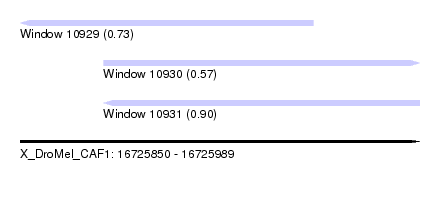

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 16,725,850 – 16,725,989 |
| Length | 139 |
| Max. P | 0.898911 |

| Location | 16,725,850 – 16,725,952 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 78.30 |
| Mean single sequence MFE | -24.90 |
| Consensus MFE | -7.70 |
| Energy contribution | -7.98 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.31 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.730729 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16725850 102 - 22224390 UCUUCAUGCAUGCCUACAAUGCCGUACCGCACUGUAUCCUACAUAUAUGUACAU-------ACAGGCAUUACAGGCAAUCGCAUCGCAUCGAUCUUAAUCGCAUAUGUA----------- ...((((((((((......(((((((..((.((((((..((((....)))).))-------))))))..))).))))...)))).)))).)).................----------- ( -24.70) >DroSec_CAF1 6719 103 - 1 ACUUCAUGCAUGCCUACAAUGCCGUACCGCACUGUAUCCUA--UAUAUGUAAAGUCUGGUCGCAGUCAUU----GUAAUCGCAUCGCAUCGAUCUCAAUCGCAUAUGGA----------- ...((((((((((.((((((((.((((((.(((.(((....--.....))).))).)))).)).).))))----)))...)))).)))).)).................----------- ( -21.80) >DroSim_CAF1 6112 103 - 1 ACUUCAUGCAUGCCUACAAUGCCGUACCGCACUGUAUCCUA--UAUAUGUAAAGUCUGGUCGCAGUCAUU----GUAAUCGCAUCGCAUCGAUCUCAAUCGCAUAUGGA----------- ...((((((((((.((((((((.((((((.(((.(((....--.....))).))).)))).)).).))))----)))...)))).)))).)).................----------- ( -21.80) >DroEre_CAF1 5374 108 - 1 UUUUCAUGCUUACCUACAAUGCCGUAACGCACUGUAUCCCA--UAUA------GUCUAGUCGCAGGCAUU----GUGAUCGCAUCCCAUCGAUCUCAAUCGAACAUGGAUACCGCCAGUU ..((((((......((((((((((..((..(((((((...)--))))------))...))..).))))))----)))...........(((((....))))).))))))........... ( -28.00) >DroYak_CAF1 5666 97 - 1 GUGUCAUGCAUACCUACAAUCCCGUACCGCACUGUAUCCUA--UAUA------GUGCAGUACCAGGCAUU----GUAAUCGCAUCCCAUCGAUCUCAAUCGCAAAUGGA----------- .....((((.....((((((((.((((.(((((((((...)--))))------)))).))))..)).)))----)))...)))).((((((((....))))...)))).----------- ( -28.20) >consensus ACUUCAUGCAUGCCUACAAUGCCGUACCGCACUGUAUCCUA__UAUAUGUA_AGUCUAGUCGCAGGCAUU____GUAAUCGCAUCGCAUCGAUCUCAAUCGCAUAUGGA___________ .....((((.(((....((((((.........................................))))))....)))...)))).((((((((....))))...))))............ ( -7.70 = -7.98 + 0.28)

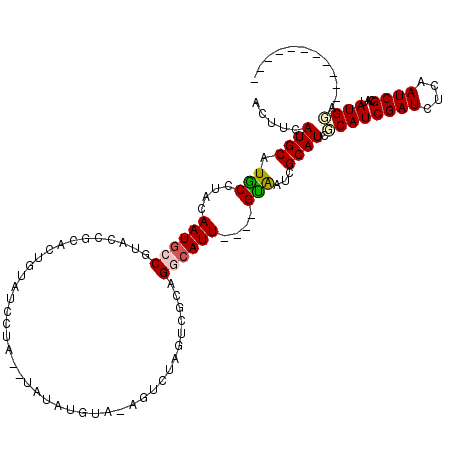
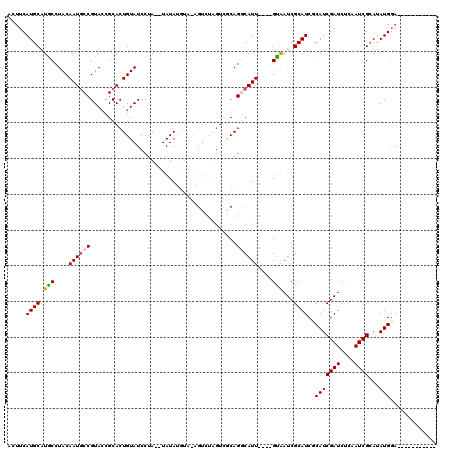
| Location | 16,725,879 – 16,725,989 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 75.66 |
| Mean single sequence MFE | -29.16 |
| Consensus MFE | -18.64 |
| Energy contribution | -18.40 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.572867 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16725879 110 + 22224390 GAUUGCCUGUAAUGCCUGU-------AUGUACAUAUAUGUAGGAUACAGUGCGGUACGGCAUUGUAGGCAUGCAUGAAGAUAUGCUGCCCU--UACUGAGAACUUUU-UUGUUGUGCUAU ....(((((((...(((((-------((((....))))))))).))))).))(((((((((.....((((.(((((....)))))))))((--(...))).......-.))))))))).. ( -33.50) >DroSec_CAF1 6748 113 + 1 GAUUAC----AAUGACUGCGACCAGACUUUACAUAUA--UAGGAUACAGUGCGGUACGGCAUUGUAGGCAUGCAUGAAGUUAUGCUGCCCUUUUACUAAGAACUUCU-UUGUUGUGCUAU ...(((----((..((((....)))............--(((..((((((((......))))))))((((.(((((....)))))))))......))).........-)..))))).... ( -26.90) >DroSim_CAF1 6141 113 + 1 GAUUAC----AAUGACUGCGACCAGACUUUACAUAUA--UAGGAUACAGUGCGGUACGGCAUUGUAGGCAUGCAUGAAGUUAUGCUGCCCUUUUACUAAGAACUUCU-UUGUUGUGCUAU ...(((----((..((((....)))............--(((..((((((((......))))))))((((.(((((....)))))))))......))).........-)..))))).... ( -26.90) >DroEre_CAF1 5414 102 + 1 GAUCAC----AAUGCCUGCGACUAGAC------UAUA--UGGGAUACAGUGCGUUACGGCAUUGUAGGUAAGCAUGAAAAUAUGCCGCCUG--UACUGUGAAGAUUUGUU----UUCUUG ..((((----(.(((..(((.......------....--.....((((((((......)))))))).....(((((....))))))))..)--)).))))).........----...... ( -25.10) >DroYak_CAF1 5695 101 + 1 GAUUAC----AAUGCCUGGUACUGCAC------UAUA--UAGGAUACAGUGCGGUACGGGAUUGUAGGUAUGCAUGACACUAUGCUGCCAU--AACUAAUAAGAUAU-UU----UUCUAC ...(((----(((.((((.((((((((------(.((--(...))).)))))))))))))))))))((((.(((((....)))))))))..--........(((...-..----.))).. ( -33.40) >consensus GAUUAC____AAUGCCUGCGACCAGACU_UACAUAUA__UAGGAUACAGUGCGGUACGGCAUUGUAGGCAUGCAUGAAGAUAUGCUGCCCU__UACUAAGAACUUCU_UUGUUGUGCUAU .......................................(((..((((((((......))))))))((((.(((((....)))))))))......)))...................... (-18.64 = -18.40 + -0.24)

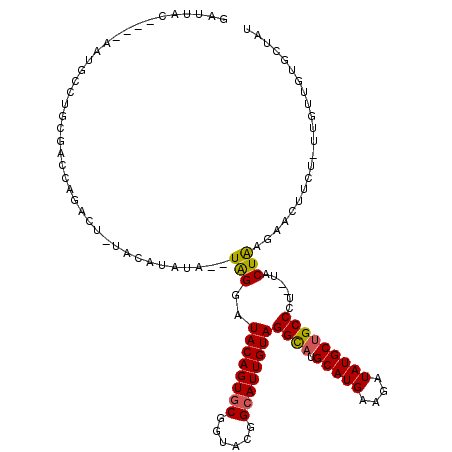
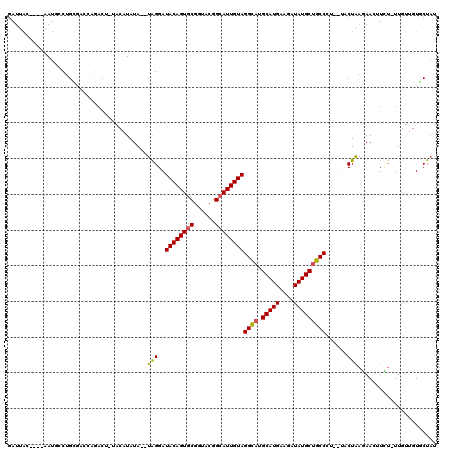
| Location | 16,725,879 – 16,725,989 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 75.66 |
| Mean single sequence MFE | -25.80 |
| Consensus MFE | -11.20 |
| Energy contribution | -11.64 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.43 |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.898911 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16725879 110 - 22224390 AUAGCACAACAA-AAAAGUUCUCAGUA--AGGGCAGCAUAUCUUCAUGCAUGCCUACAAUGCCGUACCGCACUGUAUCCUACAUAUAUGUACAU-------ACAGGCAUUACAGGCAAUC ............-..............--.(((((((((......)))).)))))....(((((((..((.((((((..((((....)))).))-------))))))..))).))))... ( -28.60) >DroSec_CAF1 6748 113 - 1 AUAGCACAACAA-AGAAGUUCUUAGUAAAAGGGCAGCAUAACUUCAUGCAUGCCUACAAUGCCGUACCGCACUGUAUCCUA--UAUAUGUAAAGUCUGGUCGCAGUCAUU----GUAAUC ........((.(-((.....))).))....(((((((((......)))).)))))(((((((.((((((.(((.(((....--.....))).))).)))).)).).))))----)).... ( -24.10) >DroSim_CAF1 6141 113 - 1 AUAGCACAACAA-AGAAGUUCUUAGUAAAAGGGCAGCAUAACUUCAUGCAUGCCUACAAUGCCGUACCGCACUGUAUCCUA--UAUAUGUAAAGUCUGGUCGCAGUCAUU----GUAAUC ........((.(-((.....))).))....(((((((((......)))).)))))(((((((.((((((.(((.(((....--.....))).))).)))).)).).))))----)).... ( -24.10) >DroEre_CAF1 5414 102 - 1 CAAGAA----AACAAAUCUUCACAGUA--CAGGCGGCAUAUUUUCAUGCUUACCUACAAUGCCGUAACGCACUGUAUCCCA--UAUA------GUCUAGUCGCAGGCAUU----GUGAUC ...(((----........)))......--.(((.(((((......)))))..)))(((((((((..((..(((((((...)--))))------))...))..).))))))----)).... ( -25.10) >DroYak_CAF1 5695 101 - 1 GUAGAA----AA-AUAUCUUAUUAGUU--AUGGCAGCAUAGUGUCAUGCAUACCUACAAUCCCGUACCGCACUGUAUCCUA--UAUA------GUGCAGUACCAGGCAUU----GUAAUC ......----..-...........((.--((((((......)))))))).....((((((((.((((.(((((((((...)--))))------)))).))))..)).)))----)))... ( -27.10) >consensus AUAGCACAACAA_AAAAGUUCUUAGUA__AGGGCAGCAUAACUUCAUGCAUGCCUACAAUGCCGUACCGCACUGUAUCCUA__UAUAUGUA_AGUCUAGUCGCAGGCAUU____GUAAUC ...............................((((((((......)))).))))((((.(((......))).))))............................................ (-11.20 = -11.64 + 0.44)
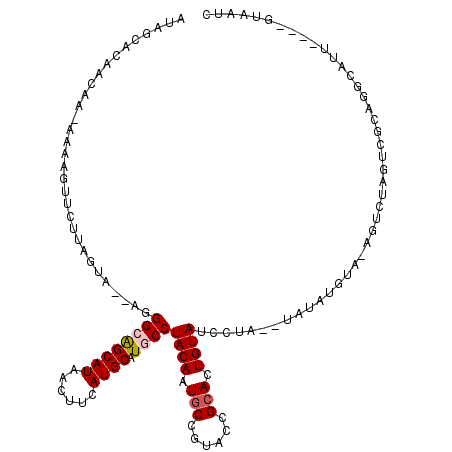


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:20:14 2006