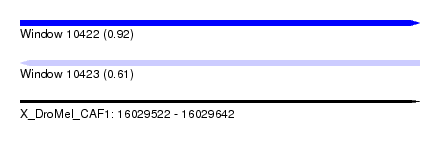
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 16,029,522 – 16,029,642 |
| Length | 120 |
| Max. P | 0.921605 |

| Location | 16,029,522 – 16,029,642 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.10 |
| Mean single sequence MFE | -35.93 |
| Consensus MFE | -34.72 |
| Energy contribution | -34.83 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.14 |
| SVM RNA-class probability | 0.921605 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16029522 120 + 22224390 AUUGAUUUUACACCCAUGUGUCCAAAGAGUCACGGUCUGGAGUAGCCGACCGAUUCGGAUUCGCUGGAAAUAAUGUCCAGUCGAUCCGUUUGCAAAUAUGUAUAUCUGGCUCAAUAAUAA .........((((....)))).....(((((((((((.((.....)))))))...((((((.((((((.......)))))).))))))..((((....))))....))))))........ ( -35.50) >DroEre_CAF1 97483 116 + 1 AUUGAUUCUACACCCAUGUGUCCAAAGAGUCACGGUCUGGAGUAGCCGACCGAUGCGGAUUCGCUGGAAAUAGUGUCCAGUCGAGCCGUUUGCAAAUA----UAUCUGGCUCAAUAAUAG .........((((....)))).....(((((((((((.((.....)))))))..((((.(((((((((.......))))).)))))))).........----....))))))........ ( -35.00) >DroYak_CAF1 80209 116 + 1 AUUGAUUCUACACCCAUGUGUCCAAAGAGUCACGGUCUGGAGUAGCCGACCGAUUCGGGUUCGCUGGAAAUAGUGUCCAGUCGACCCGUUUGCAAAUA----UAUCUGGCUCAAUAAUAA .........((((....)))).....(((((((((((.((.....)))))))...((((((.((((((.......)))))).))))))..........----....))))))........ ( -37.30) >consensus AUUGAUUCUACACCCAUGUGUCCAAAGAGUCACGGUCUGGAGUAGCCGACCGAUUCGGAUUCGCUGGAAAUAGUGUCCAGUCGACCCGUUUGCAAAUA____UAUCUGGCUCAAUAAUAA .........((((....)))).....(((((((((((.((.....)))))))...((((((.((((((.......)))))).))))))..................))))))........ (-34.72 = -34.83 + 0.11)

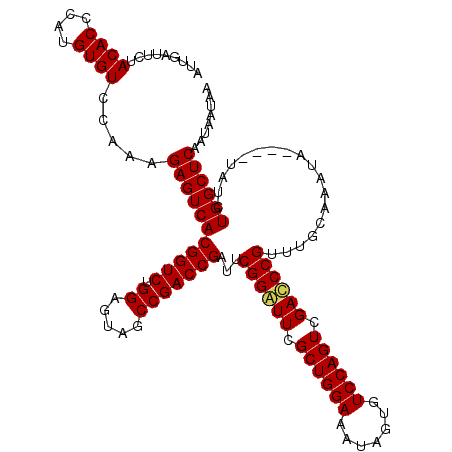
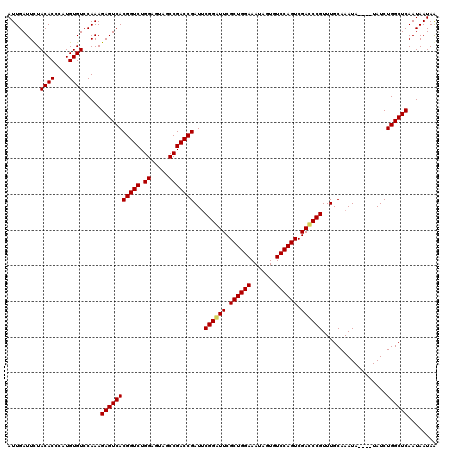
| Location | 16,029,522 – 16,029,642 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.10 |
| Mean single sequence MFE | -33.00 |
| Consensus MFE | -31.85 |
| Energy contribution | -31.97 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.605852 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 16029522 120 - 22224390 UUAUUAUUGAGCCAGAUAUACAUAUUUGCAAACGGAUCGACUGGACAUUAUUUCCAGCGAAUCCGAAUCGGUCGGCUACUCCAGACCGUGACUCUUUGGACACAUGGGUGUAAAAUCAAU .....(((((.(((((................(((((((.(((((.......))))))).)))))...(((((((.....)).)))))......)))))((((....))))....))))) ( -32.70) >DroEre_CAF1 97483 116 - 1 CUAUUAUUGAGCCAGAUA----UAUUUGCAAACGGCUCGACUGGACACUAUUUCCAGCGAAUCCGCAUCGGUCGGCUACUCCAGACCGUGACUCUUUGGACACAUGGGUGUAGAAUCAAU ......(((((((.....----...........)))))))(((((.......))))).((.((..((((((((((.....)).))))(((.((....)).)))...))))..)).))... ( -31.59) >DroYak_CAF1 80209 116 - 1 UUAUUAUUGAGCCAGAUA----UAUUUGCAAACGGGUCGACUGGACACUAUUUCCAGCGAACCCGAAUCGGUCGGCUACUCCAGACCGUGACUCUUUGGACACAUGGGUGUAGAAUCAAU .....(((((.(((((..----..........(((((((.(((((.......))))))).)))))...(((((((.....)).)))))......)))))((((....))))....))))) ( -34.70) >consensus UUAUUAUUGAGCCAGAUA____UAUUUGCAAACGGAUCGACUGGACACUAUUUCCAGCGAAUCCGAAUCGGUCGGCUACUCCAGACCGUGACUCUUUGGACACAUGGGUGUAGAAUCAAU .....(((((.(((((................(((((((.(((((.......))))))).)))))...(((((((.....)).)))))......)))))((((....))))....))))) (-31.85 = -31.97 + 0.11)

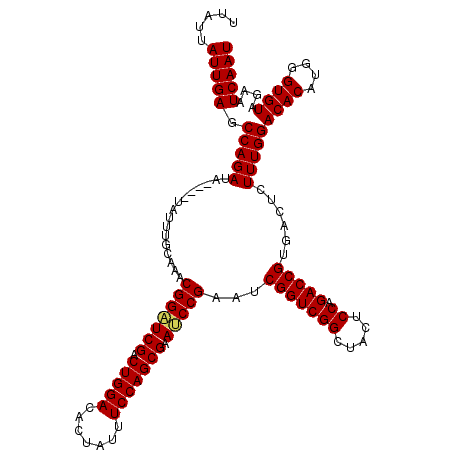
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:12:23 2006