| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,809,261 – 1,809,362 |
| Length | 101 |
| Max. P | 0.998152 |

| Location | 1,809,261 – 1,809,362 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 90.27 |
| Mean single sequence MFE | -44.58 |
| Consensus MFE | -41.62 |
| Energy contribution | -41.19 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.06 |
| Mean z-score | -3.29 |
| Structure conservation index | 0.93 |
| SVM decision value | 3.02 |
| SVM RNA-class probability | 0.998152 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1809261 101 + 22224390 CCAGUUGGCUGGCGCCAUCUGUGCA--------GCAGCCACCAGACAUAGAGGGCGCCAGAGGUUGGUGCGCCGGCUACGGCAAAGCCAAGACACCAACCAAGUCGACC ...(((((((((((((.(((((((.--------..........).)))))).))))))...(((((((((...((((.......))))..).)))))))).))))))). ( -47.60) >DroSec_CAF1 17677 99 + 1 -CAGCUGACUGGCGCCAUCUGUGCA--------GCAGCCACCAGACAUAGAGGGCGCCAGAGGUUGGUGCGCUGGCUAA-GCAAAGUCAAGACACCAACCAAGUCGACC -(.(((..((((((((.(((((((.--------..........).)))))).)))))))).(((((((((..(((((..-....))))).).)))))))).))).)... ( -44.00) >DroSim_CAF1 19626 100 + 1 -CAGCUGACUGGCGCCAUCUGUGCA--------GCAGCCACCAGACAUAGAGGGCGCCAGAGGUUGGUGCGCUGGCUAAGGCAAAGUCAAGACACCAACCAAGUCGACC -(.(((..((((((((.(((((((.--------..........).)))))).)))))))).(((((((((..(((((.......))))).).)))))))).))).)... ( -43.50) >DroYak_CAF1 18388 104 + 1 -----UGGCUGGCGCCAUCUGUGCAGCGCAGCAGCAGCCACCAGACAUAGAGGGCGCCAGAGGUUGGUGCGCCGGCUAAGGCAAAGUCAAGACACCAACCAAGUCGACG -----...((((((((.(((((((.((((....)).)).....).)))))).)))))))).(((((((((...((((.......))))..).))))))))......... ( -43.20) >consensus _CAGCUGACUGGCGCCAUCUGUGCA________GCAGCCACCAGACAUAGAGGGCGCCAGAGGUUGGUGCGCCGGCUAAGGCAAAGUCAAGACACCAACCAAGUCGACC .....(((((((((((.(((((((...................).)))))).))))))...(((((((((...((((.......))))..).)))))))).)))))... (-41.62 = -41.19 + -0.44)

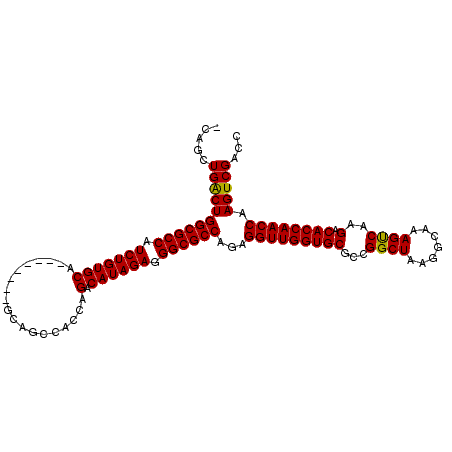
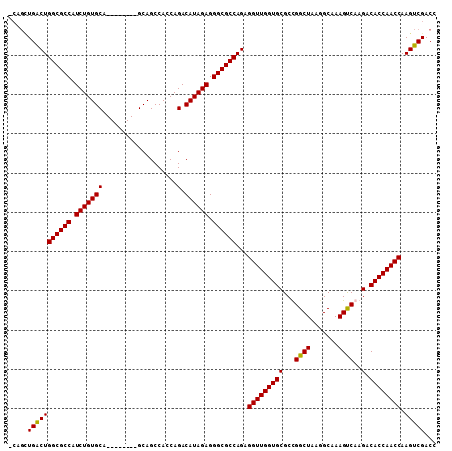
| Location | 1,809,261 – 1,809,362 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 90.27 |
| Mean single sequence MFE | -45.30 |
| Consensus MFE | -39.15 |
| Energy contribution | -38.90 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.64 |
| Structure conservation index | 0.86 |
| SVM decision value | 2.02 |
| SVM RNA-class probability | 0.985733 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1809261 101 - 22224390 GGUCGACUUGGUUGGUGUCUUGGCUUUGCCGUAGCCGGCGCACCAACCUCUGGCGCCCUCUAUGUCUGGUGGCUGC--------UGCACAGAUGGCGCCAGCCAACUGG ((.......(((((((((.((((((.......)))))).))))))))).((((((((......(((((...((...--------.)).)))))))))))))))...... ( -47.80) >DroSec_CAF1 17677 99 - 1 GGUCGACUUGGUUGGUGUCUUGACUUUGC-UUAGCCAGCGCACCAACCUCUGGCGCCCUCUAUGUCUGGUGGCUGC--------UGCACAGAUGGCGCCAGUCAGCUG- (((.(((..(((((((((.(((.((....-..)).))).)))))))))..(((((((......(((((...((...--------.)).))))))))))))))).))).- ( -43.70) >DroSim_CAF1 19626 100 - 1 GGUCGACUUGGUUGGUGUCUUGACUUUGCCUUAGCCAGCGCACCAACCUCUGGCGCCCUCUAUGUCUGGUGGCUGC--------UGCACAGAUGGCGCCAGUCAGCUG- (((.(((..(((((((((.(((.((.......)).))).)))))))))..(((((((......(((((...((...--------.)).))))))))))))))).))).- ( -43.20) >DroYak_CAF1 18388 104 - 1 CGUCGACUUGGUUGGUGUCUUGACUUUGCCUUAGCCGGCGCACCAACCUCUGGCGCCCUCUAUGUCUGGUGGCUGCUGCUGCGCUGCACAGAUGGCGCCAGCCA----- .........(((((((((.........(((......)))))))))))).((((((((......((((((..((.((....))))..).)))))))))))))...----- ( -46.50) >consensus GGUCGACUUGGUUGGUGUCUUGACUUUGCCUUAGCCAGCGCACCAACCUCUGGCGCCCUCUAUGUCUGGUGGCUGC________UGCACAGAUGGCGCCAGCCAGCUG_ .........(((((((((.(((.((.......)).))).))))))))).((((((((......(((((...((............)).)))))))))))))........ (-39.15 = -38.90 + -0.25)

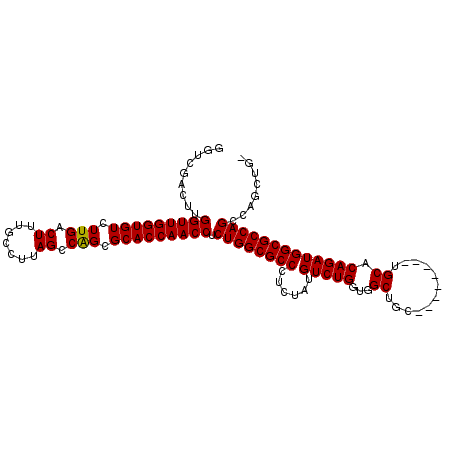
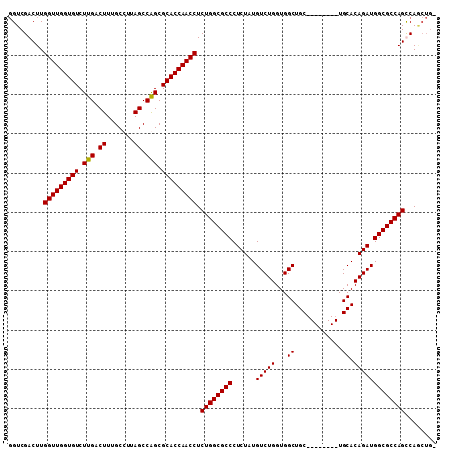
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:50:09 2006