| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,132,547 – 9,132,640 |
| Length | 93 |
| Max. P | 0.999995 |

| Location | 9,132,547 – 9,132,640 |
|---|---|
| Length | 93 |
| Sequences | 3 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 68.20 |
| Mean single sequence MFE | -23.41 |
| Consensus MFE | -14.91 |
| Energy contribution | -16.13 |
| Covariance contribution | 1.23 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.71 |
| Structure conservation index | 0.64 |
| SVM decision value | 5.90 |
| SVM RNA-class probability | 0.999995 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
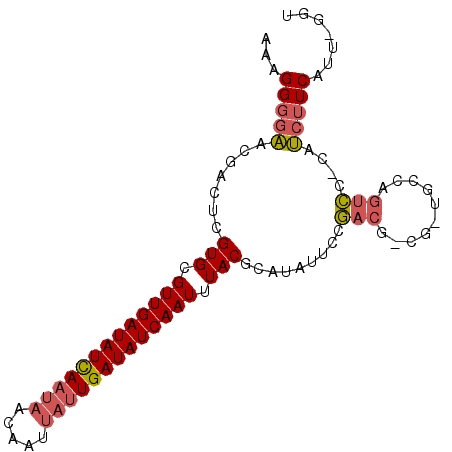
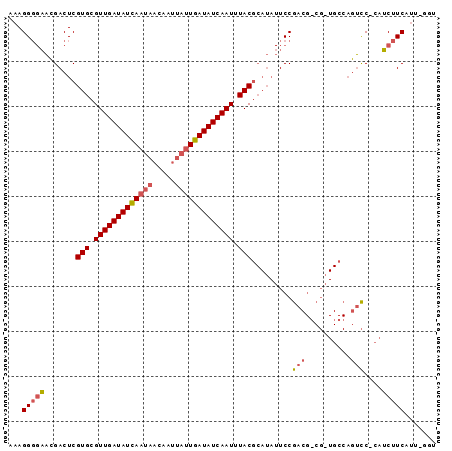
>X_DroMel_CAF1 9132547 93 - 22224390 AAAGGGGAACGACUCGUGCGUUGAUAUCAAUAACAAUUAUUGAUAUCAAUUUACGCAUAUUCCGACGGCGAUGCCAGUCC-CAUCUUCACU-GGC ...((((((.....((((.(((((((((((((.....))))))))))))).))))....))))((((((...))).))))-).........-... ( -28.90) >DroVir_CAF1 606 77 - 1 CAAGG--CGAGACUCGUGCGUUGAUAUUAAAACCAAUAACUGAUAUCAAUUUACU-----UCCAU-----------UCUCCUCGCAUCAUCGAAU ....(--((((....(((.((((((((((...........)))))))))).))).-----.....-----------....))))).......... ( -14.14) >DroSec_CAF1 1393 93 - 1 AAAGGGGAACGACUCGUGCGUUGAUAUCAAUAACAAUUAUUGAUAUCAAUUUACGCAUAUUCCGACGCCGUUGCCAGUCC-CAUCUUCAUU-GGU ....((.((((.((((((((((((((((((((.....)))))))))))....))))).....))).).)))).))....(-((.......)-)). ( -27.20) >consensus AAAGGGGAACGACUCGUGCGUUGAUAUCAAUAACAAUUAUUGAUAUCAAUUUACGCAUAUUCCGACG_CG_UGCCAGUCC_CAUCUUCAUU_GGU ...(((((.......(((.(((((((((((((.....))))))))))))).))).........(((..........)))....)))))....... (-14.91 = -16.13 + 1.23)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:58:54 2006