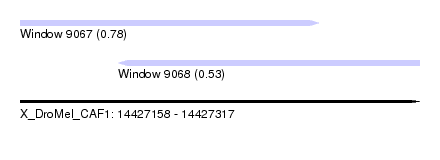
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 14,427,158 – 14,427,317 |
| Length | 159 |
| Max. P | 0.775364 |

| Location | 14,427,158 – 14,427,277 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 74.50 |
| Mean single sequence MFE | -24.27 |
| Consensus MFE | -12.38 |
| Energy contribution | -13.19 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.51 |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.775364 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 14427158 119 + 22224390 UUCUAGUUUAUGAAAAAUU-CGCAAAUCUCCAAAGAAUUAAAAAUGAUAAAUGAUUAUGGGCCAGCAGUGCGUAUACGUAAUAUAUAUCAAGUAUACGUAAUGUUGCCAGUAUGGAACAG (((((((((((.(..((((-(.............))))).....).))))))...(((.(((.((((.(((((((((..............))))))))).))))))).))))))))... ( -24.26) >DroSec_CAF1 6556 108 + 1 UUCUCGCUUAUGAAAAAUU-CGCAAAUCUCCAGAGAAAUAAAAAC--GAAAUUAUUGUGGGGUAGCAGUGCGUAUACGUGAUAUAUAUCAAGUAUACGCAACUUUACC---------CUG (((((..............-............)))))......((--((.....))))(((((((.(((((((((((.((((....)))).)))))))).))))))))---------)). ( -29.47) >DroSim_CAF1 6568 108 + 1 UUCUAGUUUAUGAAAAAAU-CGCAAAUCUGCAGAGAAAUAACAUA--UAAAUAAUUGUGGGCCAACAGUGCGUAUACGUAAUAUAUAUCAAGUAUACGUAACUUUACC---------CAG .....(((((((......(-((((....))).)).........))--))))).....((((.......(((((((((..............)))))))))......))---------)). ( -19.42) >DroYak_CAF1 6637 111 + 1 UUCUAGCA--AG-AAUCUCUCGAGAAUCUCCAGUGAAAUAAAC------AAUUAUUGUGACGCAACAGUGCGUAUACGUAAUGUACAUCAAGUAUACGUAACGUUGCCACUAUCAAACAA ........--((-(.(((....))).)))..((((......((------(.....)))...(((((..(((((((((..............)))))))))..)))))))))......... ( -23.94) >consensus UUCUAGCUUAUGAAAAAUU_CGCAAAUCUCCAGAGAAAUAAAAA___UAAAUUAUUGUGGGCCAACAGUGCGUAUACGUAAUAUAUAUCAAGUAUACGUAACGUUACC_________CAG ............................................................((((((..(((((((((..............)))))))))..))))))............ (-12.38 = -13.19 + 0.81)

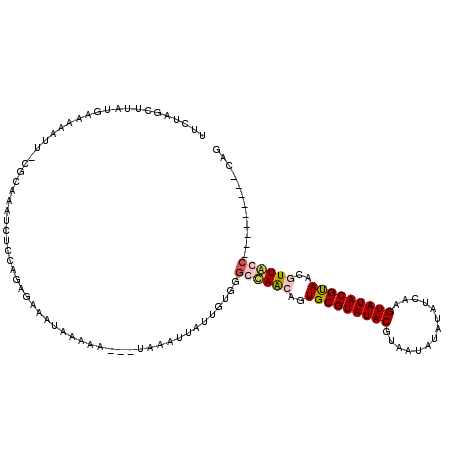
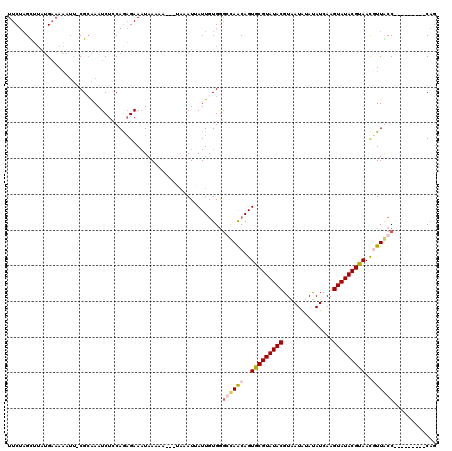
| Location | 14,427,197 – 14,427,317 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.23 |
| Mean single sequence MFE | -21.03 |
| Consensus MFE | -11.26 |
| Energy contribution | -11.26 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.80 |
| Structure conservation index | 0.54 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.526446 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
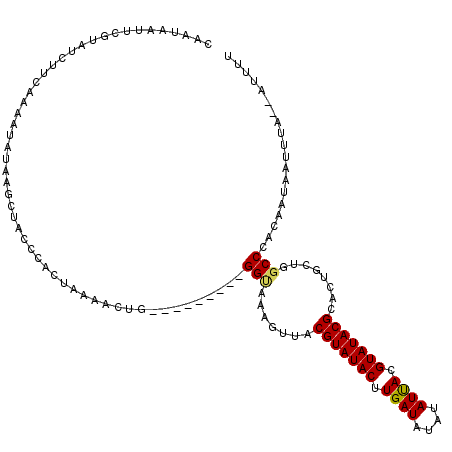
>X_DroMel_CAF1 14427197 120 - 22224390 CAAUAGUUCGUAUCUUCAAAAUAUUAGCUACCCACUAAAACUGUUCCAUACUGGCAACAUUACGUAUACUUGAUAUAUAUUACGUAUACGCACUGCUGGCCCAUAAUCAUUUAUCAUUUU ...(((((.((((.......)))).)))))....................(..(((......(((((((.((((....)))).)))))))...)))..)..................... ( -16.90) >DroSec_CAF1 6595 109 - 1 CAAUAAUUUGUAUCUUCAAAAUAUAAACUACCCACUAAAACAG---------GGUAAAGUUGCGUAUACUUGAUAUAUAUCACGUAUACGCACUGCUACCCCACAAUAAUUUC--GUUUU ..(((.((((......)))).)))..................(---------((((.(((.((((((((.((((....)))).)))))))))))..)))))............--..... ( -26.10) >DroSim_CAF1 6607 109 - 1 CAAUAAUUCGUAUCUUCAAAAUAUAAGCUACCCACUAAAACUG---------GGUAAAGUUACGUAUACUUGAUAUAUAUUACGUAUACGCACUGUUGGCCCACAAUUAUUUA--UAUGU .(((((((.(((.(((........))).)))..........((---------(((..(((..(((((((.((((....)))).))))))).)))....))))).)))))))..--..... ( -20.10) >consensus CAAUAAUUCGUAUCUUCAAAAUAUAAGCUACCCACUAAAACUG_________GGUAAAGUUACGUAUACUUGAUAUAUAUUACGUAUACGCACUGCUGGCCCACAAUAAUUUA__AUUUU ....................................................(((.......(((((((.((((....)))).)))))))........)))................... (-11.26 = -11.26 + 0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:51:34 2006