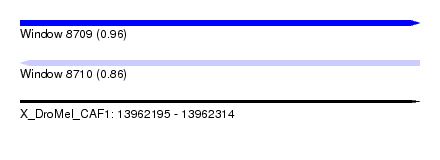

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,962,195 – 13,962,314 |
| Length | 119 |
| Max. P | 0.961113 |

| Location | 13,962,195 – 13,962,314 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.73 |
| Mean single sequence MFE | -45.30 |
| Consensus MFE | -34.79 |
| Energy contribution | -36.35 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.84 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.52 |
| SVM RNA-class probability | 0.961113 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13962195 119 + 22224390 AAAAAGCGCAGGCGGGUGGGCGCCGUUGGCCCCAACGCGAGAGUCAAGAACGUAAAGCAUAAUAAAAAGUCGUGCCUUACGUGCAGCAGUUGGAAAGCCCCCCCCCGAAUGCGCCC-ACC .....(((((..((((.(((((((...))).(((((((....))...(.((((((.(((..((.....))..))).)))))).)....)))))...))))...))))..)))))..-... ( -47.30) >DroSec_CAF1 13722 117 + 1 AAAAAGCGCAGGCGGGUGGGCGCCGUUGGCCCCAACGCGAGAGCCAAGAACGUAAAGCAUAAUAAAAAGUCGUGCCUUACGUGCAGCAGUUGGGAAGCCC---CCCGAAUGUACCCCCCC .......(((..((((.(((((((...)))((((((((....))...(.((((((.(((..((.....))..))).)))))).)....))))))..))))---))))..)))........ ( -47.70) >DroSim_CAF1 14178 116 + 1 AAAAAGCGCAGGCGGGUGGGCGCCGUUGGCCCCAACGCGAGAGCCAAGAACGUAAAGCAUAAUAAAAAGUCGUGCCUUACGUGCAGCAGUUGGGAAGCCC---CCCGAAUGUACCC-CCC .......(((..((((.(((((((...)))((((((((....))...(.((((((.(((..((.....))..))).)))))).)....))))))..))))---))))..)))....-... ( -47.70) >DroYak_CAF1 14093 114 + 1 AAUAAGCGCAGGCGGGUGGGCGCCGUUAGCCCCUUCGCGAGAGCCAAGAACGUAAAGCAUAAUAAAAAGUCUUGCCUUACGUGAAGCUUUUUGGAAUCCC---CCC---UCCACCGCCNN .....(((..((.(((.(((..(((..(((..(((.((....)).))).((((((.(((..((.....))..))).))))))...)))...)))...)))---)))---.))..)))... ( -38.50) >consensus AAAAAGCGCAGGCGGGUGGGCGCCGUUGGCCCCAACGCGAGAGCCAAGAACGUAAAGCAUAAUAAAAAGUCGUGCCUUACGUGCAGCAGUUGGGAAGCCC___CCCGAAUGCACCC_CCC .......(((..((((.((((((.....))((((((((....)).....((((((.(((..((.....))..))).))))))......))))))..))))...))))..)))........ (-34.79 = -36.35 + 1.56)

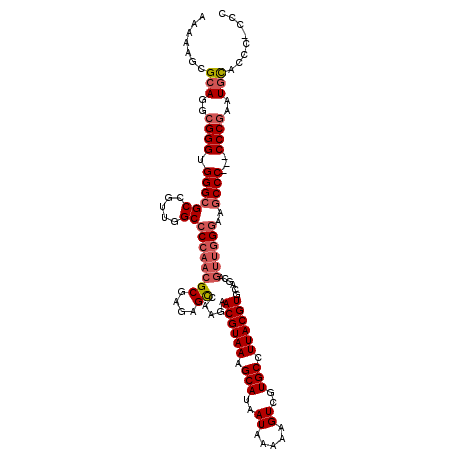
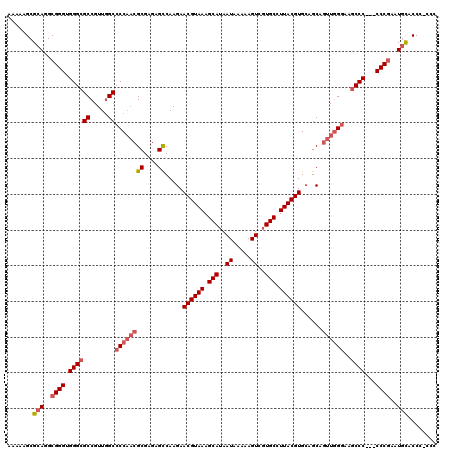
| Location | 13,962,195 – 13,962,314 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.73 |
| Mean single sequence MFE | -46.63 |
| Consensus MFE | -37.17 |
| Energy contribution | -38.30 |
| Covariance contribution | 1.13 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.81 |
| SVM RNA-class probability | 0.856732 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13962195 119 - 22224390 GGU-GGGCGCAUUCGGGGGGGGGCUUUCCAACUGCUGCACGUAAGGCACGACUUUUUAUUAUGCUUUACGUUCUUGACUCUCGCGUUGGGGCCAACGGCGCCCACCCGCCUGCGCUUUUU (((-((((((...(((((((.(((.........))).)((((((((((..(.......)..))))))))))......))))))(((((....)))))))))))))).((....))..... ( -50.60) >DroSec_CAF1 13722 117 - 1 GGGGGGGUACAUUCGGG---GGGCUUCCCAACUGCUGCACGUAAGGCACGACUUUUUAUUAUGCUUUACGUUCUUGGCUCUCGCGUUGGGGCCAACGGCGCCCACCCGCCUGCGCUUUUU .((((((..((..((((---((((..((((((.((.((((((((((((..(.......)..)))))))))).....))....))))))))(((...))))))).))))..))..)))))) ( -47.70) >DroSim_CAF1 14178 116 - 1 GGG-GGGUACAUUCGGG---GGGCUUCCCAACUGCUGCACGUAAGGCACGACUUUUUAUUAUGCUUUACGUUCUUGGCUCUCGCGUUGGGGCCAACGGCGCCCACCCGCCUGCGCUUUUU .((-(((..((..((((---((((..((((((.((.((((((((((((..(.......)..)))))))))).....))....))))))))(((...))))))).))))..))..))))). ( -47.30) >DroYak_CAF1 14093 114 - 1 NNGGCGGUGGA---GGG---GGGAUUCCAAAAAGCUUCACGUAAGGCAAGACUUUUUAUUAUGCUUUACGUUCUUGGCUCUCGCGAAGGGGCUAACGGCGCCCACCCGCCUGCGCUUAUU .....((((.(---((.---(((...........((((((((((((((.((.......)).)))))))))).....((....))))))((((.......)))).))).))).)))).... ( -40.90) >consensus GGG_GGGUACAUUCGGG___GGGCUUCCCAACUGCUGCACGUAAGGCACGACUUUUUAUUAUGCUUUACGUUCUUGGCUCUCGCGUUGGGGCCAACGGCGCCCACCCGCCUGCGCUUUUU .....((((((..((((...((((................((((((((..(.......)..))))))))((..((((((((......))))))))..)))))).))))..)))))).... (-37.17 = -38.30 + 1.13)

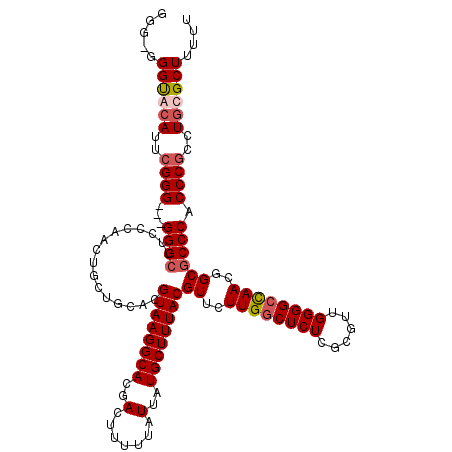
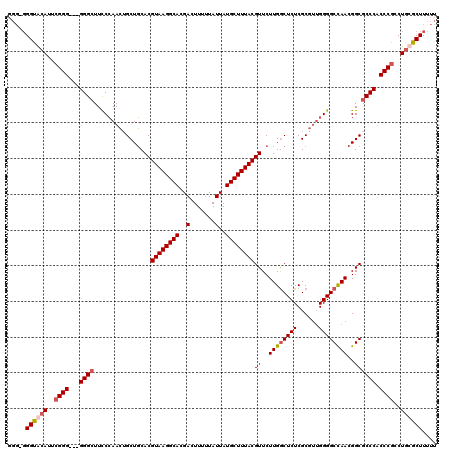
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:46:15 2006