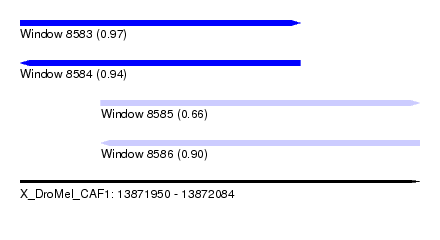
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,871,950 – 13,872,084 |
| Length | 134 |
| Max. P | 0.966666 |

| Location | 13,871,950 – 13,872,044 |
|---|---|
| Length | 94 |
| Sequences | 3 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 86.90 |
| Mean single sequence MFE | -33.13 |
| Consensus MFE | -26.37 |
| Energy contribution | -26.37 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.56 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.60 |
| SVM RNA-class probability | 0.966666 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13871950 94 + 22224390 AAAAUGCCAAACUCGCAUUGUUGUUGC---UGCAGUUGUUGUUUCGUGUUGCGUUCG-GCAGCGAGACAAACAAAAGCAACAACAACGUCACCAACGG ..(((((.......)))))((((((((---(....((((((((((((..(((.....-)))))))))).))))).)))))))))..(((.....))). ( -32.60) >DroSec_CAF1 17371 95 + 1 NNNNNNNNNNNNNNGCAUUGUUGUUGC---UGCAGUUGUUGUUUCGUGUUGCGUUCGCGCAGCGAGACAAACAAAAGCAACAACAACGUCACCAACGG .................((((((((((---(....((((((((((((..((((....))))))))))).))))).)))))))))))(((.....))). ( -34.20) >DroEre_CAF1 15154 97 + 1 AAAAUGCCAAACUCGCAUUGUUGUUGCUGCUGCUGUUGUUGUUUCGUGUUGCGUUCG-GCAGCGAGACAAACAAAAGCAACAACAACGUCACCAACGG ..(((((.......)))))(((((((.((((..((((..((((((((..(((.....-)))))))))))))))..)))).)))))))........... ( -32.60) >consensus AAAAUGCCAAACUCGCAUUGUUGUUGC___UGCAGUUGUUGUUUCGUGUUGCGUUCG_GCAGCGAGACAAACAAAAGCAACAACAACGUCACCAACGG .................((((((((((........((((((((((((..(((......)))))))))).)))))..))))))))))(((.....))). (-26.37 = -26.37 + -0.00)

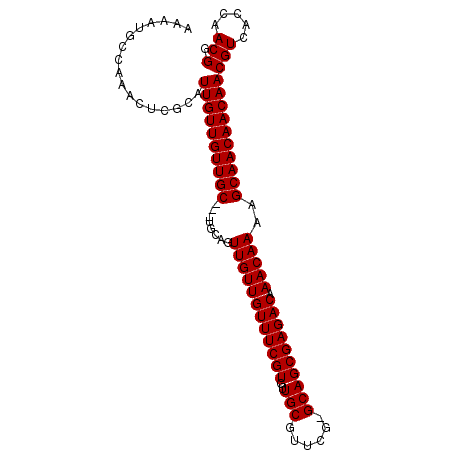
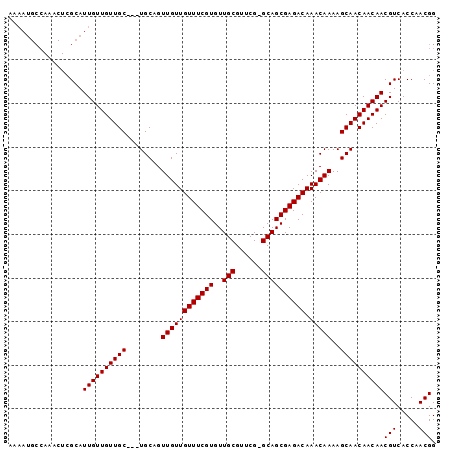
| Location | 13,871,950 – 13,872,044 |
|---|---|
| Length | 94 |
| Sequences | 3 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 86.90 |
| Mean single sequence MFE | -32.50 |
| Consensus MFE | -23.80 |
| Energy contribution | -23.80 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.39 |
| Structure conservation index | 0.73 |
| SVM decision value | 1.27 |
| SVM RNA-class probability | 0.938757 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13871950 94 - 22224390 CCGUUGGUGACGUUGUUGUUGCUUUUGUUUGUCUCGCUGC-CGAACGCAACACGAAACAACAACUGCA---GCAACAACAAUGCGAGUUUGGCAUUUU .....(....)(((((((((((..(((((.((.(((.(((-.....)))...))).)))))))..)))---))))))))(((((.......))))).. ( -31.80) >DroSec_CAF1 17371 95 - 1 CCGUUGGUGACGUUGUUGUUGCUUUUGUUUGUCUCGCUGCGCGAACGCAACACGAAACAACAACUGCA---GCAACAACAAUGCNNNNNNNNNNNNNN .(((((....)(((((((((((..(((((.((.(((.((((....))))...))).)))))))..)))---))))))))))))............... ( -31.60) >DroEre_CAF1 15154 97 - 1 CCGUUGGUGACGUUGUUGUUGCUUUUGUUUGUCUCGCUGC-CGAACGCAACACGAAACAACAACAGCAGCAGCAACAACAAUGCGAGUUUGGCAUUUU ..((((....)((((((((((((.(((((.((.(((.(((-.....)))...))).))))))).)))))))))))))))(((((.......))))).. ( -34.10) >consensus CCGUUGGUGACGUUGUUGUUGCUUUUGUUUGUCUCGCUGC_CGAACGCAACACGAAACAACAACUGCA___GCAACAACAAUGCGAGUUUGGCAUUUU .(((((....)(((((((((((..(((((.((.(((.(((......)))...))).)))))))..)))...))))))))))))............... (-23.80 = -23.80 + 0.00)
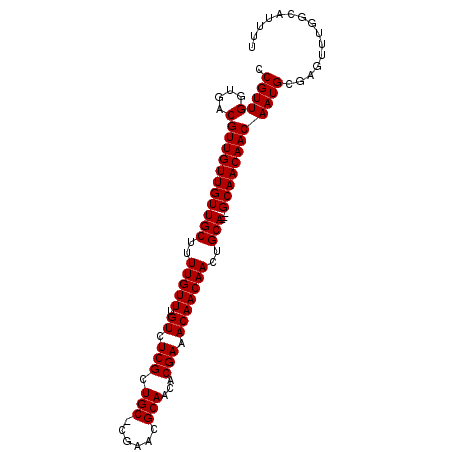


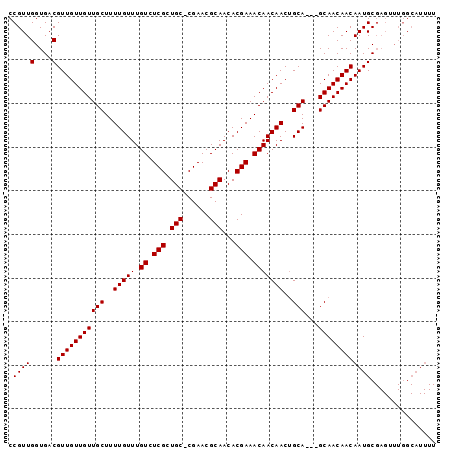
| Location | 13,871,977 – 13,872,084 |
|---|---|
| Length | 107 |
| Sequences | 3 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 97.52 |
| Mean single sequence MFE | -29.34 |
| Consensus MFE | -27.00 |
| Energy contribution | -27.00 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.661063 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
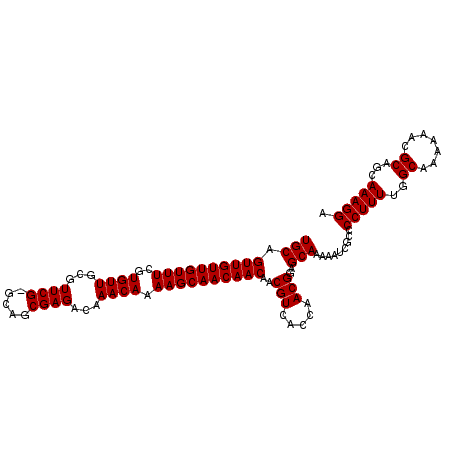
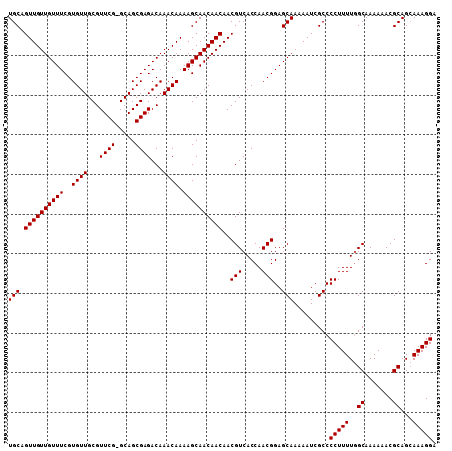
>X_DroMel_CAF1 13871977 107 + 22224390 UGCAGUUGUUGUUUCGUGUUGCGUUCG-GCAGCGAGACAAACAAAAGCAACAACAACGUCACCAACGGAGCAAAAAUCGCCCCUUUUAGCAAAAAACGCAGGAAAGGA (((.((((((((((..((((...((((-....))))...)))).))))))))))..(((.....)))..))).........((((((.((.......))..)))))). ( -29.00) >DroSec_CAF1 17398 108 + 1 UGCAGUUGUUGUUUCGUGUUGCGUUCGCGCAGCGAGACAAACAAAAGCAACAACAACGUCACCAACGGAGCAAAAAUCGCCCCUUUUGGCAAAAAACGCAGCAAAGGA (((.((((((((((..((((...(((((...)))))...)))).))))))))))..(((.....)))..))).........(((((..((.......))...))))). ( -30.10) >DroEre_CAF1 15184 107 + 1 UGCUGUUGUUGUUUCGUGUUGCGUUCG-GCAGCGAGACAAACAAAAGCAACAACAACGUCACCAACGGAGCAAAAAUCGCCCCUUUUGGCAAAAAACGCAGCAAAGGA ((((((..((((((((((((((.....-)))))..(((...................)))....))))))))).....(((......))).......))))))..... ( -28.91) >consensus UGCAGUUGUUGUUUCGUGUUGCGUUCG_GCAGCGAGACAAACAAAAGCAACAACAACGUCACCAACGGAGCAAAAAUCGCCCCUUUUGGCAAAAAACGCAGCAAAGGA (((.((((((((((..((((...((((.....))))...)))).))))))))))..(((.....)))..))).........(((((..((.......))...))))). (-27.00 = -27.00 + 0.00)

| Location | 13,871,977 – 13,872,084 |
|---|---|
| Length | 107 |
| Sequences | 3 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 97.52 |
| Mean single sequence MFE | -34.30 |
| Consensus MFE | -32.65 |
| Energy contribution | -32.43 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.95 |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.898881 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
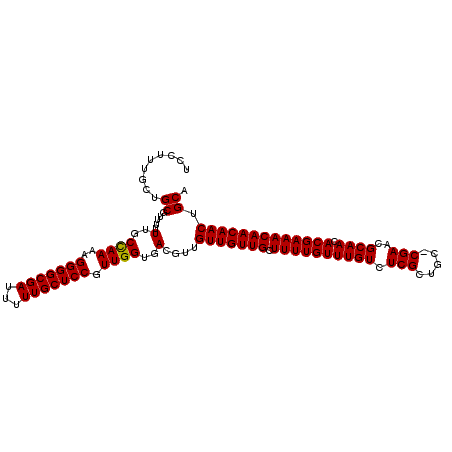
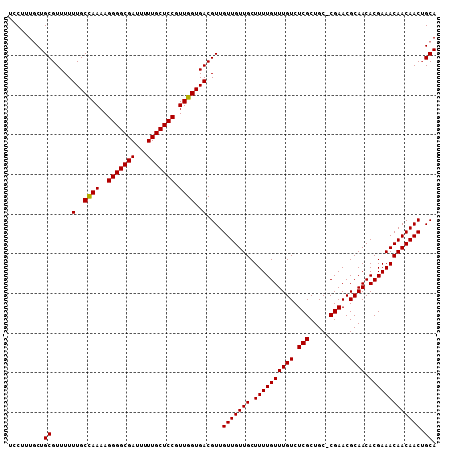
>X_DroMel_CAF1 13871977 107 - 22224390 UCCUUUCCUGCGUUUUUUGCUAAAAGGGGCGAUUUUUGCUCCGUUGGUGACGUUGUUGUUGCUUUUGUUUGUCUCGCUGC-CGAACGCAACACGAAACAACAACUGCA ................(..((((..(((((((...))))))).))))..).((.(((((((.((((((((((.(((....-)))..)))).))))))))))))).)). ( -30.10) >DroSec_CAF1 17398 108 - 1 UCCUUUGCUGCGUUUUUUGCCAAAAGGGGCGAUUUUUGCUCCGUUGGUGACGUUGUUGUUGCUUUUGUUUGUCUCGCUGCGCGAACGCAACACGAAACAACAACUGCA .....(((.(((((....(((((..(((((((...))))))).)))))))))).(((((((.((((((((((.((((...))))..)))).))))))))))))).))) ( -36.60) >DroEre_CAF1 15184 107 - 1 UCCUUUGCUGCGUUUUUUGCCAAAAGGGGCGAUUUUUGCUCCGUUGGUGACGUUGUUGUUGCUUUUGUUUGUCUCGCUGC-CGAACGCAACACGAAACAACAACAGCA ................(..((((..(((((((...))))))).))))..).((((((((((.((((((((((.(((....-)))..)))).)))))))))))))))). ( -36.20) >consensus UCCUUUGCUGCGUUUUUUGCCAAAAGGGGCGAUUUUUGCUCCGUUGGUGACGUUGUUGUUGCUUUUGUUUGUCUCGCUGC_CGAACGCAACACGAAACAACAACUGCA .........((.....(..((((..(((((((...))))))).))))..)....(((((((.((((((((((.(((.....)))..)))).))))))))))))).)). (-32.65 = -32.43 + -0.22)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:44:26 2006