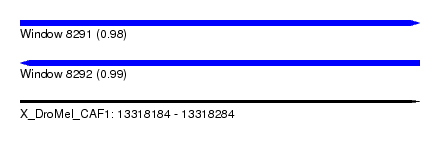
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 13,318,184 – 13,318,284 |
| Length | 100 |
| Max. P | 0.989209 |

| Location | 13,318,184 – 13,318,284 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 75.44 |
| Mean single sequence MFE | -22.33 |
| Consensus MFE | -16.94 |
| Energy contribution | -16.75 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.90 |
| SVM RNA-class probability | 0.981989 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
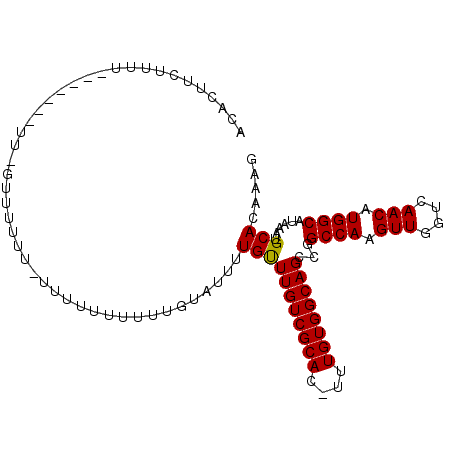
>X_DroMel_CAF1 13318184 100 + 22224390 ACCCUUCUUUUGCAUAUAUUUGUUUUUU-----UGUUUUUUAUUUUGUUUGUCGCACAUUUGUGGCAGUCGGCCAAGUUGGUCAACAUGGCAUAAAUGCACAAAG ......(((((((((.(((.........-----............((.((((((((....)))))))).))((((.(((....))).))))))).))))).)))) ( -22.00) >DroSim_CAF1 23565 105 + 1 ACCCUUCUUUUACAUAUAUUUGUUUUUUUUUUUUUUUUUGUAUUUUGUUUGUCGCACAUUUGUGGCAGUCGGCCAAGUUGGUCAACAUGGCAUAAAUGCACAAAG ..................................(((.(((((((((.((((((((....)))))))).))((((.(((....))).))))..))))))).))). ( -20.50) >DroEre_CAF1 22367 84 + 1 AUAUUUUU-------------GAUAUUU-UUUC----UUG--CUAUGCUUGUCGCAU-UUUGUGGCAGCCGGCCAAGUUGGUCAACAUGGCAUUAAUGCAAAAAG ...(((((-------------(.((((.-....----.((--(((((.(((((((..-...)))))))..((((.....))))..)))))))..)))))))))). ( -24.20) >DroYak_CAF1 22106 96 + 1 AUAUUUGUUGUU------UU-UGUGUUU-UUUUUUUUUUGGAUUUUGUUUGUCGCAU-UUUGUGGCAGCCGGCCAAGUUGGUCAACAUGGCAUGAAUGCACAAAG ............------((-(((((.(-((..............((.(((((((..-...))))))).))((((.(((....))).))))..))).))))))). ( -22.60) >consensus ACACUUCUUUU_______UU_GUUUUUU_UUUUUUUUUUGUAUUUUGUUUGUCGCAC_UUUGUGGCAGCCGGCCAAGUUGGUCAACAUGGCAUAAAUGCACAAAG .............................................(((((((((((....))))))))...((((.(((....))).))))......)))..... (-16.94 = -16.75 + -0.19)

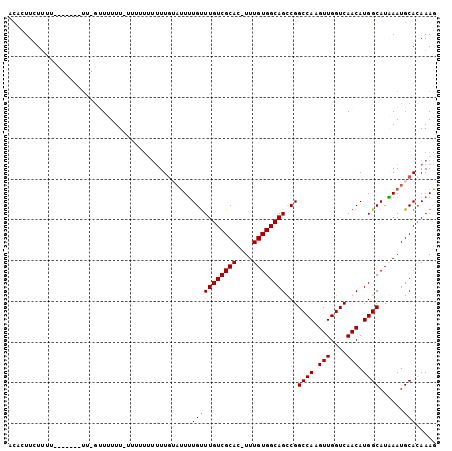
| Location | 13,318,184 – 13,318,284 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 75.44 |
| Mean single sequence MFE | -16.99 |
| Consensus MFE | -12.39 |
| Energy contribution | -11.95 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.73 |
| SVM decision value | 2.15 |
| SVM RNA-class probability | 0.989209 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
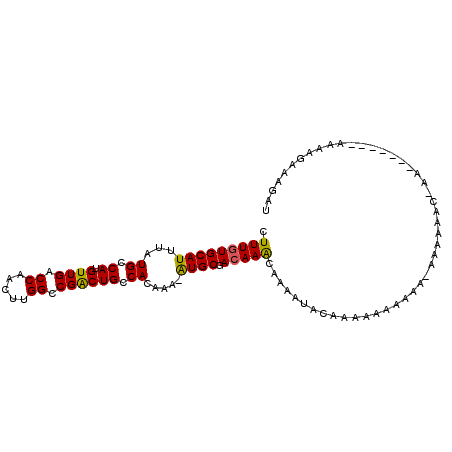
>X_DroMel_CAF1 13318184 100 - 22224390 CUUUGUGCAUUUAUGCCAUGUUGACCAACUUGGCCGACUGCCACAAAUGUGCGACAAACAAAAUAAAAAACA-----AAAAAACAAAUAUAUGCAAAAGAAGGGU ((((.(((((.(((((((.(((....))).))))....((((((....))).).))................-----.........))).)))))...))))... ( -18.50) >DroSim_CAF1 23565 105 - 1 CUUUGUGCAUUUAUGCCAUGUUGACCAACUUGGCCGACUGCCACAAAUGUGCGACAAACAAAAUACAAAAAAAAAAAAAAAAACAAAUAUAUGUAAAAGAAGGGU ..(((..(((((.((.((.((((.((.....)).)))))).)).)))))..)))................................................... ( -17.20) >DroEre_CAF1 22367 84 - 1 CUUUUUGCAUUAAUGCCAUGUUGACCAACUUGGCCGGCUGCCACAAA-AUGCGACAAGCAUAG--CAA----GAAA-AAAUAUC-------------AAAAAUAU .(((((((....((((..(((((.......((((.....))))....-...))))).)))).)--)))----))).-.......-------------........ ( -17.84) >DroYak_CAF1 22106 96 - 1 CUUUGUGCAUUCAUGCCAUGUUGACCAACUUGGCCGGCUGCCACAAA-AUGCGACAAACAAAAUCCAAAAAAAAAA-AAACACA-AA------AACAACAAAUAU .((((((((..(..((((.(((....))).)))).)..)).))))))-............................-.......-..------............ ( -14.40) >consensus CUUUGUGCAUUUAUGCCAUGUUGACCAACUUGGCCGACUGCCACAAA_AUGCGACAAACAAAAUACAAAAAAAAAA_AAAAAAC_AA_______AAAAGAAAGAU .(((((((((...((.((.((((.((.....)).)))))).)).....)))).)))))............................................... (-12.39 = -11.95 + -0.44)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:40:05 2006