
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 10,483,259 – 10,483,372 |
| Length | 113 |
| Max. P | 0.523187 |

| Location | 10,483,259 – 10,483,372 |
|---|---|
| Length | 113 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.58 |
| Mean single sequence MFE | -30.92 |
| Consensus MFE | -20.62 |
| Energy contribution | -23.73 |
| Covariance contribution | 3.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.67 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.523187 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
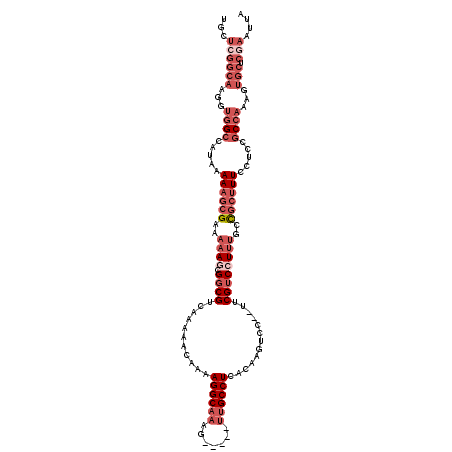
>X_DroMel_CAF1 10483259 113 + 22224390 UGCUCGGCAAGGUGGCCAUAAAAAGCGAAAAAGCGGCGUCAAAAACAAAAGGCAAAG-----UUGCCUCACAAGUCC--UUCGUCCUUUGCUGCUUUCCUCCGCCAAAGUGCUCGAAUUA ...((((((...((((.....((((((..((((.((((...........(((((...-----.))))).........--..))))))))..)))))).....))))...))).))).... ( -33.75) >DroSec_CAF1 14026 113 + 1 UGCUCGGCAAGGUGGCCAUAAAAAGCGAAAAAGCGGCGUCAAAAACAGAAGGCAAAG-----UUGCCUCACAAGUCC--UUCGUCCUUUGCCGCUUUCCUCCGCCAAAGUGCUCGAAUUA ...((((((...((((.....((((((..((((.((((...........(((((...-----.))))).........--..))))))))..)))))).....))))...))).))).... ( -35.55) >DroSim_CAF1 14215 112 + 1 UGCUCGGCAAGGUGGCCAUAAAAAGCGAAAAAGCGGCGUCAA-AACAAAAGGCAAAG-----UUGCCUCACAAGUCU--UUCGUCGUUUGCCGCUUUCCUCCGCCAAAGUGCUCGAAUUA ...((((((...((((.....((((((...((((((((....-......(((((...-----.))))).........--..))))))))..)))))).....))))...))).))).... ( -37.00) >DroEre_CAF1 15456 113 + 1 UACUCGGCAAGGUGGCCAUAAAAUGCGAAAAAGCGGCGUCAAAAACAAAAGGCAAAG-----UUGCCUCACAAGUCC--UUCGUCCUUUGCCGCUUUUCUCCGCCAAAGUGCUCAAAUUA .....((((...((((....(((.(((..((((.((((...........(((((...-----.))))).........--..))))))))..))).)))....))))...))))....... ( -29.85) >DroYak_CAF1 16171 113 + 1 UGCUCGGCAAGGUGGCCAUAAAAAGCGAAAAAGCGGCGUCAAAAACAAAAGGCAAAG-----UUGCCUCACAAGUCC--UUCGUCCUUUGCCGCUUUUCUCCGCCAAAGUGCUCGAAUUA ...((((((...((((....(((((((..((((.((((...........(((((...-----.))))).........--..))))))))..)))))))....))))...))).))).... ( -36.65) >DroAna_CAF1 14773 103 + 1 C----------CUGGCCAUAAAAACCGAAUUA-CGCCGACAAAAACAAAAGGCCAAAAACUUCUGCCUG-CCAGUCCUGUUCGUCCUUUUUUUUUUUUGUUUCC-----UUUUCAAAUUA .----------..(((..(((........)))-.)))((((((((.((((((.(.((.....(((....-.))).....)).).))))))...))))))))...-----........... ( -12.70) >consensus UGCUCGGCAAGGUGGCCAUAAAAAGCGAAAAAGCGGCGUCAAAAACAAAAGGCAAAG_____UUGCCUCACAAGUCC__UUCGUCCUUUGCCGCUUUCCUCCGCCAAAGUGCUCGAAUUA ...((((((...((((.....((((((..((((.((((...........((((((.......)))))).............))))))))..)))))).....))))...))).))).... (-20.62 = -23.73 + 3.11)


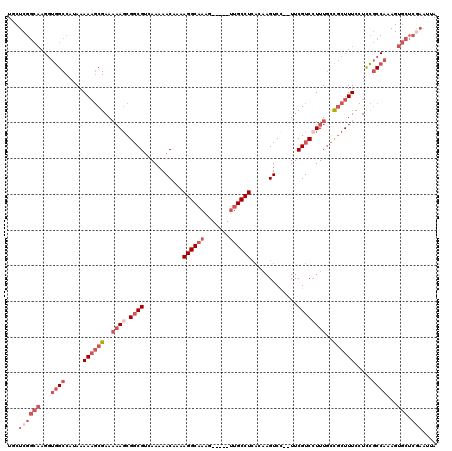
| Location | 10,483,259 – 10,483,372 |
|---|---|
| Length | 113 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.58 |
| Mean single sequence MFE | -35.47 |
| Consensus MFE | -24.71 |
| Energy contribution | -27.77 |
| Covariance contribution | 3.06 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.70 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 10483259 113 - 22224390 UAAUUCGAGCACUUUGGCGGAGGAAAGCAGCAAAGGACGAA--GGACUUGUGAGGCAA-----CUUUGCCUUUUGUUUUUGACGCCGCUUUUUCGCUUUUUAUGGCCACCUUGCCGAGCA ...((((.(((...((((.(..(((((((((...((.((((--((((....((((((.-----...))))))..))))))..))))))).....))))))..).))))...))))))).. ( -34.60) >DroSec_CAF1 14026 113 - 1 UAAUUCGAGCACUUUGGCGGAGGAAAGCGGCAAAGGACGAA--GGACUUGUGAGGCAA-----CUUUGCCUUCUGUUUUUGACGCCGCUUUUUCGCUUUUUAUGGCCACCUUGCCGAGCA ........((...((((((.(((((((((((.(((((((((--((.(....(((....-----))).)))))).))))))...))))))))...(((......)))..))))))))))). ( -40.90) >DroSim_CAF1 14215 112 - 1 UAAUUCGAGCACUUUGGCGGAGGAAAGCGGCAAACGACGAA--AGACUUGUGAGGCAA-----CUUUGCCUUUUGUU-UUGACGCCGCUUUUUCGCUUUUUAUGGCCACCUUGCCGAGCA ...((((.(((...((((.(..((((((((.(((((.((.(--((((....((((((.-----...))))))..)))-))..)).)).))).))))))))..).))))...))))))).. ( -39.30) >DroEre_CAF1 15456 113 - 1 UAAUUUGAGCACUUUGGCGGAGAAAAGCGGCAAAGGACGAA--GGACUUGUGAGGCAA-----CUUUGCCUUUUGUUUUUGACGCCGCUUUUUCGCAUUUUAUGGCCACCUUGCCGAGUA ..(((((.(((...((((.((((((.(((((.(((((((((--((.(....(((....-----))).).)))))))))))...))))))))))).(((...)))))))...)))))))). ( -39.00) >DroYak_CAF1 16171 113 - 1 UAAUUCGAGCACUUUGGCGGAGAAAAGCGGCAAAGGACGAA--GGACUUGUGAGGCAA-----CUUUGCCUUUUGUUUUUGACGCCGCUUUUUCGCUUUUUAUGGCCACCUUGCCGAGCA ...((((.(((...((((.(.(((((((((.(((((.((((--((((....((((((.-----...))))))..))))))..)).).)))).))))))))).).))))...))))))).. ( -39.30) >DroAna_CAF1 14773 103 - 1 UAAUUUGAAAA-----GGAAACAAAAAAAAAAAAGGACGAACAGGACUGG-CAGGCAGAAGUUUUUGGCCUUUUGUUUUUGUCGGCG-UAAUUCGGUUUUUAUGGCCAG----------G ......((...-----((((((((((.......((..(.....)..))((-(.(((....)))....))))))))))))).))((((-(((........)))).)))..----------. ( -19.70) >consensus UAAUUCGAGCACUUUGGCGGAGAAAAGCGGCAAAGGACGAA__GGACUUGUGAGGCAA_____CUUUGCCUUUUGUUUUUGACGCCGCUUUUUCGCUUUUUAUGGCCACCUUGCCGAGCA ...((((.(((...((((.(.(((((((((.(((((.((....((((....(((((((.......)))))))..))))....)).).)))).))))))))).).))))...))))))).. (-24.71 = -27.77 + 3.06)

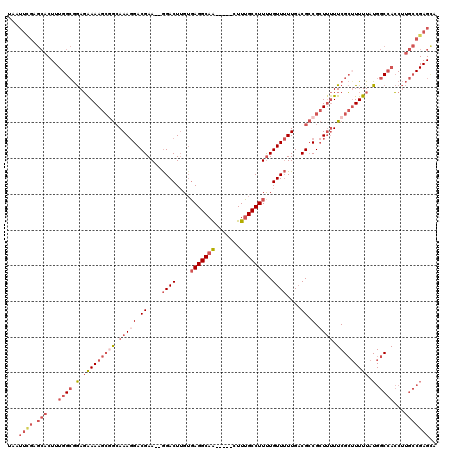
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:11:58 2006