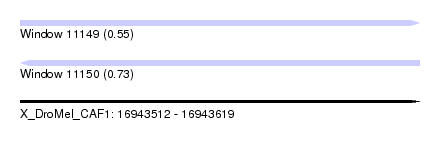

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 16,943,512 – 16,943,619 |
| Length | 107 |
| Max. P | 0.731371 |

| Location | 16,943,512 – 16,943,619 |
|---|---|
| Length | 107 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.36 |
| Mean single sequence MFE | -31.53 |
| Consensus MFE | -21.98 |
| Energy contribution | -22.47 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.546931 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16943512 107 + 22224390 UUACUAUUCGGCCACAAAGCCCAU--------UCCCAGCGACUGGAAUUAU----GUAAAUUGCUCAAAUUGGCAUCGCU-GGGAAUGGCAAGGCCAGUAAAUGAAUAGACCGGCUAAAU ...((((((((((.....(((.((--------(((((((((....((((..----...))))(((......))).)))))-)))))))))..)))).......))))))........... ( -35.41) >DroSec_CAF1 26448 107 + 1 UUACUAUUCGGCCACAAAGCCCAU--------UCCCAGCGACUGGAAUUAU----GUAAAUUGCUCAAAUUGGCAUCGCU-GGGAAUGGCAAGGCCAGUAAAUGAAUAGACCGGCUAAAU ...((((((((((.....(((.((--------(((((((((....((((..----...))))(((......))).)))))-)))))))))..)))).......))))))........... ( -35.41) >DroEre_CAF1 13175 112 + 1 UUACUAUUGGGCCACAAAACCCAU--------UCCCAGCGACUGGAAUUAUGUAUGUAAAUUGCUCAAAUUGGCAUCACUUGGGAAUGGCAAGGCCAGUGAAUGAAUAGCCCGGCUAAAU (((((....((((.......((((--------((((((.((......((((....))))..((((......)))))).).)))))))))...))))))))).....((((...))))... ( -27.10) >DroYak_CAF1 12804 116 + 1 UUACUAUUGGGCCACAAAACCCAUUCCCCCAUUCCCAGCGACUGGAAUUAU----GUAAAUUGCUCAAAUUGGCAUCACUUGGGAAUGGCAAGGCCAGUAAAUGAAUAGCCCGGCUAAAU (((((....((((.......((((((((......((((...))))......----......((((......))))......))))))))...))))))))).....((((...))))... ( -28.20) >consensus UUACUAUUCGGCCACAAAACCCAU________UCCCAGCGACUGGAAUUAU____GUAAAUUGCUCAAAUUGGCAUCACU_GGGAAUGGCAAGGCCAGUAAAUGAAUAGACCGGCUAAAU .........((((...................(((((((((......((((....))))..((((......))))))))).)))).((((...))))...............)))).... (-21.98 = -22.47 + 0.50)

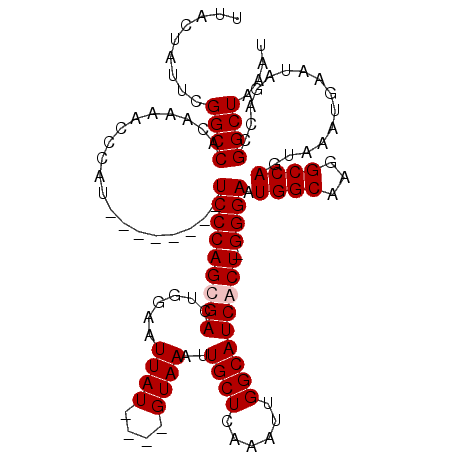
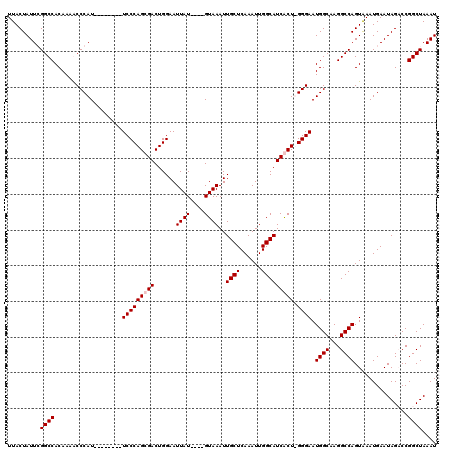
| Location | 16,943,512 – 16,943,619 |
|---|---|
| Length | 107 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.36 |
| Mean single sequence MFE | -35.49 |
| Consensus MFE | -25.32 |
| Energy contribution | -25.32 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.73 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.731371 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16943512 107 - 22224390 AUUUAGCCGGUCUAUUCAUUUACUGGCCUUGCCAUUCCC-AGCGAUGCCAAUUUGAGCAAUUUAC----AUAAUUCCAGUCGCUGGGA--------AUGGGCUUUGUGGCCGAAUAGUAA (((..(((((((............))))...((((((((-((((((...((((.((....))...----..))))...))))))))))--------)))).......)))..)))..... ( -37.60) >DroSec_CAF1 26448 107 - 1 AUUUAGCCGGUCUAUUCAUUUACUGGCCUUGCCAUUCCC-AGCGAUGCCAAUUUGAGCAAUUUAC----AUAAUUCCAGUCGCUGGGA--------AUGGGCUUUGUGGCCGAAUAGUAA (((..(((((((............))))...((((((((-((((((...((((.((....))...----..))))...))))))))))--------)))).......)))..)))..... ( -37.60) >DroEre_CAF1 13175 112 - 1 AUUUAGCCGGGCUAUUCAUUCACUGGCCUUGCCAUUCCCAAGUGAUGCCAAUUUGAGCAAUUUACAUACAUAAUUCCAGUCGCUGGGA--------AUGGGUUUUGUGGCCCAAUAGUAA .....(((((((((.........))))))..(((((((((.(((((...((((..................))))...))))))))))--------)))).......))).......... ( -30.77) >DroYak_CAF1 12804 116 - 1 AUUUAGCCGGGCUAUUCAUUUACUGGCCUUGCCAUUCCCAAGUGAUGCCAAUUUGAGCAAUUUAC----AUAAUUCCAGUCGCUGGGAAUGGGGGAAUGGGUUUUGUGGCCCAAUAGUAA ......((((((((.........))))))..(((((((((.(((((...((((.((....))...----..))))...))))))))))))))))...((((((....))))))....... ( -36.00) >consensus AUUUAGCCGGGCUAUUCAUUUACUGGCCUUGCCAUUCCC_AGCGAUGCCAAUUUGAGCAAUUUAC____AUAAUUCCAGUCGCUGGGA________AUGGGCUUUGUGGCCCAAUAGUAA .....(((((............)))))..(((...((((.((((((...((((..................))))...)))))))))).........((((((....))))))...))). (-25.32 = -25.32 + 0.00)

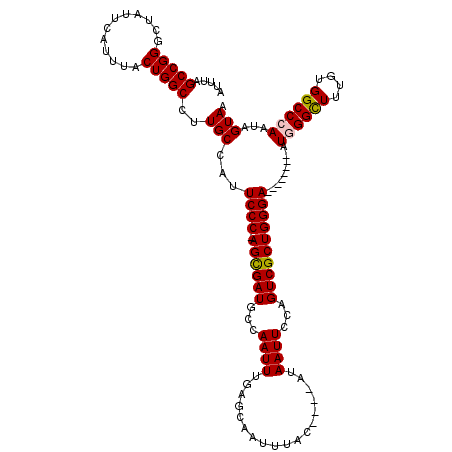
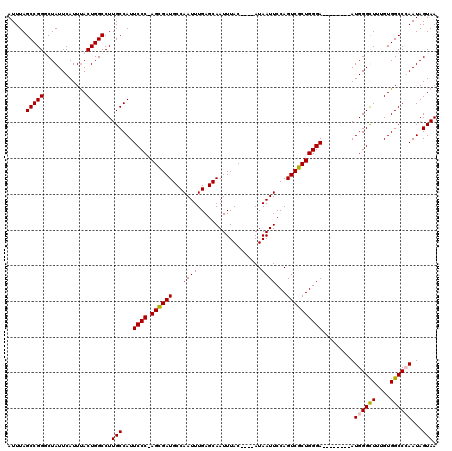
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:23:27 2006