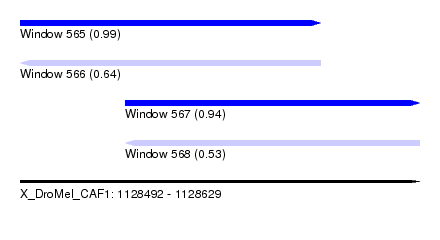

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 1,128,492 – 1,128,629 |
| Length | 137 |
| Max. P | 0.987851 |

| Location | 1,128,492 – 1,128,595 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 75.89 |
| Mean single sequence MFE | -20.16 |
| Consensus MFE | -13.86 |
| Energy contribution | -13.46 |
| Covariance contribution | -0.40 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.69 |
| SVM decision value | 2.10 |
| SVM RNA-class probability | 0.987851 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
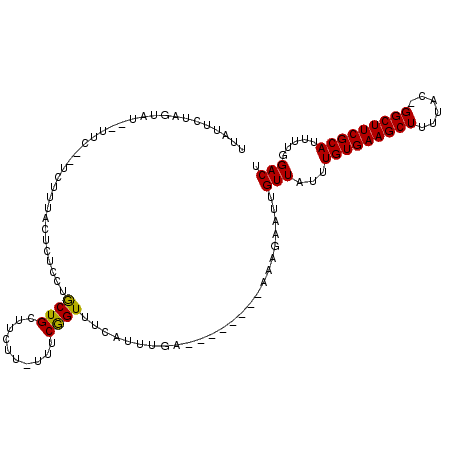
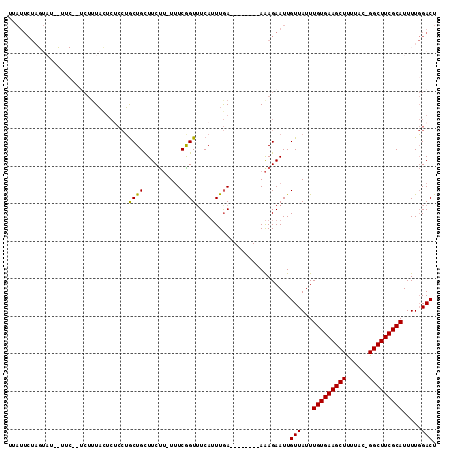
>X_DroMel_CAF1 1128492 103 + 22224390 UUAUUCUAGUAU-UUUCUCUCUUUACUCUCCUGCUGCUUCUU-UUUCGGUAUCAUUUGA--------AAAGAAUUGUUAUUUGUGAAGCUUUUAC-GGCUUCGCAUUUUGGACU ............-...............(((......(((((-((.(((......))))--------))))))........(((((((((.....-)))))))))....))).. ( -18.60) >DroSim_CAF1 26553 103 + 1 UUAUUCUAGUAU-UUUCUCUCUUUACUCUCCUGCUGCUUCUU-UUUCGGUUUCAUUUGA--------AAAGAAUUGUUAUUUGUGAAGCUUUUAC-GGCUUCGCAUUUUGGACU ............-...............(((......(((((-((.(((......))))--------))))))........(((((((((.....-)))))))))....))).. ( -18.60) >DroEre_CAF1 26359 105 + 1 UUAUUCUAGUAUACUUCGGUCUGUAUUCGCCUGCUGCUUCUUUUUUCAGUUUCAUUUGA--------AAAGAAUUGUUAUUUGUGAAGCUUUUAC-GGCUUCGCAUUUUGGACU .................(((((..................(((((((((......))))--------))))).........(((((((((.....-)))))))))....))))) ( -21.90) >DroYak_CAF1 25965 99 + 1 CGAUUCU-------------CUUUAUUCGCCUGCUGCUUCUU-UUUCAGUUUCAUUUGAACAUUUGGAAAGAAUUGUUAUUUGUGAAGCUUUUAC-GGCUUCGCAUUUUGGACU (((((((-------------.((((((((..((..(((....-....)))..))..))))....)))).))))))).....(((((((((.....-)))))))))......... ( -20.60) >DroAna_CAF1 48895 79 + 1 ----------------------UUC---UAAGACUGUUUCC--UCUCGGUUUCAGUUAA--------AGAGAAUUGUUAUUUGUGAAGCUCUUACUGGCUUCGCAUUUUCGACU ----------------------(((---(..(((((...((--....))...)))))..--------))))..(((.....(((((((((......)))))))))....))).. ( -21.10) >consensus UUAUUCUAGUAU__UUC__UCUUUACUCUCCUGCUGCUUCUU_UUUCGGUUUCAUUUGA________AAAGAAUUGUUAUUUGUGAAGCUUUUAC_GGCUUCGCAUUUUGGACU ................................((((..........)))).........................(((...(((((((((......))))))))).....))). (-13.86 = -13.46 + -0.40)

| Location | 1,128,492 – 1,128,595 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 75.89 |
| Mean single sequence MFE | -15.42 |
| Consensus MFE | -10.11 |
| Energy contribution | -10.03 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.635985 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
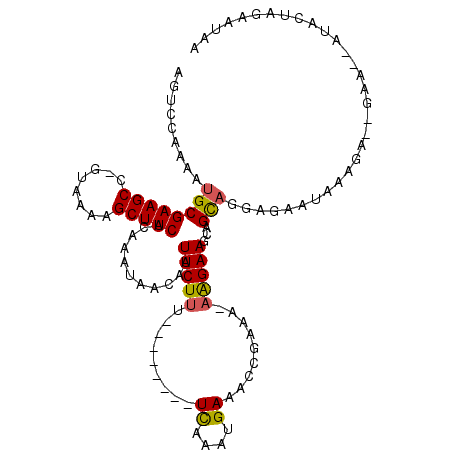
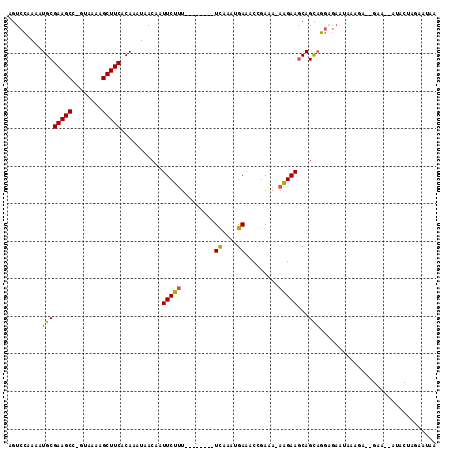
>X_DroMel_CAF1 1128492 103 - 22224390 AGUCCAAAAUGCGAAGCC-GUAAAAGCUUCACAAAUAACAAUUCUUU--------UCAAAUGAUACCGAAA-AAGAAGCAGCAGGAGAGUAAAGAGAGAAA-AUACUAGAAUAA ..(((....((((((((.-......)))))...........((((((--------((..........).))-))))))))...)))...............-............ ( -14.90) >DroSim_CAF1 26553 103 - 1 AGUCCAAAAUGCGAAGCC-GUAAAAGCUUCACAAAUAACAAUUCUUU--------UCAAAUGAAACCGAAA-AAGAAGCAGCAGGAGAGUAAAGAGAGAAA-AUACUAGAAUAA ..(((....((((((((.-......)))))...........((((((--------((..........).))-))))))))...)))...............-............ ( -14.90) >DroEre_CAF1 26359 105 - 1 AGUCCAAAAUGCGAAGCC-GUAAAAGCUUCACAAAUAACAAUUCUUU--------UCAAAUGAAACUGAAAAAAGAAGCAGCAGGCGAAUACAGACCGAAGUAUACUAGAAUAA .(((.....((((((((.-......)))))...........((((((--------(((........)))..)))))))))...)))..((((........)))).......... ( -16.10) >DroYak_CAF1 25965 99 - 1 AGUCCAAAAUGCGAAGCC-GUAAAAGCUUCACAAAUAACAAUUCUUUCCAAAUGUUCAAAUGAAACUGAAA-AAGAAGCAGCAGGCGAAUAAAG-------------AGAAUCG .(((.....((((((((.-......)))))...........((((((.......(((....)))......)-))))))))...)))........-------------....... ( -15.42) >DroAna_CAF1 48895 79 - 1 AGUCGAAAAUGCGAAGCCAGUAAGAGCUUCACAAAUAACAAUUCUCU--------UUAACUGAAACCGAGA--GGAAACAGUCUUA---GAA---------------------- .........((.(((((........))))).))...........(((--------...((((...((....--))...))))...)---)).---------------------- ( -15.80) >consensus AGUCCAAAAUGCGAAGCC_GUAAAAGCUUCACAAAUAACAAUUCUUU________UCAAAUGAAACCGAAA_AAGAAGCAGCAGGAGAAUAAAGA__GAA__AUACUAGAAUAA .........((((((((........)))))...........(((((.........((....)).........)))))...)))............................... (-10.11 = -10.03 + -0.08)

| Location | 1,128,528 – 1,128,629 |
|---|---|
| Length | 101 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 93.42 |
| Mean single sequence MFE | -28.00 |
| Consensus MFE | -24.15 |
| Energy contribution | -24.82 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.25 |
| SVM RNA-class probability | 0.935698 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1128528 101 + 22224390 UUCUU-UUUCGGUAUCAUUUGA--------AAAGAAUUGUUAUUUGUGAAGCUUUUAC-GGCUUCGCAUUUUGGACUGUCGACAAGUUGACAAAUGGCAAACCGGUUUGGC ..((.-..(((((....(((..--------..))).(((((((((((((((((.....-))))))))..........(((((....)))))))))))))))))))...)). ( -28.00) >DroSec_CAF1 25160 101 + 1 UUCUU-UUUCGGUUUCAUUUGA--------AAAGAAUUGUUAUUUGUGAAGCUUUUAC-GGCUUCGCAUUUUGGACUGUCGACAAGUUGACAAAUGGCAAACCGGUUUGGC ..((.-..((((((((((((..--------........(((...(((((((((.....-))))))))).....))).(((((....)))))))))))..))))))...)). ( -27.60) >DroSim_CAF1 26589 101 + 1 UUCUU-UUUCGGUUUCAUUUGA--------AAAGAAUUGUUAUUUGUGAAGCUUUUAC-GGCUUCGCAUUUUGGACUGUCGACAAGUUGACAAAUGGCAAACCGGUUUGGC ..((.-..((((((((((((..--------........(((...(((((((((.....-))))))))).....))).(((((....)))))))))))..))))))...)). ( -27.60) >DroEre_CAF1 26396 102 + 1 UUCUUUUUUCAGUUUCAUUUGA--------AAAGAAUUGUUAUUUGUGAAGCUUUUAC-GGCUUCGCAUUUUGGACUGUCGACAAGUUGACAAAUGGCAAACCGGUUUGGC ...(((((((((......))))--------))))).(((((((((((((((((.....-))))))))..........(((((....))))))))))))))........... ( -28.10) >DroYak_CAF1 25989 109 + 1 UUCUU-UUUCAGUUUCAUUUGAACAUUUGGAAAGAAUUGUUAUUUGUGAAGCUUUUAC-GGCUUCGCAUUUUGGACUGUCGACAAGUUGACAAAUGGCAAACCGGUUUGGC (((((-((..((((((....))).)))..)))))))(((((((((((((((((.....-))))))))..........(((((....))))))))))))))........... ( -28.50) >DroAna_CAF1 48907 101 + 1 UUCC--UCUCGGUUUCAGUUAA--------AGAGAAUUGUUAUUUGUGAAGCUCUUACUGGCUUCGCAUUUUCGACUGUCGACAAGUUGACAAAUGGCAAACCGGUUUGGC ..((--..((((((((......--------.)))).(((((((((((((((((......))))))))....((((((.......)))))).))))))))).))))...)). ( -28.20) >consensus UUCUU_UUUCGGUUUCAUUUGA________AAAGAAUUGUUAUUUGUGAAGCUUUUAC_GGCUUCGCAUUUUGGACUGUCGACAAGUUGACAAAUGGCAAACCGGUUUGGC ........((((((((((((..................(((...(((((((((......))))))))).....))).(((((....)))))))))))..))))))...... (-24.15 = -24.82 + 0.67)

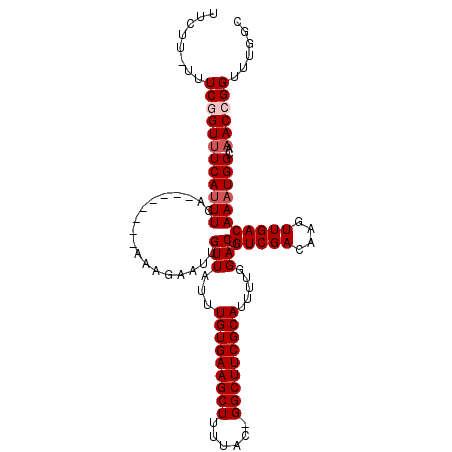
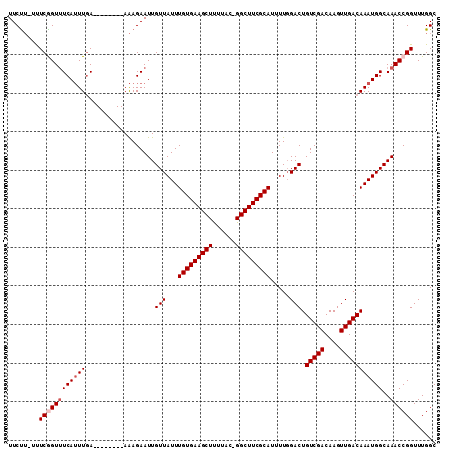
| Location | 1,128,528 – 1,128,629 |
|---|---|
| Length | 101 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 93.42 |
| Mean single sequence MFE | -17.02 |
| Consensus MFE | -16.92 |
| Energy contribution | -16.87 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.06 |
| Mean z-score | -0.66 |
| Structure conservation index | 0.99 |
| SVM decision value | 0.00 |
| SVM RNA-class probability | 0.534377 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 1128528 101 - 22224390 GCCAAACCGGUUUGCCAUUUGUCAACUUGUCGACAGUCCAAAAUGCGAAGCC-GUAAAAGCUUCACAAAUAACAAUUCUUU--------UCAAAUGAUACCGAAA-AAGAA .......((((....((((((((........))).........((.(((((.-......))))).))..............--------..)))))..))))...-..... ( -16.40) >DroSec_CAF1 25160 101 - 1 GCCAAACCGGUUUGCCAUUUGUCAACUUGUCGACAGUCCAAAAUGCGAAGCC-GUAAAAGCUUCACAAAUAACAAUUCUUU--------UCAAAUGAAACCGAAA-AAGAA .......((((((..((((((((........))).........((.(((((.-......))))).))..............--------..)))))))))))...-..... ( -17.90) >DroSim_CAF1 26589 101 - 1 GCCAAACCGGUUUGCCAUUUGUCAACUUGUCGACAGUCCAAAAUGCGAAGCC-GUAAAAGCUUCACAAAUAACAAUUCUUU--------UCAAAUGAAACCGAAA-AAGAA .......((((((..((((((((........))).........((.(((((.-......))))).))..............--------..)))))))))))...-..... ( -17.90) >DroEre_CAF1 26396 102 - 1 GCCAAACCGGUUUGCCAUUUGUCAACUUGUCGACAGUCCAAAAUGCGAAGCC-GUAAAAGCUUCACAAAUAACAAUUCUUU--------UCAAAUGAAACUGAAAAAAGAA .......((((((..((((((((........))).........((.(((((.-......))))).))..............--------..)))))))))))......... ( -15.70) >DroYak_CAF1 25989 109 - 1 GCCAAACCGGUUUGCCAUUUGUCAACUUGUCGACAGUCCAAAAUGCGAAGCC-GUAAAAGCUUCACAAAUAACAAUUCUUUCCAAAUGUUCAAAUGAAACUGAAA-AAGAA .......((((((..((((((.((..(((....))).......((.(((((.-......))))).))...................))..))))))))))))...-..... ( -15.80) >DroAna_CAF1 48907 101 - 1 GCCAAACCGGUUUGCCAUUUGUCAACUUGUCGACAGUCGAAAAUGCGAAGCCAGUAAGAGCUUCACAAAUAACAAUUCUCU--------UUAACUGAAACCGAGA--GGAA .((....((((((.....(((((........))))).......((.(((((........))))).))..............--------.......))))))...--)).. ( -18.40) >consensus GCCAAACCGGUUUGCCAUUUGUCAACUUGUCGACAGUCCAAAAUGCGAAGCC_GUAAAAGCUUCACAAAUAACAAUUCUUU________UCAAAUGAAACCGAAA_AAGAA .......((((((.....(((((........))))).......((.(((((........))))).)).............................))))))......... (-16.92 = -16.87 + -0.06)

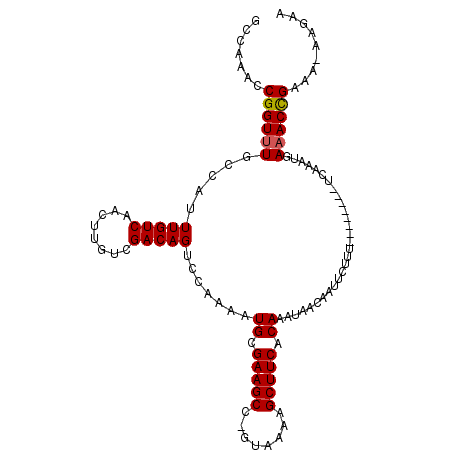
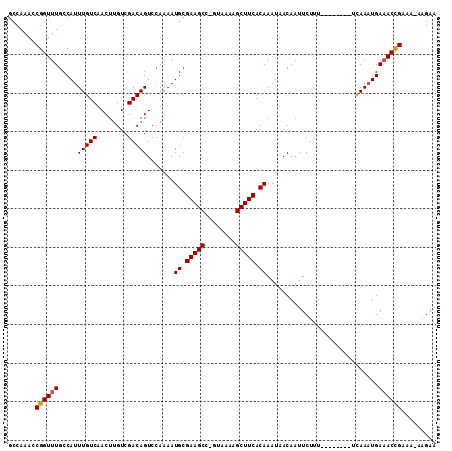
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:42:57 2006