
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 12,781,757 – 12,781,848 |
| Length | 91 |
| Max. P | 0.997752 |

| Location | 12,781,757 – 12,781,848 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 80.94 |
| Mean single sequence MFE | -36.44 |
| Consensus MFE | -25.04 |
| Energy contribution | -25.52 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.31 |
| Mean z-score | -3.27 |
| Structure conservation index | 0.69 |
| SVM decision value | 2.92 |
| SVM RNA-class probability | 0.997752 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
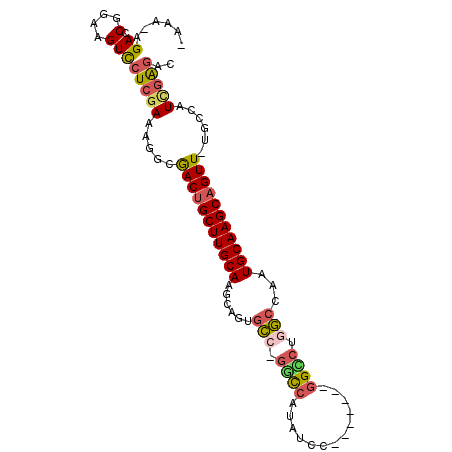
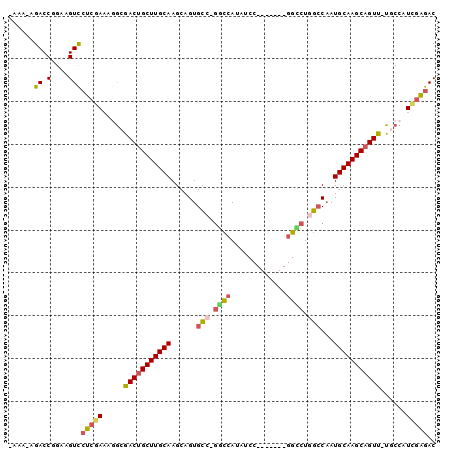
>X_DroMel_CAF1 12781757 91 + 22224390 -AAAUAGACCGGAAGUCCUCGAAAGGCGACUGCUUGCAAGCAGUGGCCUCUCAUAUCC-------GGUCUGCCCUAUGCAAGCAGUU-UGCCAUCGAGAC -.....((.(....)))(((((..((((((((((((((.((((..(((..........-------)))))))....))))))))).)-)))).))))).. ( -35.90) >DroSec_CAF1 5119 90 + 1 -AAA-AGACCGGAAGUCCUCGAACAGCGACUGCUUGCAAGCAGUGCCUGGCCAUAUCC-------GGCCUGGCCAAUGCAAGCUGUU-UGCCAUCGAGAC -...-.((.(....)))(((((...(((((.(((((((....(.(((.((((......-------)))).))))..))))))).).)-)))..))))).. ( -35.40) >DroSim_CAF1 6825 90 + 1 -AAA-AGACCGGAAGUCCUCGAACAGCGACUGCUUGCAAGCAGUGCCUGGCCAUAUCC-------GGCCUGGCCAAUGCAAGCAGUU-UGCCAUCGAGAC -...-.((.(....)))(((((...(((((((((((((....(.(((.((((......-------)))).))))..))))))))).)-)))..))))).. ( -40.00) >DroEre_CAF1 5352 97 + 1 CAAU-CGACCGGAAGUUCUCGAAAGGCGACUGCUUGCAAGCAGUGCG-GUCAACAUCCGGAGUCCGGACUGGCCAAUGCAAGCAGUU-UACCAUGCGGAC ...(-(((.(....)...))))..((.(((((((((((.((...))(-((((...(((((...))))).)))))..)))))))))))-..))........ ( -36.20) >DroYak_CAF1 6662 90 + 1 -AAG-AGACCGGAAGUCAUCGAUGGACAACUGCUUGCAAGCAGUGCG-GUCCAUAUCC-------GGGCUAGACAAUGCAAGCAGUUUUACCAUUGGGAC -...-.........(((.(((((((..(((((((((((..((((.((-(.......))-------).))).)....)))))))))))...)))))))))) ( -34.70) >consensus _AAA_AGACCGGAAGUCCUCGAAAGGCGACUGCUUGCAAGCAGUGCC_GGCCAUAUCC_______GGCCUGGCCAAUGCAAGCAGUU_UGCCAUCGAGAC ......((.(....)))(((((.....(((((((((((......(((.((((.............)))).)))...)))))))))))......))))).. (-25.04 = -25.52 + 0.48)

| Location | 12,781,757 – 12,781,848 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 80.94 |
| Mean single sequence MFE | -35.88 |
| Consensus MFE | -21.90 |
| Energy contribution | -23.62 |
| Covariance contribution | 1.72 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.58 |
| Structure conservation index | 0.61 |
| SVM decision value | 1.61 |
| SVM RNA-class probability | 0.967441 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
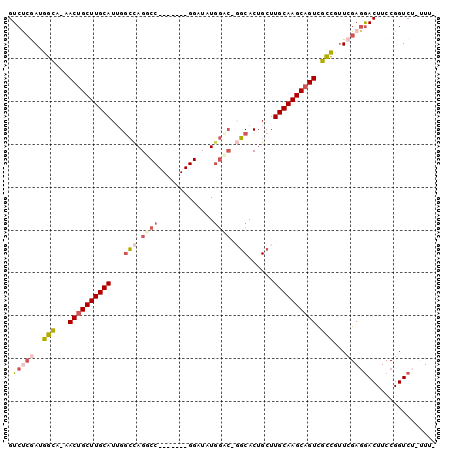
>X_DroMel_CAF1 12781757 91 - 22224390 GUCUCGAUGGCA-AACUGCUUGCAUAGGGCAGACC-------GGAUAUGAGAGGCCACUGCUUGCAAGCAGUCGCCUUUCGAGGACUUCCGGUCUAUUU- ((((.....(((-(.....))))...))))(((((-------(((..((((((((.((((((....)))))).))))))))......))))))))....- ( -39.80) >DroSec_CAF1 5119 90 - 1 GUCUCGAUGGCA-AACAGCUUGCAUUGGCCAGGCC-------GGAUAUGGCCAGGCACUGCUUGCAAGCAGUCGCUGUUCGAGGACUUCCGGUCU-UUU- ..(((((((((.-.((.(((((((..((((.((((-------(....))))).))).)....))))))).)).)))).)))))(((.....))).-...- ( -37.00) >DroSim_CAF1 6825 90 - 1 GUCUCGAUGGCA-AACUGCUUGCAUUGGCCAGGCC-------GGAUAUGGCCAGGCACUGCUUGCAAGCAGUCGCUGUUCGAGGACUUCCGGUCU-UUU- ..(((((((((.-.((((((((((..((((.((((-------(....))))).))).)....)))))))))).)))).)))))(((.....))).-...- ( -42.30) >DroEre_CAF1 5352 97 - 1 GUCCGCAUGGUA-AACUGCUUGCAUUGGCCAGUCCGGACUCCGGAUGUUGAC-CGCACUGCUUGCAAGCAGUCGCCUUUCGAGAACUUCCGGUCG-AUUG (((((..((((.-...((....))...))))...))))).((((((.((((.-.((((((((....)))))).))...)))).)...)))))...-.... ( -32.10) >DroYak_CAF1 6662 90 - 1 GUCCCAAUGGUAAAACUGCUUGCAUUGUCUAGCCC-------GGAUAUGGAC-CGCACUGCUUGCAAGCAGUUGUCCAUCGAUGACUUCCGGUCU-CUU- ......((((...(((((((((((..((..((..(-------((.......)-))..)))).)))))))))))..)))).((.(((.....))))-)..- ( -28.20) >consensus GUCUCGAUGGCA_AACUGCUUGCAUUGGCCAGGCC_______GGAUAUGGAC_GGCACUGCUUGCAAGCAGUCGCCGUUCGAGGACUUCCGGUCU_UUU_ .(((((..(((...((((((((((...(((.((((.............)))).)))......)))))))))).)))...)))))................ (-21.90 = -23.62 + 1.72)
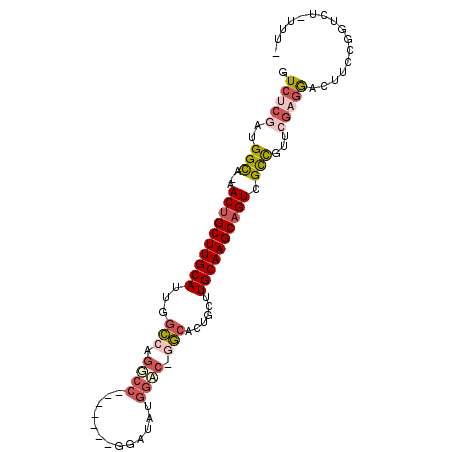


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:35:50 2006