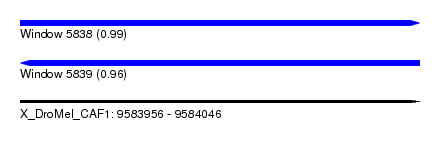

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,583,956 – 9,584,046 |
| Length | 90 |
| Max. P | 0.992322 |

| Location | 9,583,956 – 9,584,046 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 85.24 |
| Mean single sequence MFE | -33.46 |
| Consensus MFE | -26.06 |
| Energy contribution | -26.50 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.87 |
| Structure conservation index | 0.78 |
| SVM decision value | 2.32 |
| SVM RNA-class probability | 0.992322 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
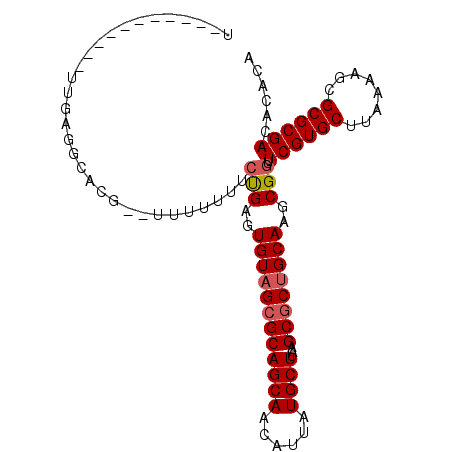
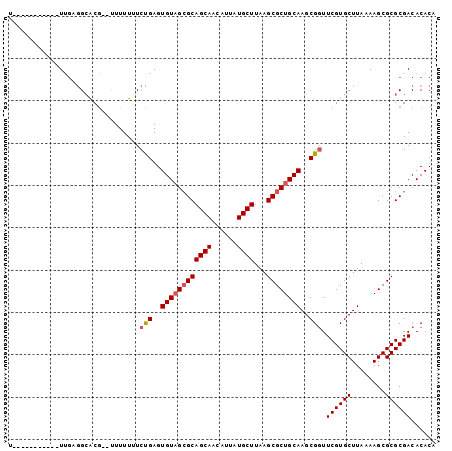
>X_DroMel_CAF1 9583956 90 + 22224390 U-----------UCGAGGCACAUAUUUUUGUCUGAGUGUAGCGCAGCAACAUUAUGCUUAAGCGCUGCAAGCGCUUCGUGCUUAAAAGCGCGCGACACACA .-----------((.(((((........)))))))((((..(((((((......))))...(((((..((((((...))))))...))))))))..)))). ( -32.50) >DroSec_CAF1 42813 88 + 1 U-----------UUGUGGCACG--UUUUUUUCUGAGUGUAGCGCAGCAACAUUAUGCUUAAGCGCUGCAAGCGGUUCGUGCUUAAAAGCGCGCGACACACA .-----------.(((.((...--.......(((..((((((((((((......))))...))))))))..)))...(((((....))))))).))).... ( -32.80) >DroSim_CAF1 29317 88 + 1 U-----------UUGUGGCACG--UUUUUGUCUGAGUGUAGCGCAGCAACAUUAUGCUUAAGCGCUGCAAGCGGUUCGUGCUUAAAAGCGCGCGACACACA .-----------.(((.((.((--((((((.(((..((((((((((((......))))...))))))))..)))........)))))))).)).))).... ( -33.50) >DroEre_CAF1 47646 99 + 1 UCAAAUGCCACAUUGAGGCACGG-CUUUUUUCCGAGUGUAGAGCAGCAACAUUAUGCUUAAGCGCUGCAAGCGGUUCGUGCUUAAAAGCGCGCGACACAC- .....((((.......))))(((-.......))).((((...((((((......))))...(((((..(((((.....)))))...))))))).))))..- ( -30.10) >DroYak_CAF1 45081 99 + 1 UCAAAUGCCUCAUUUGAGGGCGU-UUUUUUUCUGAGUGUCGCGCAGCAACAUUAUGCUUAAGCGCUGCAAGCGGUUCGUGCUUAAAAGCGCGCGACACAC- ..(((((((((....)).)))))-)).........(((((((((.(((......)))((((((((.((.....))..))))))))....)))))))))..- ( -38.40) >consensus U___________UUGAGGCACG__UUUUUUUCUGAGUGUAGCGCAGCAACAUUAUGCUUAAGCGCUGCAAGCGGUUCGUGCUUAAAAGCGCGCGACACACA ...............................(((..((((((((((((......))))...))))))))..))).((((((........))))))...... (-26.06 = -26.50 + 0.44)

| Location | 9,583,956 – 9,584,046 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 85.24 |
| Mean single sequence MFE | -29.56 |
| Consensus MFE | -25.20 |
| Energy contribution | -25.60 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.50 |
| SVM RNA-class probability | 0.959128 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
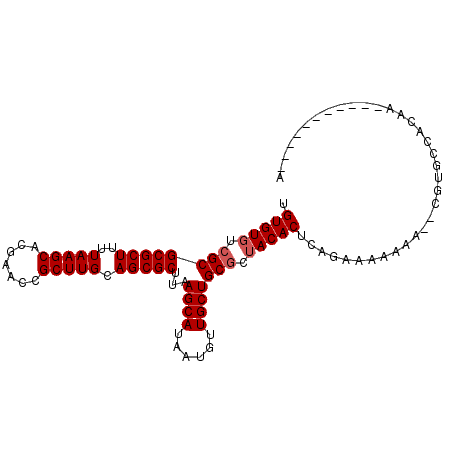
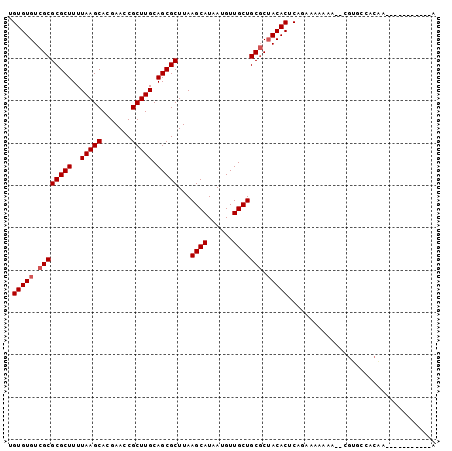
>X_DroMel_CAF1 9583956 90 - 22224390 UGUGUGUCGCGCGCUUUUAAGCACGAAGCGCUUGCAGCGCUUAAGCAUAAUGUUGCUGCGCUACACUCAGACAAAAAUAUGUGCCUCGA-----------A ((.((((.(((((((....)))...((((((.....)))))).((((......)))))))).)))).))....................-----------. ( -28.90) >DroSec_CAF1 42813 88 - 1 UGUGUGUCGCGCGCUUUUAAGCACGAACCGCUUGCAGCGCUUAAGCAUAAUGUUGCUGCGCUACACUCAGAAAAAAA--CGUGCCACAA-----------A .(((((.((((((((..(((((.......))))).)))))...((((......))))))).)))))...........--..........-----------. ( -26.40) >DroSim_CAF1 29317 88 - 1 UGUGUGUCGCGCGCUUUUAAGCACGAACCGCUUGCAGCGCUUAAGCAUAAUGUUGCUGCGCUACACUCAGACAAAAA--CGUGCCACAA-----------A .(((((.((((((((..(((((.......))))).)))))...((((......))))))).)))))...........--..........-----------. ( -26.40) >DroEre_CAF1 47646 99 - 1 -GUGUGUCGCGCGCUUUUAAGCACGAACCGCUUGCAGCGCUUAAGCAUAAUGUUGCUGCUCUACACUCGGAAAAAAG-CCGUGCCUCAAUGUGGCAUUUGA -(((((..(((((((..(((((.......))))).)))))...((((......))))))..))))).(((.......-)))((((.......))))..... ( -30.30) >DroYak_CAF1 45081 99 - 1 -GUGUGUCGCGCGCUUUUAAGCACGAACCGCUUGCAGCGCUUAAGCAUAAUGUUGCUGCGCGACACUCAGAAAAAAA-ACGCCCUCAAAUGAGGCAUUUGA -(.((((((((((((..(((((.......))))).))).....((((......))))))))))))).).........-..(((.((....)))))...... ( -35.80) >consensus UGUGUGUCGCGCGCUUUUAAGCACGAACCGCUUGCAGCGCUUAAGCAUAAUGUUGCUGCGCUACACUCAGAAAAAAA__CGUGCCACAA___________A .(((((.((((((((..(((((.......))))).)))))...((((......))))))).)))))................................... (-25.20 = -25.60 + 0.40)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:02:37 2006