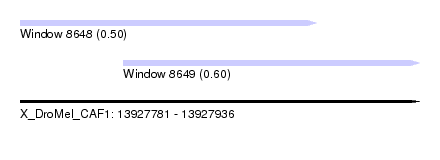
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,927,781 – 13,927,936 |
| Length | 155 |
| Max. P | 0.599747 |

| Location | 13,927,781 – 13,927,896 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 92.29 |
| Mean single sequence MFE | -26.76 |
| Consensus MFE | -21.35 |
| Energy contribution | -21.97 |
| Covariance contribution | 0.62 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.80 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13927781 115 + 22224390 AAACCCCACUGAUUGCCGUUUUUCCAGGCAUACCCACUCCAUACCCGGAAUCCCCUCUGCCAUACACACAUCUUGCAACGCAAAACGAGUGACAUUUUCAGAUGGUAUGGCCGGG .............((((.........))))..(((..(((......))).........(((((((.(((.((((((...))))...))))).((((....))))))))))).))) ( -27.80) >DroSec_CAF1 7950 114 + 1 AAACCCCACUGAUUGCCGUCUUUCCAGGCAUACCCACUCCAUACCGGGAAUCCCC-CUGCCAUACACACAUCUUACAACGCAAACCGAGUGACAUUUUCAGAUGGUAUGGCCGGG ..........(((((((.........)))...(((..........))))))).((-(.(((((((.(((.((..............))))).((((....))))))))))).))) ( -28.04) >DroSim_CAF1 10851 114 + 1 AAACCCCACUGAUUGCCGUCUUUCCAGGCAUACCCACUCCAUACCGGGAAUCCCC-CUGCCAUACACACAUCUUACAACGCAAAACGAGUGACAUUUUCAGAUGGUAUGGCCGGG ..........(((((((.........)))...(((..........))))))).((-(.(((((((.(((.((..............))))).((((....))))))))))).))) ( -28.04) >DroEre_CAF1 9688 110 + 1 AAACCCCACUGAU-GCCGUUUUUCCAGGCACACCCACUCCAUCC--AGAAUCCGC-CUGCCA-UCACACAUCUUACAACGCAAAGCGAGCGACAUUUUCAAAUGGUACGGCCGGG ...(((..(((.(-(((((((...(((((...............--.......))-)))...-...............(((.......)))........)))))))))))..))) ( -23.15) >consensus AAACCCCACUGAUUGCCGUCUUUCCAGGCAUACCCACUCCAUACCGGGAAUCCCC_CUGCCAUACACACAUCUUACAACGCAAAACGAGUGACAUUUUCAGAUGGUAUGGCCGGG ..............(((.........)))...(((..(((......))).........(((((((.............(((.......))).((((....))))))))))).))) (-21.35 = -21.97 + 0.62)

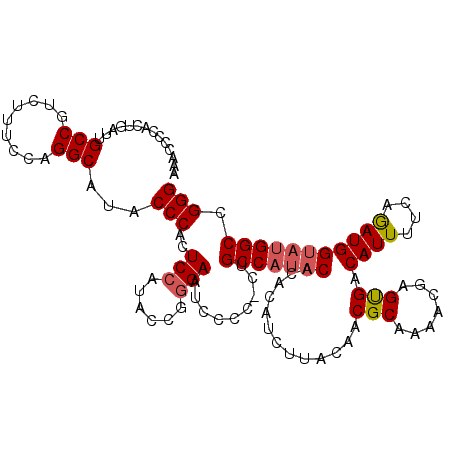
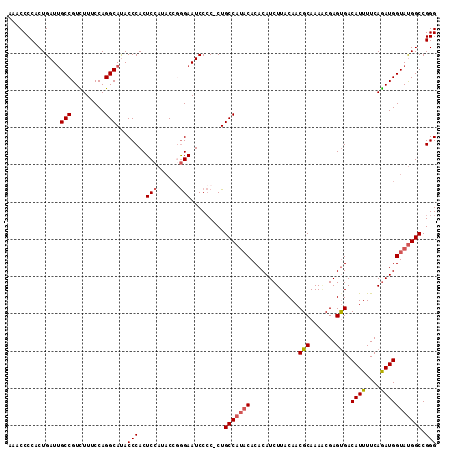
| Location | 13,927,821 – 13,927,936 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 84.57 |
| Mean single sequence MFE | -30.74 |
| Consensus MFE | -18.60 |
| Energy contribution | -21.47 |
| Covariance contribution | 2.87 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.599747 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
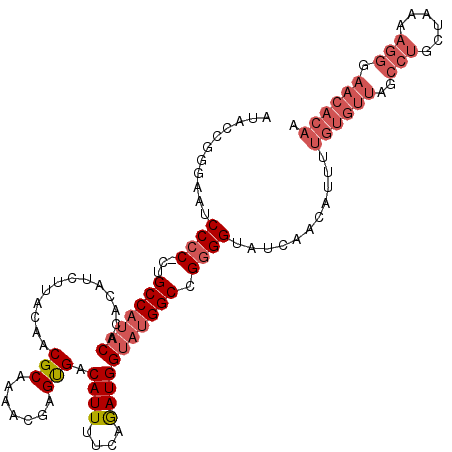
>X_DroMel_CAF1 13927821 115 + 22224390 AUACCCGGAAUCCCCUCUGCCAUACACACAUCUUGCAACGCAAAACGAGUGACAUUUUCAGAUGGUAUGGCCGGGGUAUCAACAUUUUGUGUUAGCCUGCUAAAAGGGAACACAA ((((((((.....)).(.(((((((.(((.((((((...))))...))))).((((....))))))))))).))))))).......(((((((..(((......))).))))))) ( -34.90) >DroSec_CAF1 7990 114 + 1 AUACCGGGAAUCCCC-CUGCCAUACACACAUCUUACAACGCAAACCGAGUGACAUUUUCAGAUGGUAUGGCCGGGGUAUCAACAUUUUGUGUUAUCCUGCUAAAAGGGAACACAA ((((((((...)))(-(.(((((((.(((.((..............))))).((((....))))))))))).))))))).......(((((((.((((......))))))))))) ( -34.54) >DroSim_CAF1 10891 114 + 1 AUACCGGGAAUCCCC-CUGCCAUACACACAUCUUACAACGCAAAACGAGUGACAUUUUCAGAUGGUAUGGCCGGGGUAUCAACAUUUUGUGUUAGCCUGCUAAAAGGGAACACAA ((((((((...)))(-(.(((((((.(((.((..............))))).((((....))))))))))).))))))).......(((((((..(((......))).))))))) ( -33.94) >DroEre_CAF1 9727 95 + 1 AUCC--AGAAUCCGC-CUGCCA-UCACACAUCUUACAACGCAAAGCGAGCGACAUUUUCAAAUGGUACGGCCGGGGUAUAAACAUUUUGGGCUAGACUG---------------- .(((--(((((((((-((((((-(..............(((...)))...((.....))..)))))).)).))))((....)).)))))))........---------------- ( -19.60) >consensus AUACCGGGAAUCCCC_CUGCCAUACACACAUCUUACAACGCAAAACGAGUGACAUUUUCAGAUGGUAUGGCCGGGGUAUCAACAUUUUGUGUUAGCCUGCUAAAAGGGAACACAA ...........((((...(((((((.............(((.......))).((((....))))))))))).))))...........((((((..(((......))).)))))). (-18.60 = -21.47 + 2.87)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:45:23 2006