
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,668,679 – 9,668,795 |
| Length | 116 |
| Max. P | 0.602647 |

| Location | 9,668,679 – 9,668,795 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 83.76 |
| Mean single sequence MFE | -22.83 |
| Consensus MFE | -16.35 |
| Energy contribution | -17.48 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.602647 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9668679 116 + 22224390 UAUUUCGUGAUAUAUAUACAAUAUUUUUAUCGCACAAAAUGCAAUGUUUGAAAGCUAAAAGCAAAUUCACGAGCUCUUUUCGAAUCCUUCGAGAUUUCUUUGCCUCUAUGAACACU ...(((((((.....................(((.....)))..(((((.........)))))...)))))))....((((((.....))))))...................... ( -15.80) >DroSec_CAF1 4279 114 + 1 --UUUCGUGAUAAAUAUACAAUAUUUUUAUCGCACAAAAUGCAUUGUUUGAAAGCUAGAGGCAAAUUCAUGAGCUCUUUUUGAAUCCUUCGGGAUUUCUUUGCCUCUAUAAACACU --....((((((((.(((...))).))))))))...........((((((.....((((((((((.....(....).....((((((....)))))).)))))))))))))))).. ( -29.30) >DroSim_CAF1 17859 116 + 1 UAUUUCGUGAUAAAUAUACAAUAUUUUUAUCGCACAAAAUGCAAUGUUUGAAAGCUAGAGGCAAAUUCAUGAGCUCUUUUUGAAUCCUUCCGGAUUUCUUUGCCUCUAUGAACACU ......((((((((.(((...))).))))))))...........((((((.....((((((((((.....(....).....((((((....)))))).)))))))))))))))).. ( -25.10) >DroEre_CAF1 11156 111 + 1 UA-----UUAUAUAUGCGCCAUAUUUGUAUUACCCAAAAUGCAAUGUUCGAAAUCUAAGAGCGAAUUCACGAGGUCAUUUCUAAUCCUGCGGGUUUUCUUUGCCUUUGUGAACACU ..-----.................(((((((......)))))))(((((.........)))))..(((((((((.((.....(((((...))))).....)).))))))))).... ( -21.10) >consensus UAUUUCGUGAUAAAUAUACAAUAUUUUUAUCGCACAAAAUGCAAUGUUUGAAAGCUAAAAGCAAAUUCACGAGCUCUUUUCGAAUCCUUCGGGAUUUCUUUGCCUCUAUGAACACU ......((((((((...........))))))))...........((((((.....((((((((((.....(((.....)))((((((....)))))).)))))))))))))))).. (-16.35 = -17.48 + 1.12)

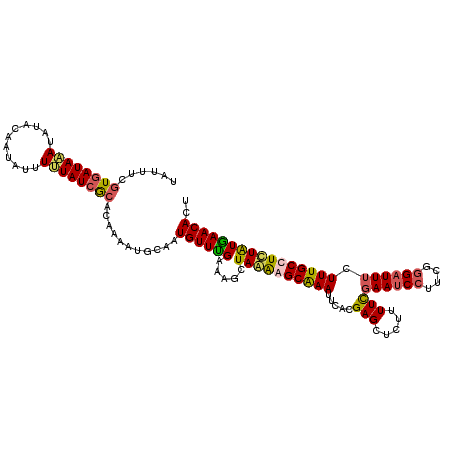
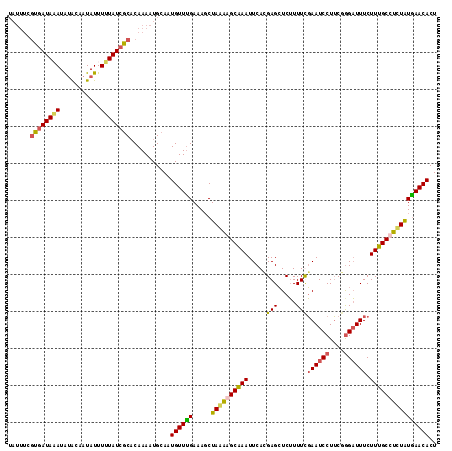
| Location | 9,668,679 – 9,668,795 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 83.76 |
| Mean single sequence MFE | -24.08 |
| Consensus MFE | -18.56 |
| Energy contribution | -19.25 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.77 |
| SVM decision value | -0.00 |
| SVM RNA-class probability | 0.531171 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9668679 116 - 22224390 AGUGUUCAUAGAGGCAAAGAAAUCUCGAAGGAUUCGAAAAGAGCUCGUGAAUUUGCUUUUAGCUUUCAAACAUUGCAUUUUGUGCGAUAAAAAUAUUGUAUAUAUAUCACGAAAUA .(((....((((((((((....((.(((..(.(((.....))))))).)).))))))))))((...........))....(((((((((....))))))))).....)))...... ( -21.30) >DroSec_CAF1 4279 114 - 1 AGUGUUUAUAGAGGCAAAGAAAUCCCGAAGGAUUCAAAAAGAGCUCAUGAAUUUGCCUCUAGCUUUCAAACAAUGCAUUUUGUGCGAUAAAAAUAUUGUAUAUUUAUCACGAAA-- .(((....(((((((((((((.(((....)))))).....(....).....))))))))))((...........))....(((((((((....))))))))).....)))....-- ( -25.70) >DroSim_CAF1 17859 116 - 1 AGUGUUCAUAGAGGCAAAGAAAUCCGGAAGGAUUCAAAAAGAGCUCAUGAAUUUGCCUCUAGCUUUCAAACAUUGCAUUUUGUGCGAUAAAAAUAUUGUAUAUUUAUCACGAAAUA .(((....(((((((((((((.(((....)))))).....(....).....))))))))))((...........))....(((((((((....))))))))).....)))...... ( -29.90) >DroEre_CAF1 11156 111 - 1 AGUGUUCACAAAGGCAAAGAAAACCCGCAGGAUUAGAAAUGACCUCGUGAAUUCGCUCUUAGAUUUCGAACAUUGCAUUUUGGGUAAUACAAAUAUGGCGCAUAUAUAA-----UA .((((((.(((((((((........((.(((.(((....))))))))((((.((.......)).))))....)))).))))).).)))))..(((((...)))))....-----.. ( -19.40) >consensus AGUGUUCAUAGAGGCAAAGAAAUCCCGAAGGAUUCAAAAAGAGCUCAUGAAUUUGCCUCUAGCUUUCAAACAUUGCAUUUUGUGCGAUAAAAAUAUUGUAUAUAUAUCACGAAAUA (((((((.((((((((((..(((((....)))))......(....).....))))))))))......)))))))......(((((((((....))))))))).............. (-18.56 = -19.25 + 0.69)

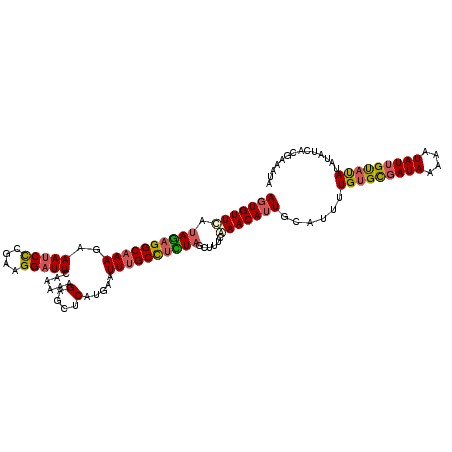
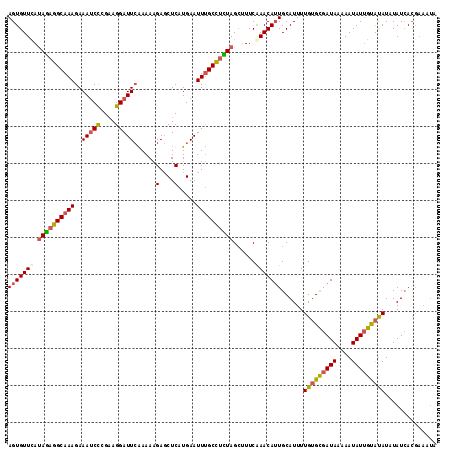
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:03:43 2006