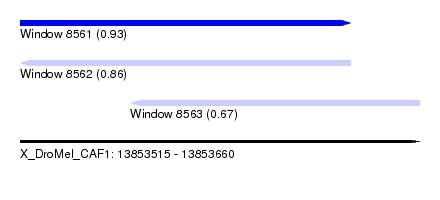
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,853,515 – 13,853,660 |
| Length | 145 |
| Max. P | 0.931947 |

| Location | 13,853,515 – 13,853,635 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 99.50 |
| Mean single sequence MFE | -35.00 |
| Consensus MFE | -35.00 |
| Energy contribution | -35.00 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.56 |
| Structure conservation index | 1.00 |
| SVM decision value | 1.22 |
| SVM RNA-class probability | 0.931947 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 13853515 120 + 22224390 CUGCGUUACGCUCGUCACAUUUUAUUAGCCAGUUUGCAGGCUGCAGGACAUUGAACGUCUGUCUCCCUGCGCGAGUCCUUUGUGUCGCAAUAAAUAACACACACAUCAGUGUGUUAGUGU .((((.(((((((((...........((((........))))((((((((.........)))...))))))))))).....))).)))).....((((((((......)))))))).... ( -35.00) >DroSec_CAF1 1753 120 + 1 CUGCGUUACGCUCGUCACAUUUUAUUAGCCAGUUUGCAGGCUGCAGGACAUUGAACGUCUGUCUCCCUGCGCGAGUCCUUUGUGUCGCAAUAAAUAACACACACAUCAGUGUGUUAAUGU .((((.(((((((((...........((((........))))((((((((.........)))...))))))))))).....))).)))).....((((((((......)))))))).... ( -35.00) >DroSim_CAF1 6384 120 + 1 CUGCGUUACGCUCGUCACAUUUUAUUAGCCAGUUUGCAGGCUGCAGGACAUUGAACGUCUGUCUCCCUGCGCGAGUCCUUUGUGUCGCAAUAAAUAACACACACAUCAGUGUGUUAAUGU .((((.(((((((((...........((((........))))((((((((.........)))...))))))))))).....))).)))).....((((((((......)))))))).... ( -35.00) >DroEre_CAF1 1708 120 + 1 CUGCGUUACGCUCGUCACAUUUUAUUAGCCAGUUUGCAGGCUGCAGGACAUUGAACGUCUGUCUCCCUGCGCGAGUCCUUUGUGUCGCAAUAAAUAACACACACAUCAGUGUGUUAGUGU .((((.(((((((((...........((((........))))((((((((.........)))...))))))))))).....))).)))).....((((((((......)))))))).... ( -35.00) >DroYak_CAF1 3749 120 + 1 CUGCGUUACGCUCGUCACAUUUUAUUAGCCAGUUUGCAGGCUGCAGGACAUUGAACGUCUGUCUCCCUGCGCGAGUCCUUUGUGUCGCAAUAAAUAACACACACAUCAGUGUGUUAGUGU .((((.(((((((((...........((((........))))((((((((.........)))...))))))))))).....))).)))).....((((((((......)))))))).... ( -35.00) >consensus CUGCGUUACGCUCGUCACAUUUUAUUAGCCAGUUUGCAGGCUGCAGGACAUUGAACGUCUGUCUCCCUGCGCGAGUCCUUUGUGUCGCAAUAAAUAACACACACAUCAGUGUGUUAGUGU .((((.(((((((((...........((((........))))((((((((.........)))...))))))))))).....))).)))).....((((((((......)))))))).... (-35.00 = -35.00 + 0.00)

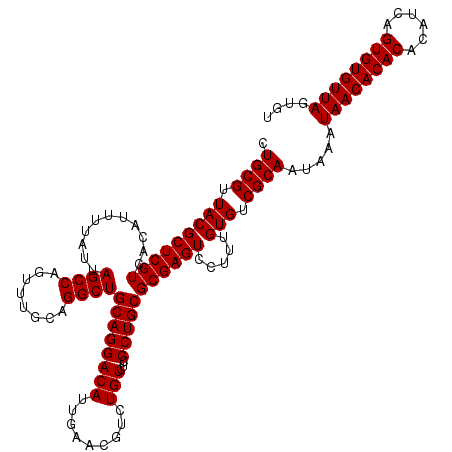
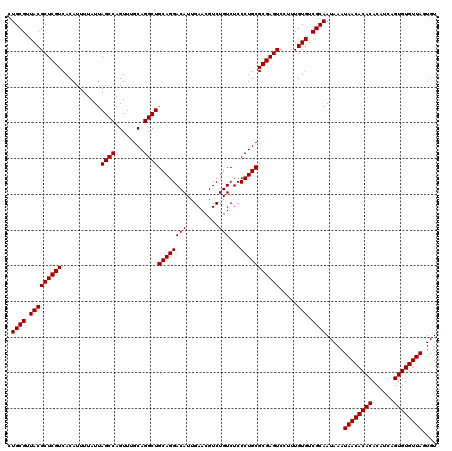
| Location | 13,853,515 – 13,853,635 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 99.50 |
| Mean single sequence MFE | -34.92 |
| Consensus MFE | -34.74 |
| Energy contribution | -34.74 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.99 |
| SVM decision value | 0.84 |
| SVM RNA-class probability | 0.864565 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
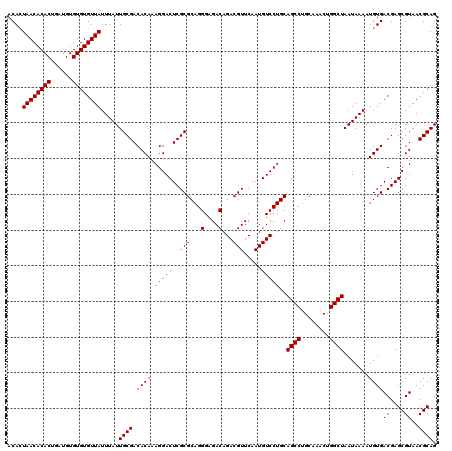
>X_DroMel_CAF1 13853515 120 - 22224390 ACACUAACACACUGAUGUGUGUGUUAUUUAUUGCGACACAAAGGACUCGCGCAGGGAGACAGACGUUCAAUGUCCUGCAGCCUGCAAACUGGCUAAUAAAAUGUGACGAGCGUAACGCAG .(((((((((((......))))))))((((((((((..........))))(((((....).(((((...)))))))))((((........))))))))))..)))....(((...))).. ( -35.00) >DroSec_CAF1 1753 120 - 1 ACAUUAACACACUGAUGUGUGUGUUAUUUAUUGCGACACAAAGGACUCGCGCAGGGAGACAGACGUUCAAUGUCCUGCAGCCUGCAAACUGGCUAAUAAAAUGUGACGAGCGUAACGCAG ....((((((((......)))))))).....((((.((((.(((((..(((...(....)...))).....)))))..((((........)))).......))))((....))..)))). ( -34.80) >DroSim_CAF1 6384 120 - 1 ACAUUAACACACUGAUGUGUGUGUUAUUUAUUGCGACACAAAGGACUCGCGCAGGGAGACAGACGUUCAAUGUCCUGCAGCCUGCAAACUGGCUAAUAAAAUGUGACGAGCGUAACGCAG ....((((((((......)))))))).....((((.((((.(((((..(((...(....)...))).....)))))..((((........)))).......))))((....))..)))). ( -34.80) >DroEre_CAF1 1708 120 - 1 ACACUAACACACUGAUGUGUGUGUUAUUUAUUGCGACACAAAGGACUCGCGCAGGGAGACAGACGUUCAAUGUCCUGCAGCCUGCAAACUGGCUAAUAAAAUGUGACGAGCGUAACGCAG .(((((((((((......))))))))((((((((((..........))))(((((....).(((((...)))))))))((((........))))))))))..)))....(((...))).. ( -35.00) >DroYak_CAF1 3749 120 - 1 ACACUAACACACUGAUGUGUGUGUUAUUUAUUGCGACACAAAGGACUCGCGCAGGGAGACAGACGUUCAAUGUCCUGCAGCCUGCAAACUGGCUAAUAAAAUGUGACGAGCGUAACGCAG .(((((((((((......))))))))((((((((((..........))))(((((....).(((((...)))))))))((((........))))))))))..)))....(((...))).. ( -35.00) >consensus ACACUAACACACUGAUGUGUGUGUUAUUUAUUGCGACACAAAGGACUCGCGCAGGGAGACAGACGUUCAAUGUCCUGCAGCCUGCAAACUGGCUAAUAAAAUGUGACGAGCGUAACGCAG ....((((((((......)))))))).....((((.((((.(((((..(((...(....)...))).....)))))..((((........)))).......))))((....))..)))). (-34.74 = -34.74 + -0.00)

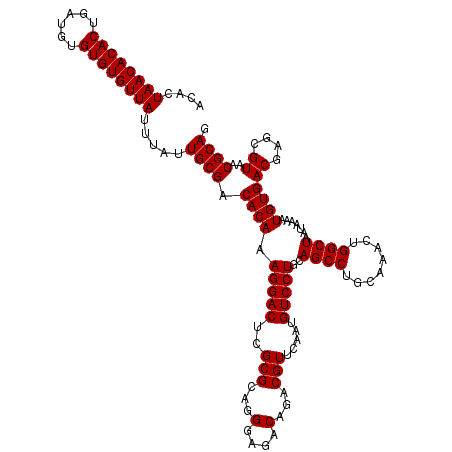
| Location | 13,853,555 – 13,853,660 |
|---|---|
| Length | 105 |
| Sequences | 3 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 95.61 |
| Mean single sequence MFE | -27.73 |
| Consensus MFE | -23.73 |
| Energy contribution | -23.73 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.19 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.671478 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
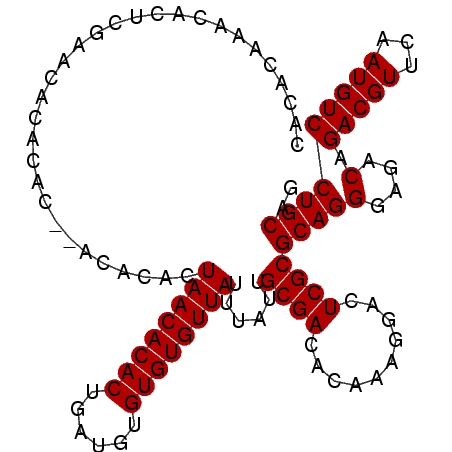
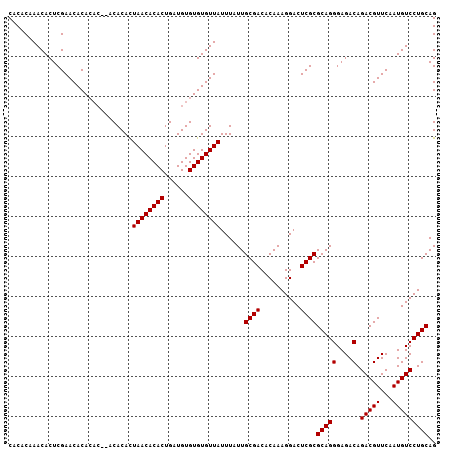
>X_DroMel_CAF1 13853555 105 - 22224390 CACACAAACACUCGAACACACAC--ACACACUAACACACUGAUGUGUGUGUUAUUUAUUGCGACACAAAGGACUCGCGCAGGGAGACAGACGUUCAAUGUCCUGCAG ..............(((((((((--(((...........)).)))))))))).......((((..........))))(((((....).(((((...))))))))).. ( -26.70) >DroSim_CAF1 6424 107 - 1 CACACAAACACUCGAACACACACACACACAUUAACACACUGAUGUGUGUGUUAUUUAUUGCGACACAAAGGACUCGCGCAGGGAGACAGACGUUCAAUGUCCUGCAG .....................((((((((((((......))))))))))))........((((..........))))(((((....).(((((...))))))))).. ( -29.50) >DroEre_CAF1 1748 101 - 1 CACACAAACACUCGAACACA------CACACUAACACACUGAUGUGUGUGUUAUUUAUUGCGACACAAAGGACUCGCGCAGGGAGACAGACGUUCAAUGUCCUGCAG ..............((((((------((((.((......)).)))))))))).......((((..........))))(((((....).(((((...))))))))).. ( -27.00) >consensus CACACAAACACUCGAACACACAC__ACACACUAACACACUGAUGUGUGUGUUAUUUAUUGCGACACAAAGGACUCGCGCAGGGAGACAGACGUUCAAUGUCCUGCAG ...............................((((((((......))))))))......((((..........))))(((((....).(((((...))))))))).. (-23.73 = -23.73 + -0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:44:06 2006