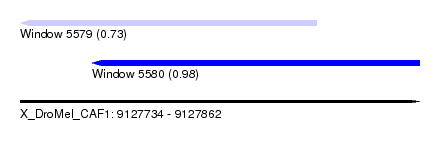

| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,127,734 – 9,127,862 |
| Length | 128 |
| Max. P | 0.978580 |

| Location | 9,127,734 – 9,127,829 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 97.22 |
| Mean single sequence MFE | -26.02 |
| Consensus MFE | -24.08 |
| Energy contribution | -24.07 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.93 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.726670 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9127734 95 - 22224390 CAAUUGGGAUUUGUGAUUAGGC-GAAGCGCAGUCUUUUGUCCUGUUCUAAAAAUAGAACUACGCAUUAGGCACAACGCACACAUGAGCGCCCAAAA ...(((((..(((((.((((((-(.((.((((....)))).))((((((....))))))..))).)))).)))))(((........)))))))).. ( -26.50) >DroSec_CAF1 3562 95 - 1 CAAUUGGGAUUUGUGAUUAGGC-GAAGCGCAGUCUUUUGUCCCGUUCUAAAAAUAGAACUACGCAUUAGGCACAACGCACACAUGAGCGCCCAAAA ...(((((...((((.....((-(...))).((((..(((...((((((....))))))...)))..))))........))))......))))).. ( -25.30) >DroEre_CAF1 3551 96 - 1 CAAUUGGGAUUUGUGAUUAGGCUGAAGCGCAGUCUUUUGUCCCGUUCUAAAAAUAGAACAGCGCAUUAGGCACAACGCACACAUGAGCGCCCAAAA ...(((((...........(.((((.((((.(.(....).)..((((((....)))))).)))).)))).)....(((........)))))))).. ( -27.00) >DroYak_CAF1 3629 96 - 1 CAAUUGGGAUUUGUGAUUAGGCUGAAGCGCAGUCUUUUGUACCGUUCUAAAAAUAGAACUACGCAUUAGGCACAACGCACACAUGAGCGCCCAAAA ...(((((..(((((...((((((.....))))))..(((...((((((....))))))...))).....)))))(((........)))))))).. ( -25.30) >consensus CAAUUGGGAUUUGUGAUUAGGC_GAAGCGCAGUCUUUUGUCCCGUUCUAAAAAUAGAACUACGCAUUAGGCACAACGCACACAUGAGCGCCCAAAA ...(((((...((((.....((......)).((((..(((...((((((....))))))...)))..))))........))))......))))).. (-24.08 = -24.07 + -0.00)

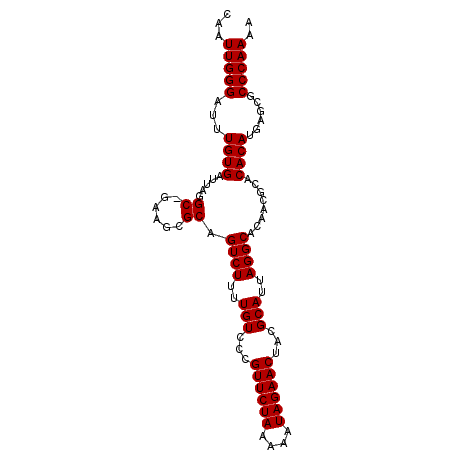
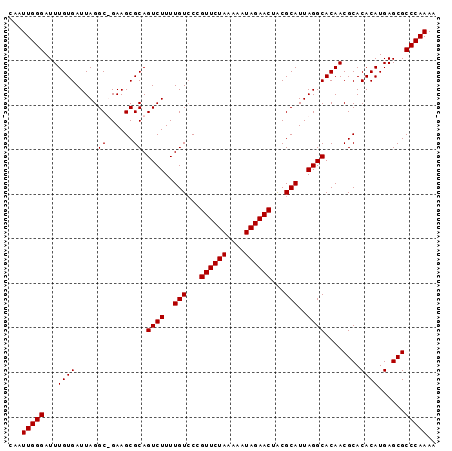
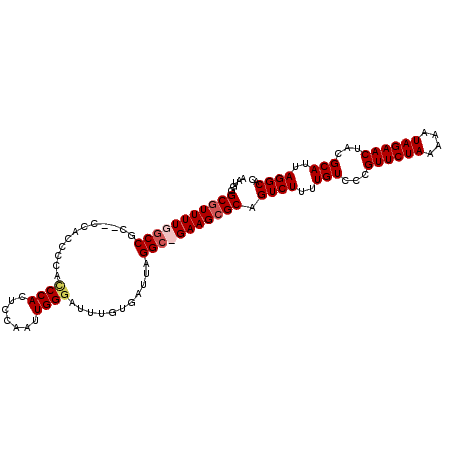
| Location | 9,127,757 – 9,127,862 |
|---|---|
| Length | 105 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 88.99 |
| Mean single sequence MFE | -34.67 |
| Consensus MFE | -28.60 |
| Energy contribution | -29.73 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.67 |
| Structure conservation index | 0.82 |
| SVM decision value | 1.82 |
| SVM RNA-class probability | 0.978580 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>X_DroMel_CAF1 9127757 105 - 22224390 AAUGGGCGUUUUGCCCGU--CCACCCCACCCACUCCAAUUGGGAUUUGUGAUUAGGC-GAAGCGCAGUCUUUUGUCCUGUUCUAAAAAUAGAACUACGCAUUAGGCAC .....((((((((((.(.--.(((....((((.......))))....)))..).)))-))))))).((((..(((...((((((....))))))...)))..)))).. ( -37.50) >DroSec_CAF1 3585 107 - 1 AAUGGGCGUUUUGGCCGCCACCACCCCACCCACUCCAAUUGGGAUUUGUGAUUAGGC-GAAGCGCAGUCUUUUGUCCCGUUCUAAAAAUAGAACUACGCAUUAGGCAC .....((((((((.((.....(((....((((.......))))....)))....)))-))))))).((((..(((...((((((....))))))...)))..)))).. ( -33.00) >DroEre_CAF1 3574 97 - 1 AAUGGGCCUUUUGGCCGC-----------CCACUCCAAUUGGGAUUUGUGAUUAGGCUGAAGCGCAGUCUUUUGUCCCGUUCUAAAAAUAGAACAGCGCAUUAGGCAC ....((((....))))((-----------(..........(((((........((((((.....))))))...)))))((((((....)))))).........))).. ( -31.80) >DroYak_CAF1 3652 108 - 1 AAUGGGCGUUUUGGCCGACCCACCCCCCUCCACUCCAAUUGGGAUUUGUGAUUAGGCUGAAGCGCAGUCUUUUGUACCGUUCUAAAAAUAGAACUACGCAUUAGGCAC .....((((((..(((....(((..(((............)))....)))....)))..)))))).((((..(((...((((((....))))))...)))..)))).. ( -36.40) >consensus AAUGGGCGUUUUGGCCGC__CCACCCCACCCACUCCAAUUGGGAUUUGUGAUUAGGC_GAAGCGCAGUCUUUUGUCCCGUUCUAAAAAUAGAACUACGCAUUAGGCAC .....(((((((((((............((((.......))))...........))))))))))).((((..(((...((((((....))))))...)))..)))).. (-28.60 = -29.73 + 1.12)


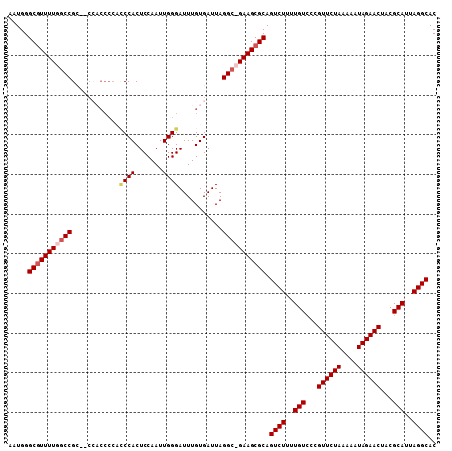
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:58:42 2006