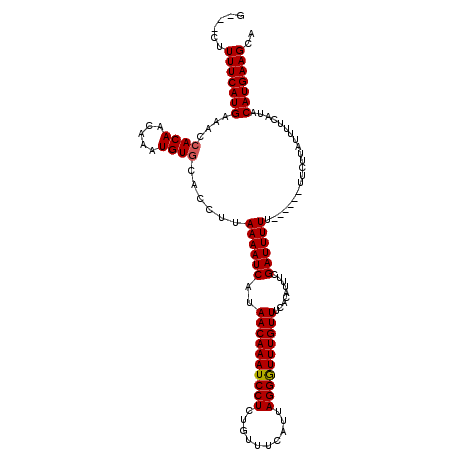
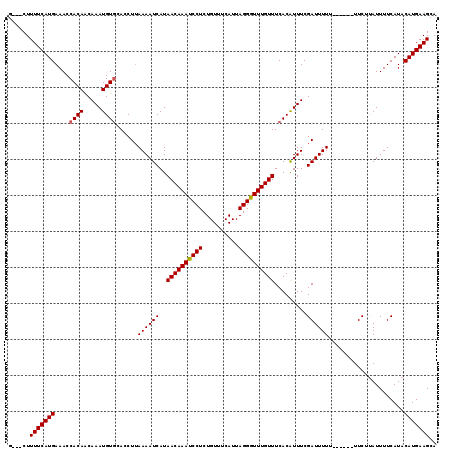
>X_DroMel_CAF1 17321541 116 - 22224390 G---CUUUUCAUGAAACCACAACAAAUGUGCAACUAAAAAUCAUAACAAAUCCUCUGUUUCAUUAGGAUUUGUUUCACGUUUCGAUUUUUUUUUGUUUCUCAUUUUCAUACAUGAAGCA (---((((..((((((.((((.....))))(((..(((((((..((((((((((..........)))))))))).........)))))))..)))........))))))....))))). ( -23.60) >DroSec_CAF1 38596 113 - 1 GGUGCUUUUCAUGAAACCACAUCAAAUGUCCACCUUAAAAUCAUAACAAAUCCUCUGUUUCAUUAGGGUUUGUUUCACAUUUCGAUUUUU------UUCUUAUUUUCAUACAUGAAGCA ..((((((..((((((.......((((((...............((((((((((..........))))))))))..)))))).((.....------.))....))))))....)))))) ( -17.53) >DroSim_CAF1 28488 110 - 1 G---CUUUUCAUGAAACCACAGCAAAUGUGCACCUUAAAAUCAUAACAAAUCCUCUGUUUCAUUAGGGUUUGUUUCACAUUUCGAUUUUU------UUCUUAUUUUCAUACAUGAAGCA (---((((..((((((.....(.(((((((..............((((((((((..........)))))))))).))))))))((.....------.))....))))))....))))). ( -20.96) >consensus G___CUUUUCAUGAAACCACAACAAAUGUGCACCUUAAAAUCAUAACAAAUCCUCUGUUUCAUUAGGGUUUGUUUCACAUUUCGAUUUUU______UUCUUAUUUUCAUACAUGAAGCA ......(((((((....((((.....))))......((((((..((((((((((..........)))))))))).........)))))).....................))))))).. (-16.69 = -16.80 + 0.11)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:26:01 2006