
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 11,061,485 – 11,061,576 |
| Length | 91 |
| Max. P | 0.776595 |

| Location | 11,061,485 – 11,061,576 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 83.51 |
| Mean single sequence MFE | -17.62 |
| Consensus MFE | -10.06 |
| Energy contribution | -12.00 |
| Covariance contribution | 1.94 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.553173 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
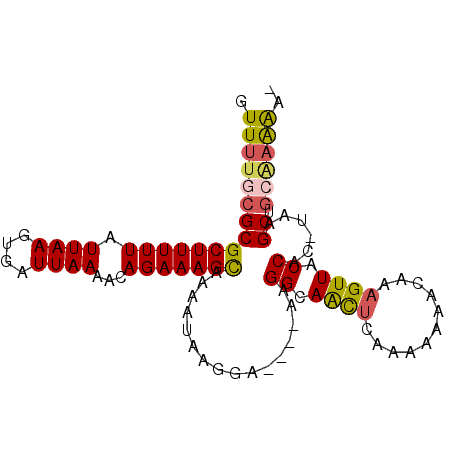
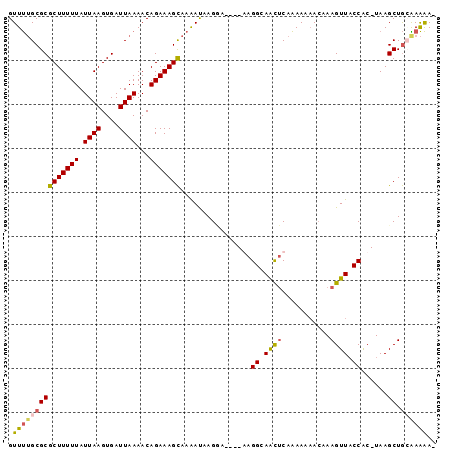
>X_DroMel_CAF1 11061485 91 + 22224390 GUUUUGCGCGCUUUUUAUUAAGUGAUUAAAACAGAAAGCAAAAUAAGGA----AAGGCAACUCAAAAAAACAAAGUUACCAC-UAAGCUGCAAAAA- .(((((((((((((((.((((....))))...)))))))..........----..((.((((...........)))).))..-...)).)))))).- ( -18.10) >DroPse_CAF1 38785 95 + 1 GUUCAGCGCGCUUUUUAUUAAGUGAUUAAAACAGAAAGUACGACAAGCAAAAAAAGGGAGUGCAGAA-AAGAAAGUUUCCGCUUAAGCUAAGAGGA- ..........(((((..(((((((.............((((..(............)..)))).(((-(......)))))))))))...)))))..- ( -14.50) >DroSim_CAF1 15393 91 + 1 GUUUUGCGCGCUUUUUAUUAAGUGAUUAAAACAGAAAGCAAAAUAAGGA----GAGGCAACUCAAAAAAACAAAGUUACCAC-UAAGCUGCGAAAA- .(((((((((((((((.((((....))))...))))))).......((.----(((....))).....(((...))).))..-...)).)))))).- ( -20.70) >DroEre_CAF1 15757 91 + 1 GUUUUGCGCGCUUUUUAUUAAGUGAUUAAAACAGAAAGCAAAAUAAGUA----GAGGCAACUCAAAAA-ACAAAGUUACCAC-UAAGCUGGAAAAAC (((((....(((((((.((((....))))...)))))))..........----(((....)))..)))-)).......(((.-.....)))...... ( -17.20) >consensus GUUUUGCGCGCUUUUUAUUAAGUGAUUAAAACAGAAAGCAAAAUAAGGA____AAGGCAACUCAAAAAAACAAAGUUACCAC_UAAGCUGCAAAAA_ .(((((((((((((((.((((....))))...)))))))................((.((((...........)))).))......)).)))))).. (-10.06 = -12.00 + 1.94)

| Location | 11,061,485 – 11,061,576 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 83.51 |
| Mean single sequence MFE | -19.23 |
| Consensus MFE | -10.93 |
| Energy contribution | -12.18 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.57 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.776595 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
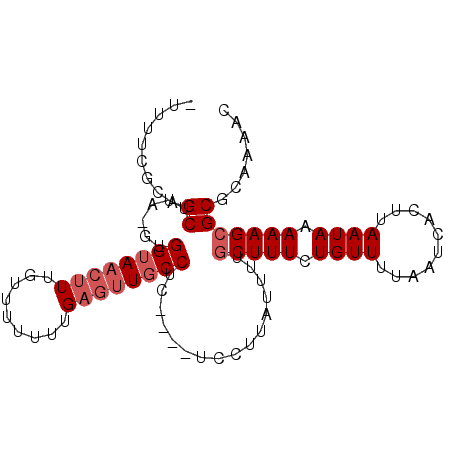
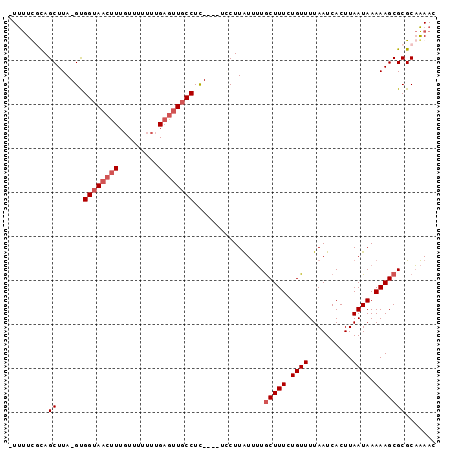
>X_DroMel_CAF1 11061485 91 - 22224390 -UUUUUGCAGCUUA-GUGGUAACUUUGUUUUUUUGAGUUGCCUU----UCCUUAUUUUGCUUUCUGUUUUAAUCACUUAAUAAAAAGCGCGCAAAAC -.((((((.((...-..((((((((.........))))))))..----..........(((((.((((..........)))).))))))))))))). ( -21.10) >DroPse_CAF1 38785 95 - 1 -UCCUCUUAGCUUAAGCGGAAACUUUCUU-UUCUGCACUCCCUUUUUUUGCUUGUCGUACUUUCUGUUUUAAUCACUUAAUAAAAAGCGCGCUGAAC -.....(((((....(((((((......)-))))))....................(..((((.((((..........)))).))))..)))))).. ( -16.60) >DroSim_CAF1 15393 91 - 1 -UUUUCGCAGCUUA-GUGGUAACUUUGUUUUUUUGAGUUGCCUC----UCCUUAUUUUGCUUUCUGUUUUAAUCACUUAAUAAAAAGCGCGCAAAAC -(((((((.....(-(.((((((((.........)))))))).)----).........(((((.((((..........)))).)))))))).)))). ( -19.30) >DroEre_CAF1 15757 91 - 1 GUUUUUCCAGCUUA-GUGGUAACUUUGU-UUUUUGAGUUGCCUC----UACUUAUUUUGCUUUCUGUUUUAAUCACUUAAUAAAAAGCGCGCAAAAC (((((....((.((-(.((((((((...-.....)))))))).)----))........(((((.((((..........)))).)))))))..))))) ( -19.90) >consensus _UUUUCGCAGCUUA_GUGGUAACUUUGUUUUUUUGAGUUGCCUC____UCCUUAUUUUGCUUUCUGUUUUAAUCACUUAAUAAAAAGCGCGCAAAAC .........((......((((((((.........))))))))................(((((.((((..........)))).)))))))....... (-10.93 = -12.18 + 1.25)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:19:34 2006