
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,910,003 – 10,910,095 |
| Length | 92 |
| Max. P | 0.930861 |

| Location | 10,910,003 – 10,910,095 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 76.23 |
| Mean single sequence MFE | -20.93 |
| Consensus MFE | -16.33 |
| Energy contribution | -16.00 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.35 |
| Structure conservation index | 0.78 |
| SVM decision value | 1.10 |
| SVM RNA-class probability | 0.914552 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
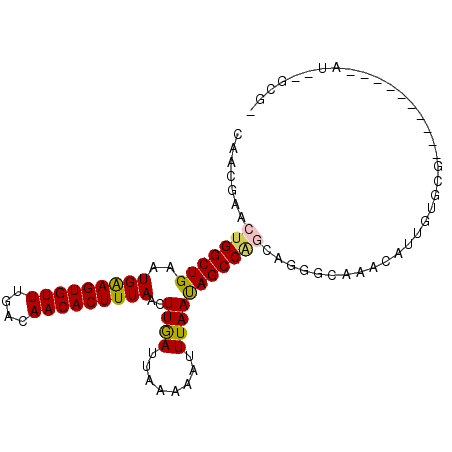
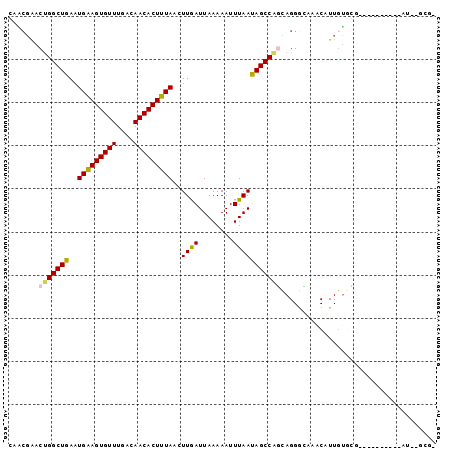
>X_DroMel_CAF1 10910003 92 + 22224390 AAACGAACUGGCUGAAUGAAGUGUUUGACAACACUUUAACUUGAUUAAAAAUUUAAUAGCCAGCAGGGCAAACAUUGUGCGAAUGG-----AU--GGAU .......(((((((..(((((((((....)))))))))..((((........))))))))))).....((..((((.....)))).-----.)--)... ( -20.30) >DroPse_CAF1 23306 86 + 1 CAGCGAACUGGCUGAAUGGAGUGUUUGACAACACUUUAACUUAAUUAAAAAUUUAACAGCCGUUAGGGGAUGCAUUGUGGG----------AU--GCG- .....(((.(((((..(((((((((....)))))))))..((((........)))))))))))).......(((((....)----------))--)).- ( -20.60) >DroYak_CAF1 8652 94 + 1 GUACGAACUGGCUGAAUGAAGUGUUUGACAACACUUUAACUUGAUUAAAAAUUUAAUAGCCAGCAGGGCAAACAUUGUGCGAAUGG-----GCUGAGGA ((((((.(((((((..(((((((((....)))))))))..((((........)))))))))))..(......).))))))......-----........ ( -22.60) >DroMoj_CAF1 11867 77 + 1 -GUCUGACGGGCUGAAUGAAGUGUUUGGCAACACUUUAACUUGAUUAAAAAUUUAAUAGCCAGCAGGAUAAACAUUUG--------------------- -.((((...(((((..(((((((((....)))))))))..((((........)))))))))..))))...........--------------------- ( -18.60) >DroAna_CAF1 9073 95 + 1 G--GGAACUGGCUGAAUGAAGUGUUUGACAACACUUUAACUUGAUUAAAAAUUUAAUAGCCAGCAGGGCAAACAUUGUACCAUUGCUGGGGGC--GUAA .--.......(((...(((((((((....))))))))).....................(((((((.(((.....)))....))))))).)))--.... ( -22.90) >DroPer_CAF1 17058 86 + 1 CAGCGAACUGGCUGAAUGGAGUGUUUGACAACACUUUAACUUAAUUAAAAAUUUAACAGCCGUUAGGGGAUGCAUUGUGGG----------AU--GCG- .....(((.(((((..(((((((((....)))))))))..((((........)))))))))))).......(((((....)----------))--)).- ( -20.60) >consensus CAACGAACUGGCUGAAUGAAGUGUUUGACAACACUUUAACUUGAUUAAAAAUUUAAUAGCCAGCAGGGCAAACAUUGUGCG__________AU__GCG_ .......(((((((..(((((((((....)))))))))..((((........))))))))))).................................... (-16.33 = -16.00 + -0.33)

| Location | 10,910,003 – 10,910,095 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 76.23 |
| Mean single sequence MFE | -17.57 |
| Consensus MFE | -10.92 |
| Energy contribution | -11.25 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.62 |
| SVM decision value | 1.21 |
| SVM RNA-class probability | 0.930861 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
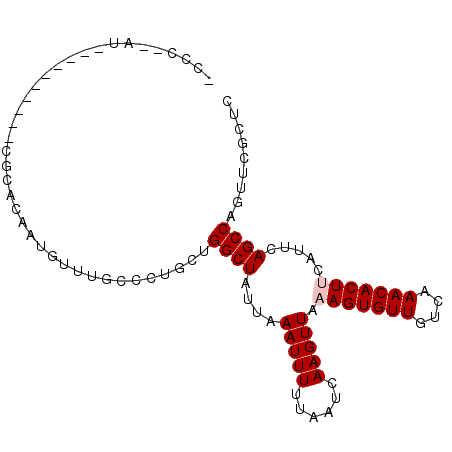
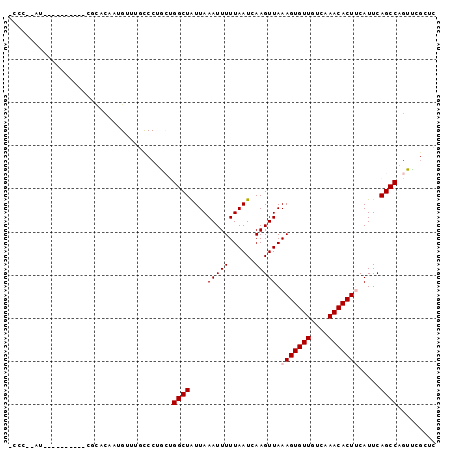
>X_DroMel_CAF1 10910003 92 - 22224390 AUCC--AU-----CCAUUCGCACAAUGUUUGCCCUGCUGGCUAUUAAAUUUUUAAUCAAGUUAAAGUGUUGUCAAACACUUCAUUCAGCCAGUUCGUUU ...(--(.-----.((((.....))))..))....(((((((....(((((......))))).(((((((....))))))).....)))))))...... ( -17.90) >DroPse_CAF1 23306 86 - 1 -CGC--AU----------CCCACAAUGCAUCCCCUAACGGCUGUUAAAUUUUUAAUUAAGUUAAAGUGUUGUCAAACACUCCAUUCAGCCAGUUCGCUG -.((--((----------......))))..........(((((...(((((......)))))..((((((....)))))).....)))))......... ( -17.30) >DroYak_CAF1 8652 94 - 1 UCCUCAGC-----CCAUUCGCACAAUGUUUGCCCUGCUGGCUAUUAAAUUUUUAAUCAAGUUAAAGUGUUGUCAAACACUUCAUUCAGCCAGUUCGUAC ......((-----......))........(((...(((((((....(((((......))))).(((((((....))))))).....)))))))..))). ( -18.70) >DroMoj_CAF1 11867 77 - 1 ---------------------CAAAUGUUUAUCCUGCUGGCUAUUAAAUUUUUAAUCAAGUUAAAGUGUUGCCAAACACUUCAUUCAGCCCGUCAGAC- ---------------------............((((.((((....(((((......))))).(((((((....))))))).....)))).).)))..- ( -14.60) >DroAna_CAF1 9073 95 - 1 UUAC--GCCCCCAGCAAUGGUACAAUGUUUGCCCUGCUGGCUAUUAAAUUUUUAAUCAAGUUAAAGUGUUGUCAAACACUUCAUUCAGCCAGUUCC--C ....--....(((....)))...............(((((((....(((((......))))).(((((((....))))))).....)))))))...--. ( -19.60) >DroPer_CAF1 17058 86 - 1 -CGC--AU----------CCCACAAUGCAUCCCCUAACGGCUGUUAAAUUUUUAAUUAAGUUAAAGUGUUGUCAAACACUCCAUUCAGCCAGUUCGCUG -.((--((----------......))))..........(((((...(((((......)))))..((((((....)))))).....)))))......... ( -17.30) >consensus _CCC__AU__________CGCACAAUGUUUGCCCUGCUGGCUAUUAAAUUUUUAAUCAAGUUAAAGUGUUGUCAAACACUUCAUUCAGCCAGUUCGCUC ......................................((((....(((((......))))).(((((((....))))))).....))))......... (-10.92 = -11.25 + 0.33)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:17:51 2006