
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 11,238,792 – 11,238,883 |
| Length | 91 |
| Max. P | 0.843574 |

| Location | 11,238,792 – 11,238,883 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 86.57 |
| Mean single sequence MFE | -21.25 |
| Consensus MFE | -17.00 |
| Energy contribution | -17.28 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.843574 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
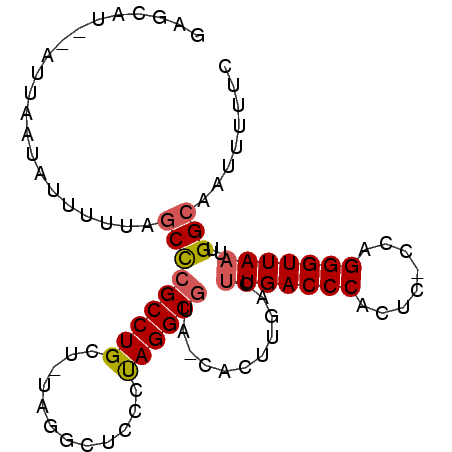
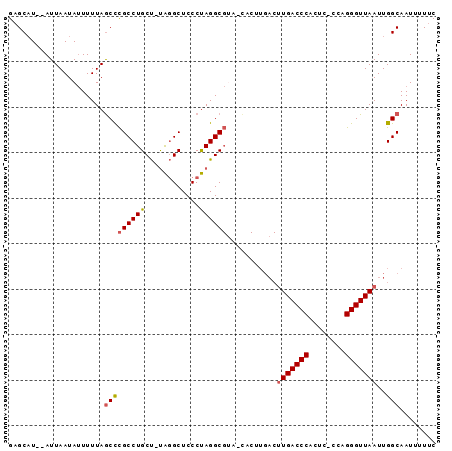
>X_DroMel_CAF1 11238792 91 + 22224390 GAGCAUAUAUUAAUAUUUUUAGCUCGCCUGCU-UAGGCUCCCUAGGCGUA-UACUUGACUUGACCCACUC-CCAGGGUUAAUUGGCAAUUUUUC ((((.................))))(((.(((-((((...)))))))...-........(((((((....-...)))))))..)))........ ( -25.93) >DroSec_CAF1 25739 92 + 1 GAGCAUAUAUUAAUAUUUUUAGCCCGCCUGCUUUAGGCUCCCUAGGCGUA-UACUUGACUUGACCCACUC-CCAGGGUUAAUUGGCAAUUUUUC .....................((((((((((.....)).....)))))..-........(((((((....-...)))))))..)))........ ( -19.80) >DroEre_CAF1 25454 90 + 1 GAGCAU--AUUAAUAUUUUUAGCCCGCCUGCU-UAGGCUCCCUAGGCGAACCACUUGACUUGACCCACUC-CCAGGGUUAAUUGGCAAUUUUUC ......--.................(((((((-((((...))))))))...........(((((((....-...)))))))..)))........ ( -22.00) >DroYak_CAF1 17634 89 + 1 GAGCAU--AUUAAUAUUUUUAGCCCGCCUGCU-UAGGCUCCCUAGGCGUA-CACUUGACUUGACCCACUC-CCAGGGUUAAUUGGCAAUUUUUC ......--.................(((.(((-((((...)))))))...-........(((((((....-...)))))))..)))........ ( -21.90) >DroAna_CAF1 20231 79 + 1 GCGCUU--AUUAAUACUUUUAACCUGCCUGCC-CGAGCUCCCCAGGC------------CUGACCCACUCCCCAGGGUUAAUUGGCAAUUUUUC ..(((.--((((((........((((...((.-...))....))))(------------(((..........)))))))))).)))........ ( -16.60) >consensus GAGCAU__AUUAAUAUUUUUAGCCCGCCUGCU_UAGGCUCCCUAGGCGUA_CACUUGACUUGACCCACUC_CCAGGGUUAAUUGGCAAUUUUUC .....................(((((((((............))))))...........(((((((........)))))))..)))........ (-17.00 = -17.28 + 0.28)

| Location | 11,238,792 – 11,238,883 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 86.57 |
| Mean single sequence MFE | -24.53 |
| Consensus MFE | -16.21 |
| Energy contribution | -16.33 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.66 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.510775 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
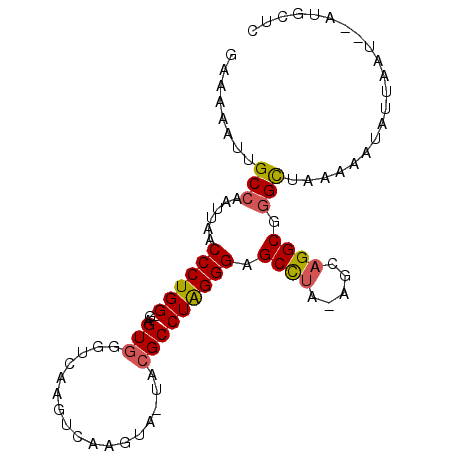
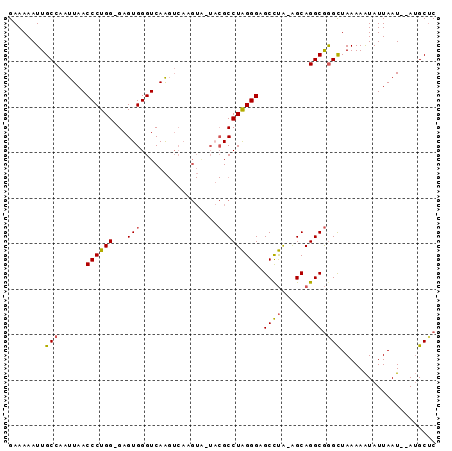
>X_DroMel_CAF1 11238792 91 - 22224390 GAAAAAUUGCCAAUUAACCCUGG-GAGUGGGUCAAGUCAAGUA-UACGCCUAGGGAGCCUA-AGCAGGCGAGCUAAAAAUAUUAAUAUAUGCUC ........(((......((((((-(.(((...(.......)..-))).))))))).((...-.)).)))((((.................)))) ( -23.93) >DroSec_CAF1 25739 92 - 1 GAAAAAUUGCCAAUUAACCCUGG-GAGUGGGUCAAGUCAAGUA-UACGCCUAGGGAGCCUAAAGCAGGCGGGCUAAAAAUAUUAAUAUAUGCUC ........(((......((((((-(.(((...(.......)..-))).))))))).((((.....)))).)))..................... ( -24.80) >DroEre_CAF1 25454 90 - 1 GAAAAAUUGCCAAUUAACCCUGG-GAGUGGGUCAAGUCAAGUGGUUCGCCUAGGGAGCCUA-AGCAGGCGGGCUAAAAAUAUUAAU--AUGCUC ........(((......((((((-(...(..(((.......)))..).))))))).((((.-...)))).))).............--...... ( -26.60) >DroYak_CAF1 17634 89 - 1 GAAAAAUUGCCAAUUAACCCUGG-GAGUGGGUCAAGUCAAGUG-UACGCCUAGGGAGCCUA-AGCAGGCGGGCUAAAAAUAUUAAU--AUGCUC ........(((......((((((-(.(((.(.(.......)).-))).))))))).((((.-...)))).))).............--...... ( -25.10) >DroAna_CAF1 20231 79 - 1 GAAAAAUUGCCAAUUAACCCUGGGGAGUGGGUCAG------------GCCUGGGGAGCUCG-GGCAGGCAGGUUAAAAGUAUUAAU--AAGCGC ......(((((.....((((........))))...------------(((((((...))))-))).)))))...............--...... ( -22.20) >consensus GAAAAAUUGCCAAUUAACCCUGG_GAGUGGGUCAAGUCAAGUA_UACGCCUAGGGAGCCUA_AGCAGGCGGGCUAAAAAUAUUAAU__AUGCUC ........(((......((((((...(((.................))))))))).((((.....)))).)))..................... (-16.21 = -16.33 + 0.12)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:20:55 2006