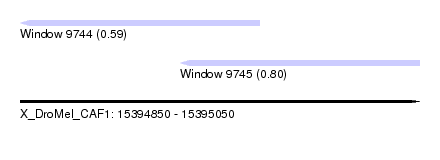
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 15,394,850 – 15,395,050 |
| Length | 200 |
| Max. P | 0.802019 |

| Location | 15,394,850 – 15,394,970 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.65 |
| Mean single sequence MFE | -26.14 |
| Consensus MFE | -20.27 |
| Energy contribution | -21.90 |
| Covariance contribution | 1.63 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.590269 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
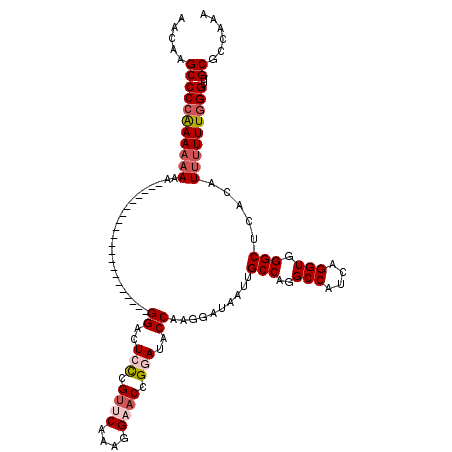
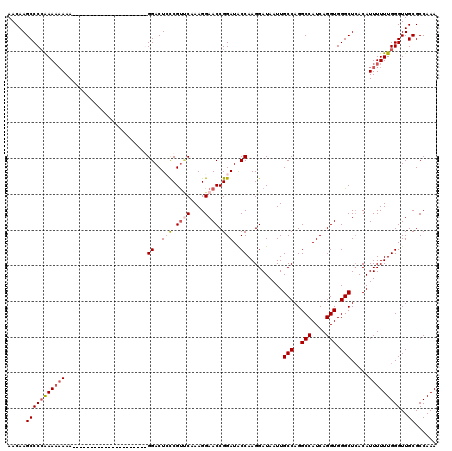
>X_DroMel_CAF1 15394850 120 - 22224390 AACAAGCCCAAAAAAAAAAAAAAAAAAACAACAAAAUAGGGACUCCCGUCCCAAGGAACCGGAUACCAAGGAUAAUUGCCAGGCCAUCAGGUGGGCUCACAUUUUUUGGGUUGCGCCAAA ....(((((((((((.......................(((((....)))))...((.((.........))......(((..(((....))).)))))...)))))))))))........ ( -30.30) >DroSec_CAF1 41247 110 - 1 AACAAGCCCCAAAAAAAA----AAAA------AAAAUAGGGACUACCGUUCAAAGGAACCGGAUGCCAAGGAUAAUUGCCAGGCCAUCAGGUGGGCUCACAUUUUUUGGGUUGCGCCAAA .....(((((((((((..----....------......(((.(((((((((....))))..((((((..((.......)).)).)))).))))).)))...)))))))))..))...... ( -27.06) >DroEre_CAF1 32159 97 - 1 AACAAGCCCCGAA--AAA---------------------GGACUGCCGUUCAAAGGAACCGGAUACCAAGGAUAAUUGCCAGGCCAUCAGGUGGGCUCACAUUUUUUGGGUUGCGCCAAA .....((((((((--(((---------------------((....((((((....))).)))...))..........(((..(((....))).))).....)))))))))..))...... ( -26.20) >DroYak_CAF1 23865 99 - 1 AACAAGCCCCAAAAAAAA---------------------GGACUCUCGCACAAAGGAACCGGAUACCAAGGAUAAUUGCCAGGCCAUCAGGUGGGCUAACAUUUUUUGGGUUGCGCCAAA .....(((((((((((..---------------------((..(((........))).)).................(((..(((....))).))).....)))))))))..))...... ( -21.00) >consensus AACAAGCCCCAAAAAAAA_____________________GGACUCCCGUUCAAAGGAACCGGAUACCAAGGAUAAUUGCCAGGCCAUCAGGUGGGCUCACAUUUUUUGGGUUGCGCCAAA .....(((((((((((.......................((..(((.((((....)))).)))..))..........(((..(((....))).))).....)))))))))..))...... (-20.27 = -21.90 + 1.63)

| Location | 15,394,930 – 15,395,050 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.13 |
| Mean single sequence MFE | -30.20 |
| Consensus MFE | -27.00 |
| Energy contribution | -27.12 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.802019 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
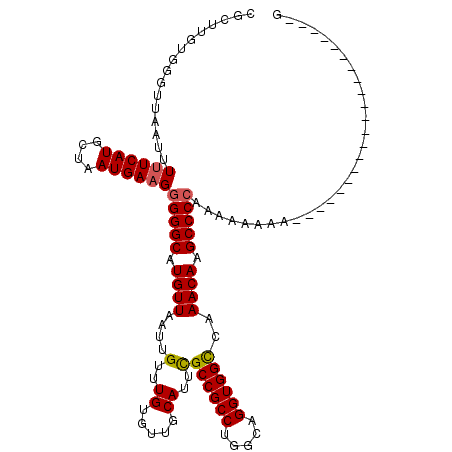
>X_DroMel_CAF1 15394930 120 - 22224390 CGCUUGUGGUUUAAUUUUUCAUGCUAAUGAAGGGGGCAUGUUAAUUGUUUGUGUUGCAUUCGCCGCCUGGCAGGUGGCCAAACAAGCCCAAAAAAAAAAAAAAAAAAACAACAAAAUAGG ...((((.((((...(((((((....)))))))((((.((((....(..((.....))..)((((((.....))))))..)))).))))................)))).))))...... ( -31.50) >DroSec_CAF1 41327 110 - 1 CGCUUGUGGGUUAAUUUUUCAUGCUAAUGAAGGGGGCAUGUUAAUUGUUUGUGUUGCAUUCGCCGCCUAGCAGGUGGCCAAACAAGCCCCAAAAAAAA----AAAA------AAAAUAGG ..(((((.........((((((....))))))(((((.((((....(..((.....))..)((((((.....))))))..)))).)))))........----....------...))))) ( -31.10) >DroEre_CAF1 32239 97 - 1 CGCUUGUGGGUUAAUUUGUCAUGCUAAUGAAGGGGGCAUGUUAAUUGUUUGUGUUGCAUUCGCCGCCUGGCAGGUGGCCAAACAAGCCCCGAA--AAA---------------------G ...(((.(((((...(((.((((((.........)))))).))).((((((.((..(....(((....)))..)..)))))))))))))))).--...---------------------. ( -29.20) >DroYak_CAF1 23945 99 - 1 CGCUUGUGGGUUAAUUUUUCAUGCUAAUGAAGGGGGCAUGUUAAUUGUUUGUGUUGCAUUUGCCGCCUGGCAGGUGGUCAAACAAGCCCCAAAAAAAA---------------------G ...(((.(((((...((..((((((.........))))))..)).((((((.....((((((((....))))))))..))))))))))))))......---------------------. ( -29.00) >consensus CGCUUGUGGGUUAAUUUUUCAUGCUAAUGAAGGGGGCAUGUUAAUUGUUUGUGUUGCAUUCGCCGCCUGGCAGGUGGCCAAACAAGCCCCAAAAAAAA_____________________G ................((((((....))))))(((((.((((....(..((.....))..)((((((.....))))))..)))).))))).............................. (-27.00 = -27.12 + 0.12)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:01:37 2006