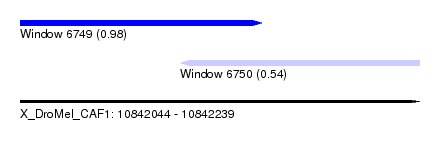
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,842,044 – 10,842,239 |
| Length | 195 |
| Max. P | 0.980762 |

| Location | 10,842,044 – 10,842,162 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.94 |
| Mean single sequence MFE | -25.62 |
| Consensus MFE | -24.64 |
| Energy contribution | -24.45 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.46 |
| Structure conservation index | 0.96 |
| SVM decision value | 1.87 |
| SVM RNA-class probability | 0.980762 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
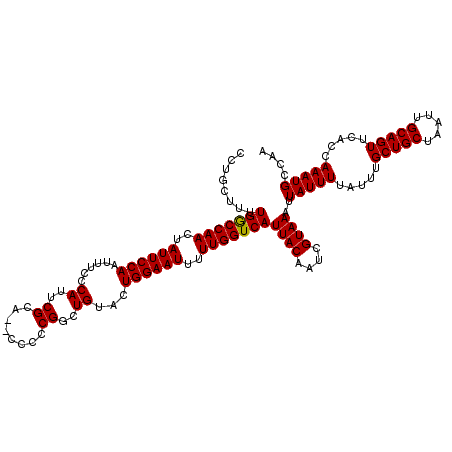
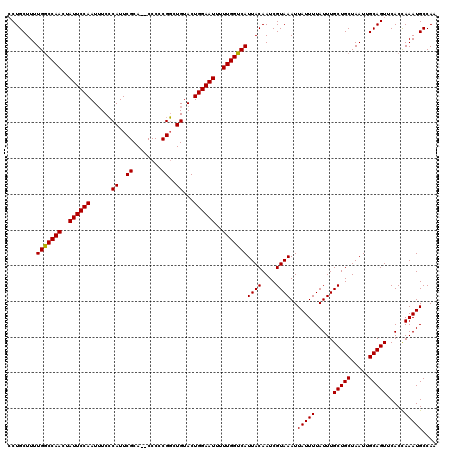
>X_DroMel_CAF1 10842044 118 + 22224390 CCUGCUUUUGGCCAACUAUUCCAAUUUCCCAUUCGCA--CCCCCGGUUGUACUGGAAUUUUUGGUCAUUACAAUCGUAAAUUAUUUUAUUUGCUGCUAAUUGCAGUUCACCAAAUGCCAA .((((...(((((((..((((((..(..((.......--.....))..)...))))))..)))))))........((((((......))))))........))))............... ( -24.40) >DroSec_CAF1 20847 118 + 1 GCUGCUUUUGACCAACUAUUCCAAUUUCCCAUUCGCA--CCCCCGGCUGUACUGGAAUUUUUGGUCAUUACAAUCGUAAAUUAUUUUAUUUGCUGCUAAUUGCAGUUCACCAAAUGCCAA (((((...(((((((..((((((......((..((..--....))..))...))))))..)))))))........((((((......))))))........))))).............. ( -26.20) >DroSim_CAF1 1709 118 + 1 CCUGCUUUUGGCCAACUAUUCCAAUUUCCCAUUCGCA--CCCCCGGCUGUACUGGAAUUUUUGGUCAUUACAAUCGUAAAUUAUUUUAUUUGCUGCUAAUUGCAGUUCACCAAAUGCCAA .((((...(((((((..((((((......((..((..--....))..))...))))))..)))))))........((((((......))))))........))))............... ( -24.40) >DroYak_CAF1 9834 120 + 1 GCUGCUUUUGGCCAACUAUUCCAAUAUCGCAUUCGCAGCCCCCCGUUUGUACUGGAAUUUUUGGUCAUUACAAUCGUAAAUUAUUUUAUUUGCUGCUAAUUGCAGUUCACUAAAUGCCAA (((((...(((((((..((((((.....(((..((........))..)))..))))))..)))))))........((((((......))))))........))))).............. ( -27.50) >consensus CCUGCUUUUGGCCAACUAUUCCAAUUUCCCAUUCGCA__CCCCCGGCUGUACUGGAAUUUUUGGUCAUUACAAUCGUAAAUUAUUUUAUUUGCUGCUAAUUGCAGUUCACCAAAUGCCAA ........(((((((..((((((......((..((........))..))...))))))..)))))))((((....))))..(((((.....(((((.....))))).....))))).... (-24.64 = -24.45 + -0.19)

| Location | 10,842,122 – 10,842,239 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.79 |
| Mean single sequence MFE | -34.50 |
| Consensus MFE | -19.48 |
| Energy contribution | -21.98 |
| Covariance contribution | 2.50 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.56 |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.543747 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
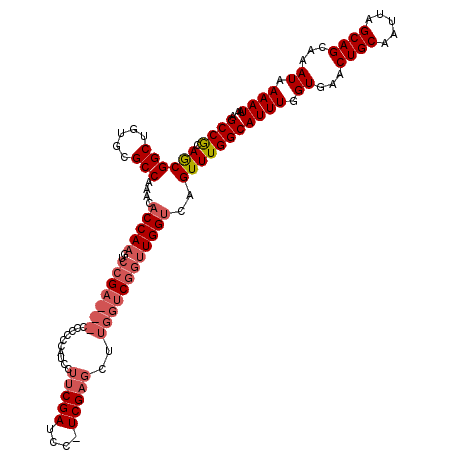
>X_DroMel_CAF1 10842122 117 - 22224390 AGCCGCAGCGGCUGUGCGCCAAACACCAAGCCCGA---ACCCUCAUCCUUCGAUCCUUCGAGCUUGGUCGGUUGGUCAGUUUGGCAUUUGGUGAACUGCAAUUAGCAGCAAAUAAAAUAA (((((...)))))...(((((((..((((((((.(---(((..(....(((((....)))))....)..))))))...))))))..)))))))..((((.....))))............ ( -36.60) >DroSec_CAF1 20925 119 - 1 AGCCGCAGCGGCUGUGCGCCAAACACCAAGUCCGACUACCCCCCAUCCUUCGAUCC-UCGAGCUUGGUCGGUUGGUCAGUUUGGCAUUUGGUGAACUGCAAUUAGCAGCAAAUAAAAUAA (((((...)))))...(((((((..(((((.(.(((((((..(((...((((....-.))))..)))..)).))))).))))))..)))))))..((((.....))))............ ( -39.50) >DroSim_CAF1 1787 119 - 1 AGCCGCAGCGGCUGUGCGCCAAACACCAAGUCCGACUACCCCCCAUCCUUCGAUCC-UCGAGCUUGGUCGGUUGGUCAGUUUGGCAUUUGGUGAACUGCAAUUAGCAGCAAAUAAAAUAA (((((...)))))...(((((((..(((((.(.(((((((..(((...((((....-.))))..)))..)).))))).))))))..)))))))..((((.....))))............ ( -39.50) >DroYak_CAF1 9914 93 - 1 AGCCACAAC----------------CCAACGGCGA----CCCCCUUUCUCCGAUCC-UCGA------CCAACUGGUCAGUUUGGCAUUUAGUGAACUGCAAUUAGCAGCAAAUAAAAUAA .((((.(((----------------....(((.((----.......)).)))....-..((------((....)))).)))))))((((.((...((((.....))))...)).)))).. ( -22.40) >consensus AGCCGCAGCGGCUGUGCGCCAAACACCAAGUCCGA___CCCCCCAUCCUUCGAUCC_UCGAGCUUGGUCGGUUGGUCAGUUUGGCAUUUGGUGAACUGCAAUUAGCAGCAAAUAAAAUAA .((((.((((((.....)))....(((((..(((((((..........(((((....)))))..))))))))))))..)))))))((((.((...((((.....))))...)).)))).. (-19.48 = -21.98 + 2.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:16:31 2006