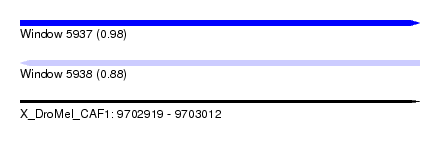

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,702,919 – 9,703,012 |
| Length | 93 |
| Max. P | 0.984915 |

| Location | 9,702,919 – 9,703,012 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 89.85 |
| Mean single sequence MFE | -39.76 |
| Consensus MFE | -30.40 |
| Energy contribution | -30.60 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.89 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.99 |
| SVM RNA-class probability | 0.984915 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9702919 93 + 22224390 CUGGCUAGCAGGUUAGCUCCUGGCUGGCAAGCCGGAUGUUAACGGCCAGCAGCUGCAGUUGCAGCAGGAAGUGGCCA--AAAAAUCAG--GGCGCAU ((((((.....((((((..((((((....))))))..))))))))))))..(((((....))))).....(((.((.--........)--).))).. ( -39.60) >DroSec_CAF1 44387 93 + 1 CUGGCUACCAGGAUAGCUCCUGGCUGGCAAGCCGGAUGUUAACGGCCAGCAGCUGCAGUUGCAGCAGGAAGUGGCCA--AAAAAUCAG--GGCGCAU ...((((((((((....)))))).))))..(((.(((......(((((.(.(((((....)))))..)...))))).--....)))..--))).... ( -38.50) >DroSim_CAF1 18817 93 + 1 CUGGCUACCAGGAUAGCUCCUGGCUGGCAAGCCGGAUGUUAACGGCCAGCAGCUGCAGUUGCAGCAGGAAGUGGCCA--AAAAAUCCG--GGCGCAU ...((((((((((....)))))).))))...((((((......(((((.(.(((((....)))))..)...))))).--....)))))--)...... ( -41.20) >DroEre_CAF1 21781 91 + 1 CUGGCC---A-GUUAGCUGCUGGCUGGCAAGCCGGAUGUUAACGGCCAGCAGCUGCAGUUGCAGCAGGAAGUGGCCAGGAAAAAUCGGG-GGCACA- ((((((---.-((((((..((((((....))))))..))))))))))))..(((((....))))).....(((.((..((....))..)-).))).- ( -45.20) >DroYak_CAF1 20762 92 + 1 CUGGCC---UGGGUUGCUCCUGGCUGGCAAGCCGGAUGUUGACGGCCAGCAGCUACAGUUGCAACAGGAAGUGGCCA--AAAAAUCGGGUGGCGCAU ...(((---(((((..((((((((((((.(((.....)))....)))))).((.......))..)))).))..))).--......)))))....... ( -34.31) >consensus CUGGCUA_CAGGAUAGCUCCUGGCUGGCAAGCCGGAUGUUAACGGCCAGCAGCUGCAGUUGCAGCAGGAAGUGGCCA__AAAAAUCAG__GGCGCAU ...(((.....(.((((..((((((....))))))..)))).)(((((.(.(((((....)))))..)...)))))..............))).... (-30.40 = -30.60 + 0.20)

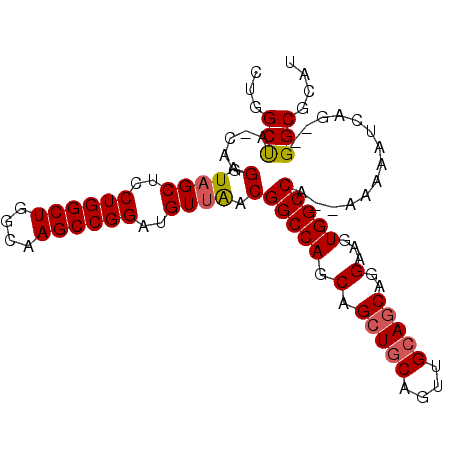
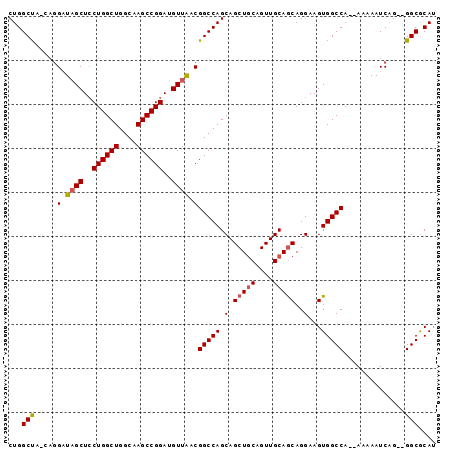
| Location | 9,702,919 – 9,703,012 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 89.85 |
| Mean single sequence MFE | -35.22 |
| Consensus MFE | -25.38 |
| Energy contribution | -25.10 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.56 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.879523 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9702919 93 - 22224390 AUGCGCC--CUGAUUUUU--UGGCCACUUCCUGCUGCAACUGCAGCUGCUGGCCGUUAACAUCCGGCUUGCCAGCCAGGAGCUAACCUGCUAGCCAG ....(((--..(((((..--((((((......(((((....)))))...))))))..)).))).)))..((.(((.(((......)))))).))... ( -31.50) >DroSec_CAF1 44387 93 - 1 AUGCGCC--CUGAUUUUU--UGGCCACUUCCUGCUGCAACUGCAGCUGCUGGCCGUUAACAUCCGGCUUGCCAGCCAGGAGCUAUCCUGGUAGCCAG ..(((((--..(((((..--((((((......(((((....)))))...))))))..)).))).)))......(((((((....))))))).))... ( -36.70) >DroSim_CAF1 18817 93 - 1 AUGCGCC--CGGAUUUUU--UGGCCACUUCCUGCUGCAACUGCAGCUGCUGGCCGUUAACAUCCGGCUUGCCAGCCAGGAGCUAUCCUGGUAGCCAG ..(((((--(((((((..--((((((......(((((....)))))...))))))..)).))))))...))..(((((((....))))))).))... ( -41.00) >DroEre_CAF1 21781 91 - 1 -UGUGCC-CCCGAUUUUUCCUGGCCACUUCCUGCUGCAACUGCAGCUGCUGGCCGUUAACAUCCGGCUUGCCAGCCAGCAGCUAAC-U---GGCCAG -......-...........(((((((.........((....))((((((((((.(....)....((....)).))))))))))...-)---)))))) ( -34.70) >DroYak_CAF1 20762 92 - 1 AUGCGCCACCCGAUUUUU--UGGCCACUUCCUGUUGCAACUGUAGCUGCUGGCCGUCAACAUCCGGCUUGCCAGCCAGGAGCAACCCA---GGCCAG .((.(((..((((....)--))).........(((((..(((..(((((.(((((........))))).).)))))))..)))))...---))))). ( -32.20) >consensus AUGCGCC__CCGAUUUUU__UGGCCACUUCCUGCUGCAACUGCAGCUGCUGGCCGUUAACAUCCGGCUUGCCAGCCAGGAGCUAACCUG_UAGCCAG ..((.((....(((.((...((((((......(((((....)))))...))))))..)).))).((((....)))).)).))............... (-25.38 = -25.10 + -0.28)

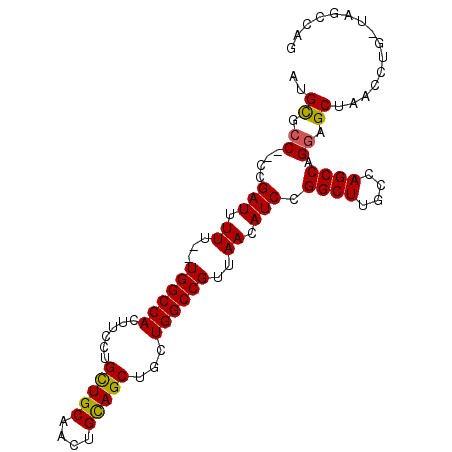
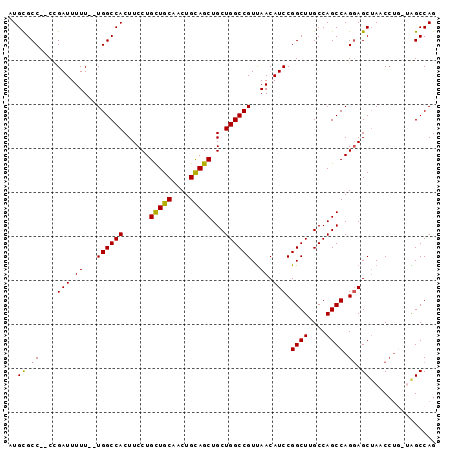
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:04:07 2006