
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 11,016,714 – 11,016,870 |
| Length | 156 |
| Max. P | 0.724516 |

| Location | 11,016,714 – 11,016,833 |
|---|---|
| Length | 119 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.44 |
| Mean single sequence MFE | -33.72 |
| Consensus MFE | -26.97 |
| Energy contribution | -28.17 |
| Covariance contribution | 1.20 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.35 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.667536 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
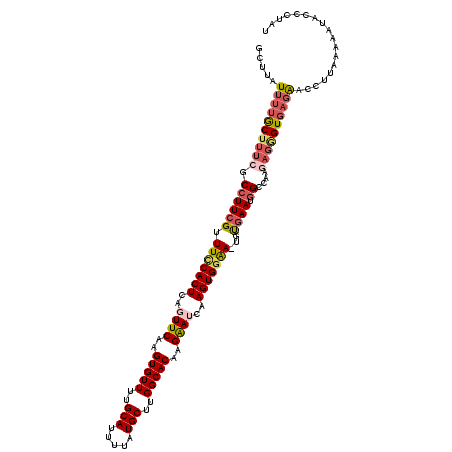
>X_DroMel_CAF1 11016714 119 - 22224390 GCUUAUUUUGCUUUCGCCUUCGUUUUCACUCAGUUCAAGUGUUUUGCAUUUUAUGCUUGGCACAAGAAUCAAGUGGAA-UGUGAAUGGCCAAGAGGGUGAGAACCUUAAAAAUACCGUAU .....(((..(((((((((((..(((((((..((((..(((((..((((...))))..)))))..))))..)))))))-...))).)))...)))))..))).................. ( -33.80) >DroSec_CAF1 9469 119 - 1 GCUUAUUUUACUGUCGCCUUCGUUUCCACUCAGUUCAAGUGUUUUGCAUUUUAUGCUUGGCACAAGAAUCAAGUGGAA-UGUGAAUGGACAACAGAGUGAGAACCUUAAAAAUACCGUAU .((((((((..((((.(.(((..(((((((..((((..(((((..((((...))))..)))))..))))..)))))))-...))).)))))..))))))))................... ( -35.00) >DroSim_CAF1 5173 118 - 1 GCUUAUUUUGCUUUCGCCUUCGUUUCCACUCAGUUCAAGUGUUUUGCAUUUUAUGCUUGGCACAAGAAUCAAGUGGAA-UGUGAAUGGACAAAAGAGUGAGAACCUUAAAAA-ACCGUAU .((((((((.......(((((..(((((((..((((..(((((..((((...))))..)))))..))))..)))))))-...))).)).....))))))))...........-....... ( -31.90) >DroEre_CAF1 3329 119 - 1 GCUUAUUUUGCUUUCGCCUUCGUUUCCACUCAGUUCAAGUGUUUUGCAUUUUAUGCUUGGCACGAGGAGCAAGUGGAA-UGCGAAUGGCCAAGAGGGUGAGAACCUAAAAAAUACCCUAU .....(((..((((((((((((((((((((..((((..(((((..((((...))))..)))))...)))).)))))))-.))))).)))...)))))..))).................. ( -41.70) >DroYak_CAF1 4622 119 - 1 GCUUAUUUUGCUUUCGCCUUCGUUUCCACUCAGUUCAAGUGUUUUGCAUUUUAUGCUUGGCACAAGAAACAAGUGGAA-UGUGAAUGGCCAGGAGGGUGAGAACCUUAAAAAUACCCUAU .....(((..(((((((((((..(((((((...(((..(((((..((((...))))..)))))..)))...)))))))-...))).)))...)))))..))).................. ( -34.90) >DroAna_CAF1 4108 120 - 1 GCUUAUUUUGCUUUCGCCUUCGUUUCCACUCAGUUCAAGUGUUUUGCAUUUUCUGCUUGGCACAAGAAUCAAGUGAGACCAUAAAUUGGAGAGUGUGUUUUGACUUUAUUAAUACCCUGU .........((((((......(((((.(((..((((..(((((..(((.....)))..)))))..))))..))))))))((.....))))))))(((((...........)))))..... ( -25.00) >consensus GCUUAUUUUGCUUUCGCCUUCGUUUCCACUCAGUUCAAGUGUUUUGCAUUUUAUGCUUGGCACAAGAAUCAAGUGGAA_UGUGAAUGGCCAAGAGGGUGAGAACCUUAAAAAUACCCUAU .....((((((((((.((((((.(((((((..((((..(((((..(((.....)))..)))))..))))..)))))))...)))).))....)))))))))).................. (-26.97 = -28.17 + 1.20)


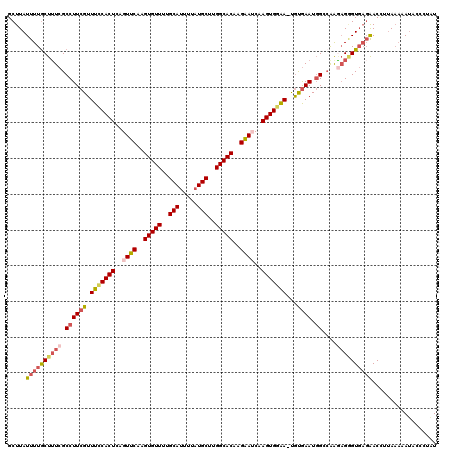
| Location | 11,016,754 – 11,016,870 |
|---|---|
| Length | 116 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.85 |
| Mean single sequence MFE | -22.02 |
| Consensus MFE | -17.59 |
| Energy contribution | -18.45 |
| Covariance contribution | 0.86 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.724516 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
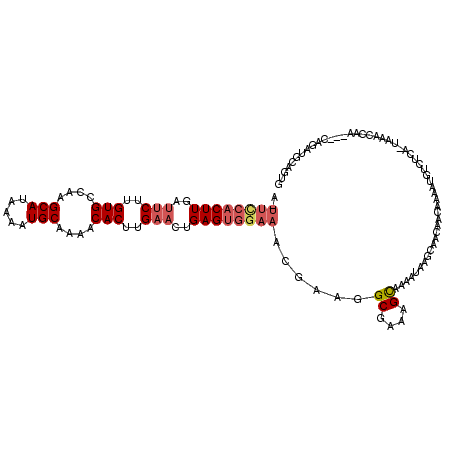
>X_DroMel_CAF1 11016754 116 + 22224390 A-UUCCACUUGAUUCUUGUGCCAAGCAUAAAAUGCAAAACACUUGAACUGAGUGAAAACGAAGGCGAAAGCAAAAUAAGCAACAACAAAAUGUCUCA-UAAACCAAC--CAGAUGCAGUG .-...(((((..(((..(((....((((...))))....)))..)))..))))).........((....))..............((...(((.((.-.........--..)).))).)) ( -19.70) >DroSec_CAF1 9509 118 + 1 A-UUCCACUUGAUUCUUGUGCCAAGCAUAAAAUGCAAAACACUUGAACUGAGUGGAAACGAAGGCGACAGUAAAAUAAGCAACAACAAAAUGUCUCA-UAAACCAAACCAAGAUGCAGUG .-((((((((..(((..(((....((((...))))....)))..)))..))))))))......(.((((.....................)))).).-...................... ( -22.20) >DroEre_CAF1 3369 115 + 1 A-UUCCACUUGCUCCUCGUGCCAAGCAUAAAAUGCAAAACACUUGAACUGAGUGGAAACGAAGGCGAAAGCAAAAUAAGCAGCAACAAAAUGUCUCA-UAAACCAA---CAGAUGCAGUG .-...((((((((.(((((..(..((((...))))....(((((.....))))))..))))..((....)).......).))))).....(((.((.-........---..)).)))))) ( -21.80) >DroYak_CAF1 4662 115 + 1 A-UUCCACUUGUUUCUUGUGCCAAGCAUAAAAUGCAAAACACUUGAACUGAGUGGAAACGAAGGCGAAAGCAAAAUAAGCAACAACAAAAUGUCUCA-UAAACCAA---CAGAUGCACUG .-((((((((..(((..(((....((((...))))....)))..)))..))))))))......((....))...................(((.((.-........---..)).)))... ( -24.30) >DroMoj_CAF1 4304 99 + 1 --GUUGUCUUUUAUCUUGUGAAAAGCAUGAAAUGCAAAACACUUGAACUGAGUGGAAAUGAAGGCGAAAGCAAAAUAAGUAACAAAACAA----------------GACAAGAAG---CG --...(..((((.((((((.....((((...)))).....(((((..((..((....))..))((....))....)))))......))))----------------)).))))..---). ( -21.20) >DroAna_CAF1 4148 120 + 1 GGUCUCACUUGAUUCUUGUGCCAAGCAGAAAAUGCAAAACACUUGAACUGAGUGGAAACGAAGGCGAAAGCAAAAUAAGCAACAACAAAAUGUCUUUCUUAACAAACACCAGAAGCACUG (((.((((((..(((..(((....(((.....)))....)))..)))..))))))........((....))....((((.((..((.....))..))))))......))).......... ( -22.90) >consensus A_UUCCACUUGAUUCUUGUGCCAAGCAUAAAAUGCAAAACACUUGAACUGAGUGGAAACGAAGGCGAAAGCAAAAUAAGCAACAACAAAAUGUCUCA_UAAACCAA___CAGAUGCAGUG ..((((((((..(((..(((....(((.....)))....)))..)))..))))))))......((....))................................................. (-17.59 = -18.45 + 0.86)

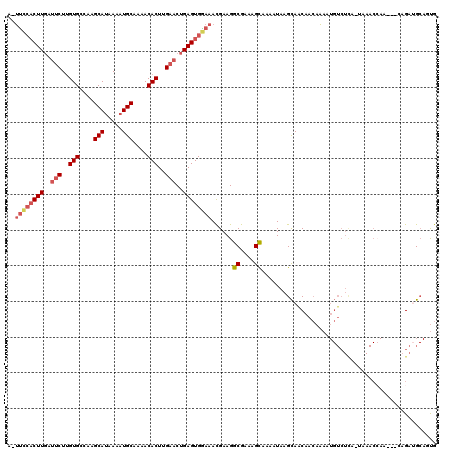
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:18:58 2006