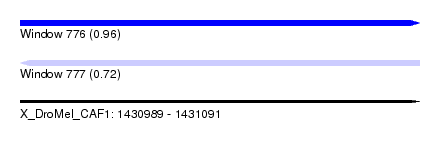

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,430,989 – 1,431,091 |
| Length | 102 |
| Max. P | 0.963259 |

| Location | 1,430,989 – 1,431,091 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 88.30 |
| Mean single sequence MFE | -42.46 |
| Consensus MFE | -31.83 |
| Energy contribution | -33.70 |
| Covariance contribution | 1.87 |
| Combinations/Pair | 1.07 |
| Mean z-score | -3.03 |
| Structure conservation index | 0.75 |
| SVM decision value | 1.55 |
| SVM RNA-class probability | 0.963259 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1430989 102 + 22224390 GGCUCAUUACGA-GCUGCCUGCAUAUUAUGUGCAG-C-CAGCAAUGCAGCGGCUCAGAAGCGACUUUGCCGUUGCCGUUGGAGCAUUGACAACUGCAGGCCGCCA (((((.....))-)))(((((((......((((..-(-((((...((((((((..((......))..)))))))).))))).)))).......)))))))..... ( -44.22) >DroSec_CAF1 35723 99 + 1 GGCUCAUUACGA-GCUGCCUGCAUAUUAUGUGCAGGC-CAGCAGUGCAUCGGCUCAGAAGCGACUUUGCCGUUGCCGUUGGAGCAUUGACAACUGCAGA----CA (((((.....))-)))((((((((.....))))))))-..(((((.((...((((((..(((((......)))))..)).))))..))...)))))...----.. ( -41.40) >DroEre_CAF1 32526 103 + 1 GGCUCAUUACGA-GCUGCCUGCAUAUUAUGUGCAGGC-CAGCAGUGCAGCGGCUCAGAAGCGACUUUGCCGUUGCCGUUGGAGCAUUGACAACUGCAGGCCGCCA (((((.....))-)))(((((((......((((...(-((((.(.((((((((..((......))..)))))))))))))).)))).......)))))))..... ( -45.52) >DroWil_CAF1 109699 99 + 1 GGCUCAUUAUGAAGUCGCCUGCAUAUUAUGUGCAGGCUCAGCGGCGCAGCGGCUCAGAACCGACUUUGCUG------UUGGAGCAUUGACAACUGCAGGCCAUCA ((((......(..(((((((((((.....))))))))(((((((((.(((((.......))).)).)))))------))))......)))..)....)))).... ( -38.70) >consensus GGCUCAUUACGA_GCUGCCUGCAUAUUAUGUGCAGGC_CAGCAGUGCAGCGGCUCAGAAGCGACUUUGCCGUUGCCGUUGGAGCAUUGACAACUGCAGGCCGCCA .((((...(((..(((((((((((.....))))))))..)))...((((((((..((......))..)))))))))))..))))..................... (-31.83 = -33.70 + 1.87)

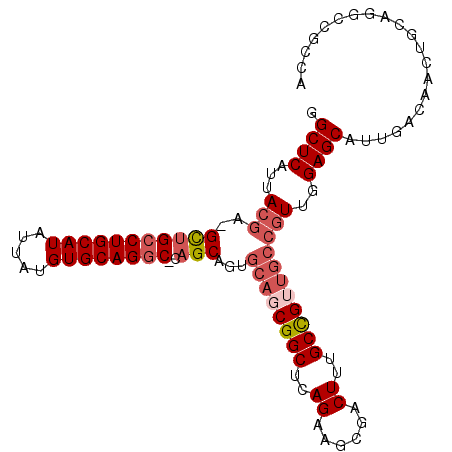
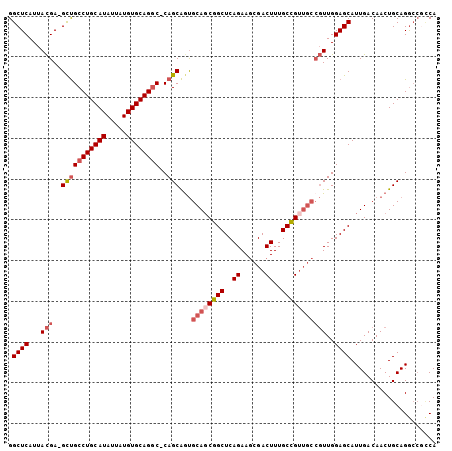
| Location | 1,430,989 – 1,431,091 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 88.30 |
| Mean single sequence MFE | -37.57 |
| Consensus MFE | -26.31 |
| Energy contribution | -28.62 |
| Covariance contribution | 2.31 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.715451 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1430989 102 - 22224390 UGGCGGCCUGCAGUUGUCAAUGCUCCAACGGCAACGGCAAAGUCGCUUCUGAGCCGCUGCAUUGCUG-G-CUGCACAUAAUAUGCAGGCAGC-UCGUAAUGAGCC (((((((.....)))))))..((((..((((((.((((..((......))..)))).)))...((((-.-(((((.......))))).))))-.)))...)))). ( -40.50) >DroSec_CAF1 35723 99 - 1 UG----UCUGCAGUUGUCAAUGCUCCAACGGCAACGGCAAAGUCGCUUCUGAGCCGAUGCACUGCUG-GCCUGCACAUAAUAUGCAGGCAGC-UCGUAAUGAGCC ..----...............((((..((((((.((((..((......))..)))).)))...((((-.((((((.......))))))))))-.)))...)))). ( -35.10) >DroEre_CAF1 32526 103 - 1 UGGCGGCCUGCAGUUGUCAAUGCUCCAACGGCAACGGCAAAGUCGCUUCUGAGCCGCUGCACUGCUG-GCCUGCACAUAAUAUGCAGGCAGC-UCGUAAUGAGCC (((((((.....)))))))..((((..((((((.((((..((......))..)))).)))...((((-.((((((.......))))))))))-.)))...)))). ( -41.30) >DroWil_CAF1 109699 99 - 1 UGAUGGCCUGCAGUUGUCAAUGCUCCAA------CAGCAAAGUCGGUUCUGAGCCGCUGCGCCGCUGAGCCUGCACAUAAUAUGCAGGCGACUUCAUAAUGAGCC (((..((.....))..)))..((((...------((((..((.((((.....)))))).....)))).(((((((.......)))))))...........)))). ( -33.40) >consensus UGGCGGCCUGCAGUUGUCAAUGCUCCAACGGCAACGGCAAAGUCGCUUCUGAGCCGCUGCACUGCUG_GCCUGCACAUAAUAUGCAGGCAGC_UCGUAAUGAGCC (((((((.....)))))))..((((..((((((.((((..((......))..)))).)))........(((((((.......))))))).....)))...)))). (-26.31 = -28.62 + 2.31)

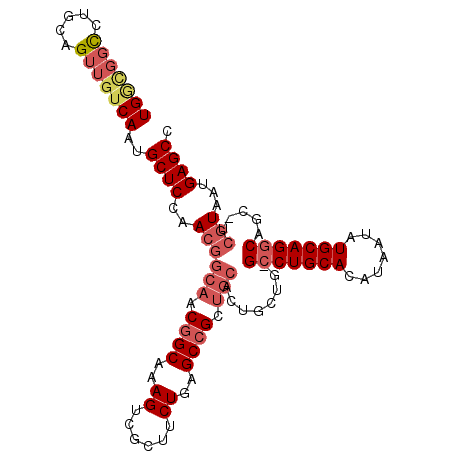
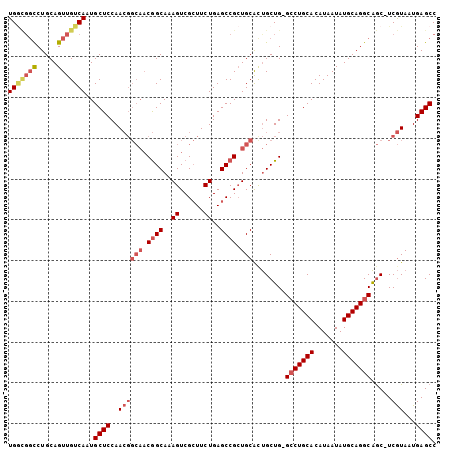
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:46:09 2006