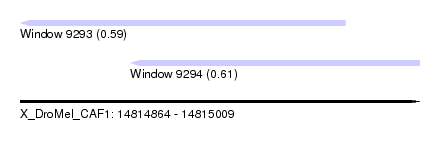

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 14,814,864 – 14,815,009 |
| Length | 145 |
| Max. P | 0.610763 |

| Location | 14,814,864 – 14,814,982 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 92.23 |
| Mean single sequence MFE | -33.83 |
| Consensus MFE | -33.16 |
| Energy contribution | -33.04 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.26 |
| Structure conservation index | 0.98 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.589852 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
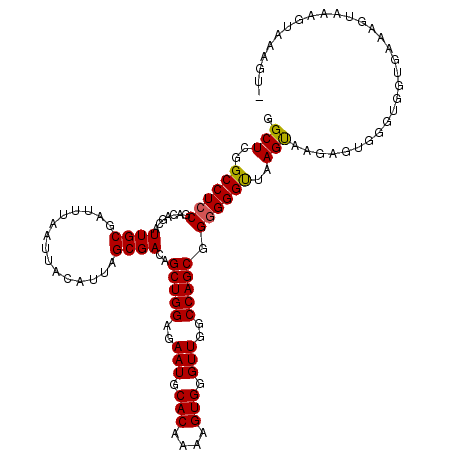
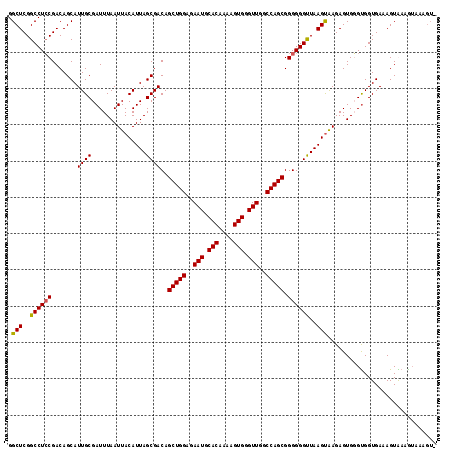
>X_DroMel_CAF1 14814864 118 - 22224390 GGCUCGGCCUACGACAGCAUUGCGAUUUAAUUACAUUAGCGACAGCUGGAGAAUGCACAAAAGUGGGUUGGCCAGCGGGGGGUUAAGCAAGAGUGUGUGGUGAAAGUGAAGUAAAGUA (((...)))(((..((.(((..((...........(((((..(.(((((..(((.(((....))).)))..)))))..)..))))).........))..)))....))..)))..... ( -30.95) >DroSim_CAF1 8093 111 - 1 GGCUCGGCCUCCGACAGCAUUGCGAUUUAAUUACAUUAGCGACAGCUGGAGAAUGCACAAAAGUGGGUUGGCCAGCGGGGGGCUAAGUGAGAGUGUGUGGUGAAAGUAAAG------- ...((((...))))...(((..((......((((.(((((..(.(((((..(((.(((....))).)))..)))))..)..))))))))).....))..))).........------- ( -35.60) >DroEre_CAF1 13943 118 - 1 GGCUCGGCCUCCGACAGCAUUGCGAUUUAAUUACAUUAGCGACAGCUGGAGAAUGCACAAAAGUGGGUUGGCCAGCGGGGGGUUAAGUAAGAGUGGGCGGUGAAAGUGAAGUACAGUU .(((..((((((.......((((...............))))..(((((..(((.(((....))).)))..))))).))))))..)))..((.((..(............)..)).)) ( -34.36) >DroYak_CAF1 1514 118 - 1 GGCUCGGCCUCCGACAGCAUUGCGAUUUAAUUACAUUAGCGACAGCUGGAGAAUGCACAAAAGUGGGUUGGCCAGCGGGGGGUUAAGUAAGAGUGGGUGGUGAAAGUAAUAUAAAGUG .(((..(((.((.((.((...)).......((((.(((((..(.(((((..(((.(((....))).)))..)))))..)..)))))))))..)).)).)))...)))........... ( -34.40) >consensus GGCUCGGCCUCCGACAGCAUUGCGAUUUAAUUACAUUAGCGACAGCUGGAGAAUGCACAAAAGUGGGUUGGCCAGCGGGGGGUUAAGUAAGAGUGGGUGGUGAAAGUAAAGUAAAGU_ .(((..((((((.......((((...............))))..(((((..(((.(((....))).)))..))))).))))))..))).............................. (-33.16 = -33.04 + -0.12)

| Location | 14,814,904 – 14,815,009 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 99.62 |
| Mean single sequence MFE | -27.80 |
| Consensus MFE | -27.80 |
| Energy contribution | -27.80 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.26 |
| Structure conservation index | 1.00 |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.610763 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
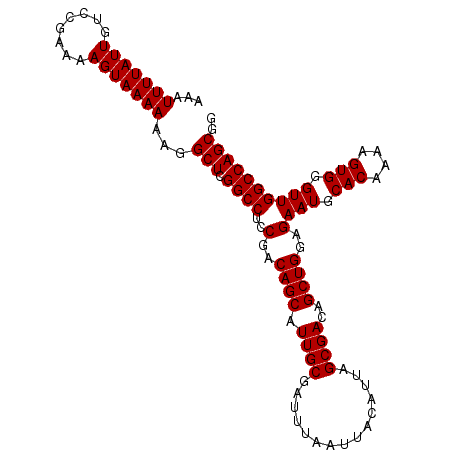
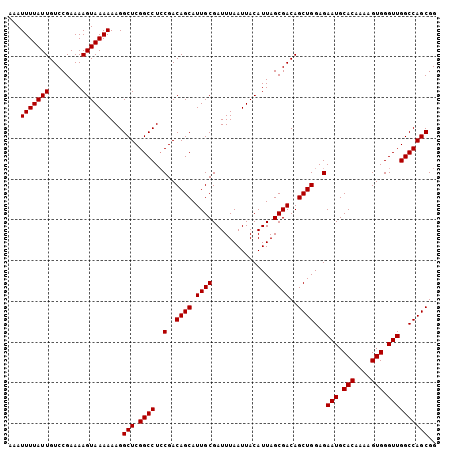
>X_DroMel_CAF1 14814904 105 - 22224390 AAAUUUUAUUGUCCGAAAAGUAAAAAAGGCUCGGCCUACGACAGCAUUGCGAUUUAAUUACAUUAGCGACAGCUGGAGAAUGCACAAAAGUGGGUUGGCCAGCGG ...(((((((........)))))))...(((.((((..(..((((.((((...............))))..))))..)(((.(((....))).)))))))))).. ( -27.96) >DroSec_CAF1 8878 105 - 1 AAAUUUUAUUGUCCGAAAAGUAAAAAAGGCUCGGCCUCCGACAGCAUUGCGAUUUAAUUACAUUAGCGACAGCUGGAGAAUGCACAAAAGUGGGUUGGCCAGCGG ...(((((((........)))))))...(((.((((..(..((((.((((...............))))..))))..)(((.(((....))).)))))))))).. ( -27.76) >DroSim_CAF1 8126 105 - 1 AAAUUUUAUUGUCCGAAAAGUAAAAAAGGCUCGGCCUCCGACAGCAUUGCGAUUUAAUUACAUUAGCGACAGCUGGAGAAUGCACAAAAGUGGGUUGGCCAGCGG ...(((((((........)))))))...(((.((((..(..((((.((((...............))))..))))..)(((.(((....))).)))))))))).. ( -27.76) >DroEre_CAF1 13983 105 - 1 AAAUUUUAUUGUCCGAAAAGUAAAAAAGGCUCGGCCUCCGACAGCAUUGCGAUUUAAUUACAUUAGCGACAGCUGGAGAAUGCACAAAAGUGGGUUGGCCAGCGG ...(((((((........)))))))...(((.((((..(..((((.((((...............))))..))))..)(((.(((....))).)))))))))).. ( -27.76) >DroYak_CAF1 1554 105 - 1 AAAUUUUAUUGUCCGAAAAGUAAAAAAGGCUCGGCCUCCGACAGCAUUGCGAUUUAAUUACAUUAGCGACAGCUGGAGAAUGCACAAAAGUGGGUUGGCCAGCGG ...(((((((........)))))))...(((.((((..(..((((.((((...............))))..))))..)(((.(((....))).)))))))))).. ( -27.76) >consensus AAAUUUUAUUGUCCGAAAAGUAAAAAAGGCUCGGCCUCCGACAGCAUUGCGAUUUAAUUACAUUAGCGACAGCUGGAGAAUGCACAAAAGUGGGUUGGCCAGCGG ...(((((((........)))))))...(((.((((..(..((((.((((...............))))..))))..)(((.(((....))).)))))))))).. (-27.80 = -27.80 + 0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:54:56 2006