
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 16,158,244 – 16,158,344 |
| Length | 100 |
| Max. P | 0.999994 |

| Location | 16,158,244 – 16,158,344 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 71.41 |
| Mean single sequence MFE | -30.72 |
| Consensus MFE | -21.69 |
| Energy contribution | -22.50 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.22 |
| Mean z-score | -3.96 |
| Structure conservation index | 0.71 |
| SVM decision value | 5.83 |
| SVM RNA-class probability | 0.999994 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16158244 100 + 22224390 ACACCAAAUUCUGACGUUCACUUUCGCAAAUCAAAGUCAGUUUAGGCG------------UUUAGCCUAACUCAUUGAUUGUAGCCAACGAAAGUGAAACUCGGAAUGGAAA ...(((..((((((..((((((((((........(((((((((((((.------------....))))))...)))))))........))))))))))..)))))))))... ( -35.79) >DroSec_CAF1 4156 86 + 1 GUGCUUAGUUCUGACGUUCACUUUCGCAAAUCAAAG----UCUA----------------------CUGACUCUUUGAUUGUAGCCAACGAAAGUGAAAAUCAGAGUGGAAA ....((..((((((..(((((((((((.((((((((----((..----------------------..))))..))))))...))....)))))))))..))))))..)).. ( -27.00) >DroSim_CAF1 5029 90 + 1 GCUCCAAGUUCUGACGUUCACUUUCGCAAAUCAAAGUCAGUCUA----------------------CUGACUCUUUGAUUGUAGCCAACGAAAGUGAAAAUCAGAGUGGAAA ..((((...(((((..(((((((((((.((((((((((((....----------------------))))))..))))))...))....)))))))))..))))).)))).. ( -35.10) >DroYak_CAF1 4940 105 + 1 -------GUUCUGACGUUUACUUUCGGAAAUCAAAGUCAAUUCGGGCAGUGCAUUCGUAUUUUACCCUAGCAUUUUGAUUGUAUGCAACGAAAGUGAAACUCGGCGUGGAAA -------...((((..((((((((((..(((((((((......(((.(((((....)))))...)))....)))))))))........))))))))))..))))........ ( -25.00) >consensus GC_CCAAGUUCUGACGUUCACUUUCGCAAAUCAAAGUCAGUCUA______________________CUAACUCUUUGAUUGUAGCCAACGAAAGUGAAAAUCAGAGUGGAAA ........((((((..((((((((((..((((((((..((((.((((.................))))))))))))))))........))))))))))..))))))...... (-21.69 = -22.50 + 0.81)
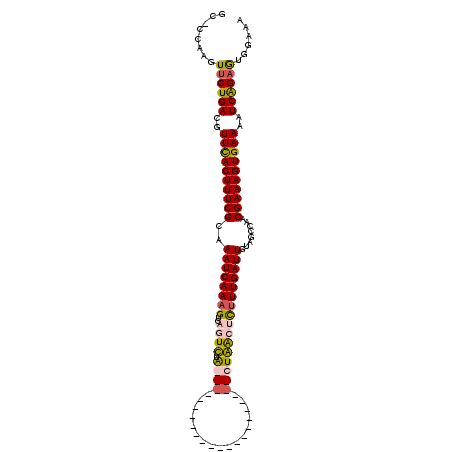


| Location | 16,158,244 – 16,158,344 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 71.41 |
| Mean single sequence MFE | -28.48 |
| Consensus MFE | -17.45 |
| Energy contribution | -20.52 |
| Covariance contribution | 3.06 |
| Combinations/Pair | 1.09 |
| Mean z-score | -3.34 |
| Structure conservation index | 0.61 |
| SVM decision value | 5.13 |
| SVM RNA-class probability | 0.999975 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16158244 100 - 22224390 UUUCCAUUCCGAGUUUCACUUUCGUUGGCUACAAUCAAUGAGUUAGGCUAAA------------CGCCUAAACUGACUUUGAUUUGCGAAAGUGAACGUCAGAAUUUGGUGU ...((((((.((..(((((((((((.......(((((((.((((((((....------------.)))).)))).)..)))))).)))))))))))..)).)))..)))... ( -29.30) >DroSec_CAF1 4156 86 - 1 UUUCCACUCUGAUUUUCACUUUCGUUGGCUACAAUCAAAGAGUCAG----------------------UAGA----CUUUGAUUUGCGAAAGUGAACGUCAGAACUAAGCAC .......((((((.(((((((((((.......(((((..(((((..----------------------..))----)))))))).))))))))))).))))))......... ( -28.50) >DroSim_CAF1 5029 90 - 1 UUUCCACUCUGAUUUUCACUUUCGUUGGCUACAAUCAAAGAGUCAG----------------------UAGACUGACUUUGAUUUGCGAAAGUGAACGUCAGAACUUGGAGC .(((((.((((((.(((((((((((.......(((((..(((((((----------------------....)))))))))))).))))))))))).))))))...))))). ( -37.20) >DroYak_CAF1 4940 105 - 1 UUUCCACGCCGAGUUUCACUUUCGUUGCAUACAAUCAAAAUGCUAGGGUAAAAUACGAAUGCACUGCCCGAAUUGACUUUGAUUUCCGAAAGUAAACGUCAGAAC------- ..........((((((.(((((((..((((.........))))..(((((..............))))).................))))))))))).)).....------- ( -18.94) >consensus UUUCCACUCCGAGUUUCACUUUCGUUGGCUACAAUCAAAGAGUCAG______________________UAAACUGACUUUGAUUUGCGAAAGUGAACGUCAGAACUUGG_GC .......((((((.(((((((((((.......((((((((((((.((((................)))).))))..)))))))).))))))))))).))))))......... (-17.45 = -20.52 + 3.06)
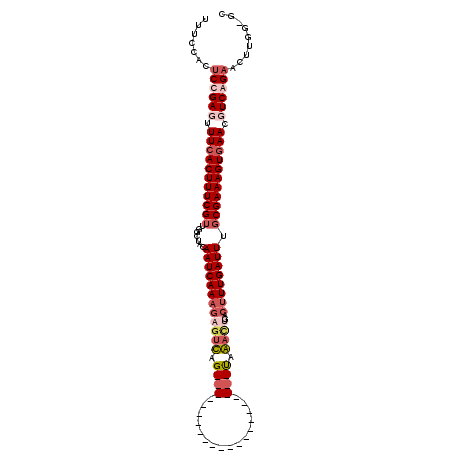


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:14:01 2006