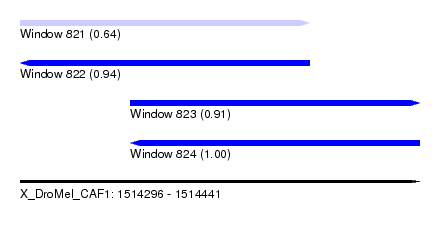

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,514,296 – 1,514,441 |
| Length | 145 |
| Max. P | 0.995771 |

| Location | 1,514,296 – 1,514,401 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.28 |
| Mean single sequence MFE | -21.72 |
| Consensus MFE | -16.98 |
| Energy contribution | -16.98 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.639650 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1514296 105 + 22224390 UCGACAAAAUUAACCGCUUCCGUUUUCAAUCCGACCCGUUUCCGGUUCGUUCUCUUCAGA-------------AAGCAUAAAAUAA--AACACGAAGAGCACAACGAACCGGAAACCACG .....................................((((((((((((((((((((...-------------.............--.....))))))...)))))))))))))).... ( -27.90) >DroSec_CAF1 3780 105 + 1 UCGACAAAAUUAACCGCUUCCGUUUUCAAUCCGACCCGUUUCCGAUUCGUUCUCUGCAGA-------------AAGCAUAAAAUAA--AACACGAAGAGCACAACGAACCGGAAACCACG .....................................(((((((.((((((((((.....-------------.............--.......))))...)))))).))))))).... ( -17.71) >DroSim_CAF1 5002 105 + 1 UCGACAAAAUUAACCGCUUCCGUUUUCAAUCCGACCCGUUUCCGAUUCGUUCUCUGCAGA-------------AAGCAGAAAAUAA--AACACGAAGAGCACAACGAACCGGAAACCACG .....................................(((((((.((((((.(((((...-------------..)))))......--.....(.....)..)))))).))))))).... ( -20.20) >DroEre_CAF1 3805 105 + 1 ACGACAAAAUUAACCGCUUCCGUUUUCAAUCCGACCCGUUUCCGAUUCGUUCUCUGAAGA-------------AAGCAUAUAAUAA--AGAAAUAAGAGCACAACGAACCGGAAACCACG .....................................(((((((.((((((((((.....-------------.............--.......))))...)))))).))))))).... ( -17.71) >DroYak_CAF1 5328 120 + 1 UCGACAAAAUUAACAGCUUCCGUUUUCAAUCCGACCCGUUUCCGAUUCGUUCUCUGGAGAAUAUUCUGCAUUAAAGCAGAAAUUAAAUAGAAAGAAGAGCACAACGAACCGGAAACCACG (((........(((.......))).......)))...(((((((.((((((((((......((((((((......)))))......)))......))))...)))))).))))))).... ( -25.06) >consensus UCGACAAAAUUAACCGCUUCCGUUUUCAAUCCGACCCGUUUCCGAUUCGUUCUCUGCAGA_____________AAGCAUAAAAUAA__AACACGAAGAGCACAACGAACCGGAAACCACG .....................................(((((((.((((((((((........................................))))...)))))).))))))).... (-16.98 = -16.98 + 0.00)

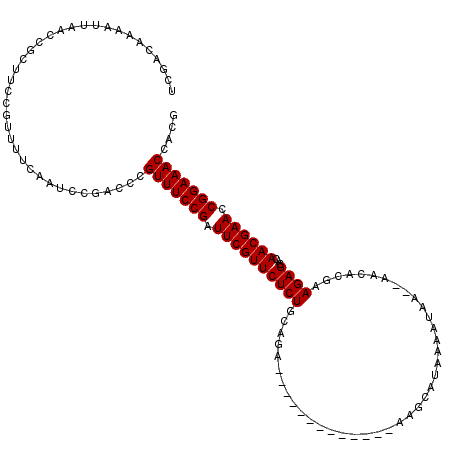
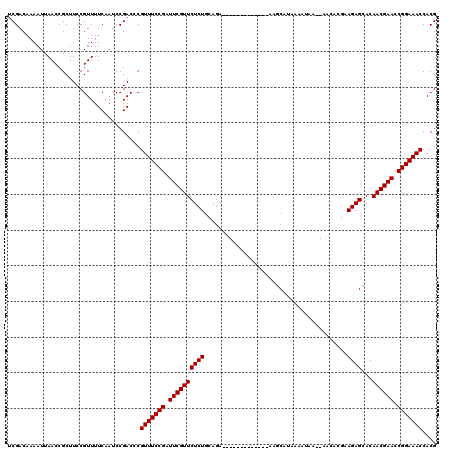
| Location | 1,514,296 – 1,514,401 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.28 |
| Mean single sequence MFE | -32.64 |
| Consensus MFE | -27.22 |
| Energy contribution | -27.06 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.83 |
| SVM decision value | 1.27 |
| SVM RNA-class probability | 0.938230 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1514296 105 - 22224390 CGUGGUUUCCGGUUCGUUGUGCUCUUCGUGUU--UUAUUUUAUGCUU-------------UCUGAAGAGAACGAACCGGAAACGGGUCGGAUUGAAAACGGAAGCGGUUAAUUUUGUCGA ....((((((((((((((...(((((((.((.--.........))..-------------..))))))))))))))))))))).((..(((((((...((....)).)))))))..)).. ( -35.70) >DroSec_CAF1 3780 105 - 1 CGUGGUUUCCGGUUCGUUGUGCUCUUCGUGUU--UUAUUUUAUGCUU-------------UCUGCAGAGAACGAAUCGGAAACGGGUCGGAUUGAAAACGGAAGCGGUUAAUUUUGUCGA ....((((((((((((((.(((.....((((.--.......))))..-------------...)))...)))))))))))))).((..(((((((...((....)).)))))))..)).. ( -30.40) >DroSim_CAF1 5002 105 - 1 CGUGGUUUCCGGUUCGUUGUGCUCUUCGUGUU--UUAUUUUCUGCUU-------------UCUGCAGAGAACGAAUCGGAAACGGGUCGGAUUGAAAACGGAAGCGGUUAAUUUUGUCGA ....(((((((((((((((..(.....)..).--.....((((((..-------------...)))))))))))))))))))).((..(((((((...((....)).)))))))..)).. ( -34.60) >DroEre_CAF1 3805 105 - 1 CGUGGUUUCCGGUUCGUUGUGCUCUUAUUUCU--UUAUUAUAUGCUU-------------UCUUCAGAGAACGAAUCGGAAACGGGUCGGAUUGAAAACGGAAGCGGUUAAUUUUGUCGU ....((((((((((((((...((((.......--.............-------------.....)))))))))))))))))).((..(((((((...((....)).)))))))..)).. ( -27.51) >DroYak_CAF1 5328 120 - 1 CGUGGUUUCCGGUUCGUUGUGCUCUUCUUUCUAUUUAAUUUCUGCUUUAAUGCAGAAUAUUCUCCAGAGAACGAAUCGGAAACGGGUCGGAUUGAAAACGGAAGCUGUUAAUUUUGUCGA ....((((((((((((((.....................((((((......))))))..(((....))))))))))))))))).((..(((((((...(....)...)))))))..)).. ( -35.00) >consensus CGUGGUUUCCGGUUCGUUGUGCUCUUCGUGUU__UUAUUUUAUGCUU_____________UCUGCAGAGAACGAAUCGGAAACGGGUCGGAUUGAAAACGGAAGCGGUUAAUUUUGUCGA ....((((((((((((((...((((........................................)))))))))))))))))).((..(((((((...(....)...)))))))..)).. (-27.22 = -27.06 + -0.16)

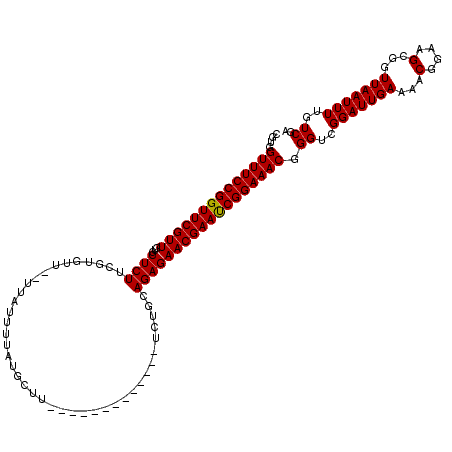
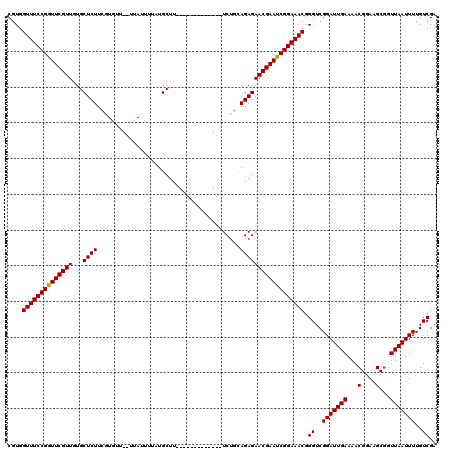
| Location | 1,514,336 – 1,514,441 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.92 |
| Mean single sequence MFE | -25.13 |
| Consensus MFE | -18.84 |
| Energy contribution | -18.52 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.75 |
| SVM decision value | 1.10 |
| SVM RNA-class probability | 0.914829 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1514336 105 + 22224390 UCCGGUUCGUUCUCUUCAGA-------------AAGCAUAAAAUAA--AACACGAAGAGCACAACGAACCGGAAACCACGAUCGGAAUAGCCAACGUAGUGGCAAAACCUAAAUCUGAAC (((((((((((((((((...-------------.............--.....))))))...)))))))))))..((......))....((((......))))................. ( -29.70) >DroSec_CAF1 3820 105 + 1 UCCGAUUCGUUCUCUGCAGA-------------AAGCAUAAAAUAA--AACACGAAGAGCACAACGAACCGGAAACCACGAUCGGAAUAGCCAACGUAGUGGCAAAACCUAAAUCUGAAC ((((.((((((((((.....-------------.............--.......))))...)))))).))))..((......))....((((......))))................. ( -19.51) >DroSim_CAF1 5042 105 + 1 UCCGAUUCGUUCUCUGCAGA-------------AAGCAGAAAAUAA--AACACGAAGAGCACAACGAACCGGAAACCACGAUCGGAAUAGCCAACGUAGUGGCAAAACCUAAAUCUGAAC ((((((.(((..(((((...-------------..)))))......--......................(....).)))))))))...((((......))))................. ( -23.20) >DroEre_CAF1 3845 105 + 1 UCCGAUUCGUUCUCUGAAGA-------------AAGCAUAUAAUAA--AGAAAUAAGAGCACAACGAACCGGAAACCACGAUCGGAGUAGCCAACGUAGUGGCAAAACCUAAAUCUGAAC ((((.((((((((((.....-------------.............--.......))))...)))))).))))........(((((...((((......))))..........))))).. ( -20.13) >DroYak_CAF1 5368 120 + 1 UCCGAUUCGUUCUCUGGAGAAUAUUCUGCAUUAAAGCAGAAAUUAAAUAGAAAGAAGAGCACAACGAACCGGAAACCACGGUCGGAAUAGCCGACGUAGUGGCAAAACCUAAAUCUGAAC ((((.((((((((((......((((((((......)))))......)))......))))...)))))).))))..((((.(((((.....)))))...)))).................. ( -33.10) >consensus UCCGAUUCGUUCUCUGCAGA_____________AAGCAUAAAAUAA__AACACGAAGAGCACAACGAACCGGAAACCACGAUCGGAAUAGCCAACGUAGUGGCAAAACCUAAAUCUGAAC ((((.((((((((((........................................))))...)))))).))))...((.(((.((....((((......))))....))...)))))... (-18.84 = -18.52 + -0.32)

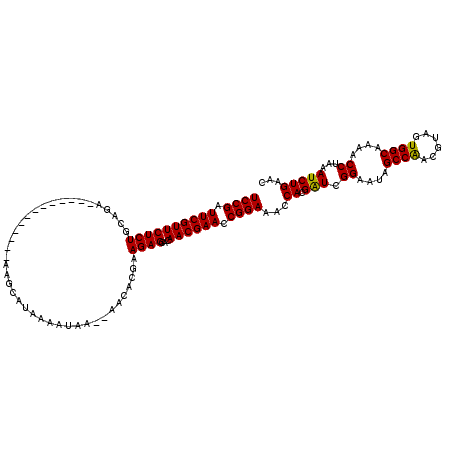
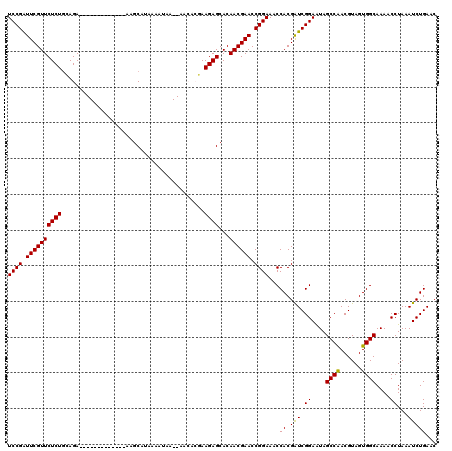
| Location | 1,514,336 – 1,514,441 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.92 |
| Mean single sequence MFE | -31.56 |
| Consensus MFE | -26.70 |
| Energy contribution | -26.38 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.46 |
| Structure conservation index | 0.85 |
| SVM decision value | 2.61 |
| SVM RNA-class probability | 0.995771 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1514336 105 - 22224390 GUUCAGAUUUAGGUUUUGCCACUACGUUGGCUAUUCCGAUCGUGGUUUCCGGUUCGUUGUGCUCUUCGUGUU--UUAUUUUAUGCUU-------------UCUGAAGAGAACGAACCGGA ...........((.....))((((((((((.....)))).)))))).(((((((((((...(((((((.((.--.........))..-------------..)))))))))))))))))) ( -34.10) >DroSec_CAF1 3820 105 - 1 GUUCAGAUUUAGGUUUUGCCACUACGUUGGCUAUUCCGAUCGUGGUUUCCGGUUCGUUGUGCUCUUCGUGUU--UUAUUUUAUGCUU-------------UCUGCAGAGAACGAAUCGGA ...........((.....))((((((((((.....)))).)))))).(((((((((((.(((.....((((.--.......))))..-------------...)))...))))))))))) ( -28.80) >DroSim_CAF1 5042 105 - 1 GUUCAGAUUUAGGUUUUGCCACUACGUUGGCUAUUCCGAUCGUGGUUUCCGGUUCGUUGUGCUCUUCGUGUU--UUAUUUUCUGCUU-------------UCUGCAGAGAACGAAUCGGA ...........((.....))((((((((((.....)))).)))))).((((((((((((..(.....)..).--.....((((((..-------------...))))))))))))))))) ( -33.00) >DroEre_CAF1 3845 105 - 1 GUUCAGAUUUAGGUUUUGCCACUACGUUGGCUACUCCGAUCGUGGUUUCCGGUUCGUUGUGCUCUUAUUUCU--UUAUUAUAUGCUU-------------UCUUCAGAGAACGAAUCGGA ...........((.....))((((((((((.....)))).)))))).(((((((((((...((((.......--.............-------------.....))))))))))))))) ( -25.91) >DroYak_CAF1 5368 120 - 1 GUUCAGAUUUAGGUUUUGCCACUACGUCGGCUAUUCCGACCGUGGUUUCCGGUUCGUUGUGCUCUUCUUUCUAUUUAAUUUCUGCUUUAAUGCAGAAUAUUCUCCAGAGAACGAAUCGGA ...........((.....))((((((((((.....)))).)))))).(((((((((((.....................((((((......))))))..(((....)))))))))))))) ( -36.00) >consensus GUUCAGAUUUAGGUUUUGCCACUACGUUGGCUAUUCCGAUCGUGGUUUCCGGUUCGUUGUGCUCUUCGUGUU__UUAUUUUAUGCUU_____________UCUGCAGAGAACGAAUCGGA ...........((.....))((((((((((.....)))).)))))).(((((((((((...((((........................................))))))))))))))) (-26.70 = -26.38 + -0.32)

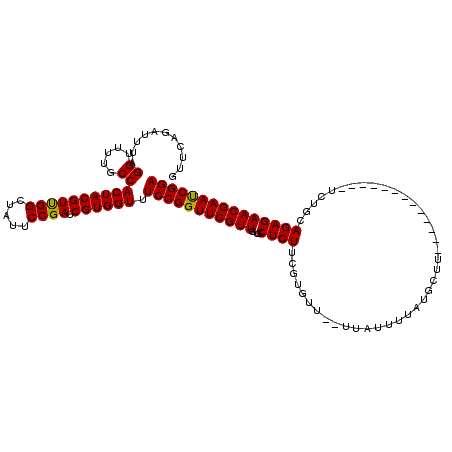
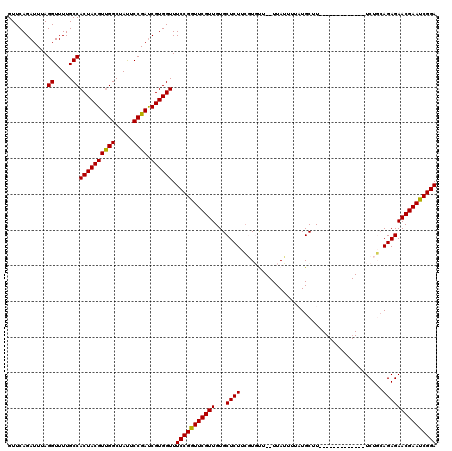
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:46:51 2006