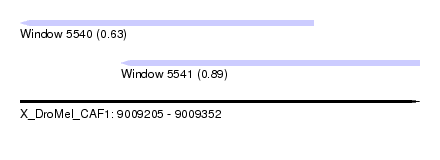
| Sequence ID | X_DroMel_CAF1 |
|---|---|
| Location | 9,009,205 – 9,009,352 |
| Length | 147 |
| Max. P | 0.892852 |

| Location | 9,009,205 – 9,009,313 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 84.97 |
| Mean single sequence MFE | -18.70 |
| Consensus MFE | -13.97 |
| Energy contribution | -13.87 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.626528 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
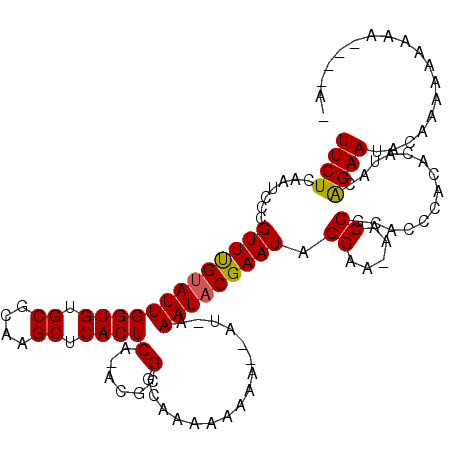
>X_DroMel_CAF1 9009205 108 - 22224390 UUCUCAAUUUCGUUCGUAUUGGUGUGCGCAAGCUCACUCA-ACGCGGCAAAAA-----AUGAAAUACGAAUACGAAAAACGGCACCCACACAUACAGAAUACAAAAAAAAA---CAC ((((...(((((((((((((((((.((....)).))))..-......((....-----.)).)))))))..))))))...((...))........))))............---... ( -19.80) >DroEre_CAF1 3593 117 - 1 UUCUCAAUCCCGUUUGCAUUGGUGUGCGCAAGCUCACUCAAACGCGGCAAAAAAAAAAAUAAAAUACGAAUACGAAGAACGACACCUACACAUACGGAAUACAAAAAAAAAAAAAAA .........((((.((....(((((((....)).................................((....)).......)))))....)).)))).................... ( -17.80) >DroYak_CAF1 4128 105 - 1 UUCUCAAUCGCGUUUGUAUUGGUGUGCGCAAGCUCACUCA-ACGCGGCAAAAAAAAG----AAAUACGAAUACGAA-AACGGCACCCACACAUACAGAAUACAAAAAAAAA------ ((((...(((..((((((((((((.((....)).))))..-................----.))))))))..))).-...((...))........))))............------ ( -18.50) >consensus UUCUCAAUCCCGUUUGUAUUGGUGUGCGCAAGCUCACUCA_ACGCGGCAAAAAAAA__AU_AAAUACGAAUACGAA_AACGGCACCCACACAUACAGAAUACAAAAAAAAA____A_ ((((.......(((((((((((((.((....)).))))(......)................))))))))).((.....))..............)))).................. (-13.97 = -13.87 + -0.11)

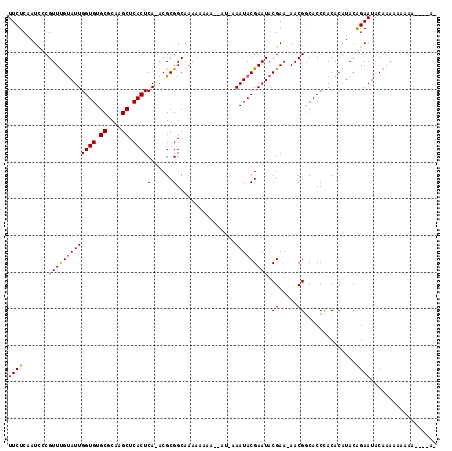
| Location | 9,009,242 – 9,009,352 |
|---|---|
| Length | 110 |
| Sequences | 3 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 88.44 |
| Mean single sequence MFE | -28.78 |
| Consensus MFE | -24.21 |
| Energy contribution | -24.43 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.52 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.892852 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
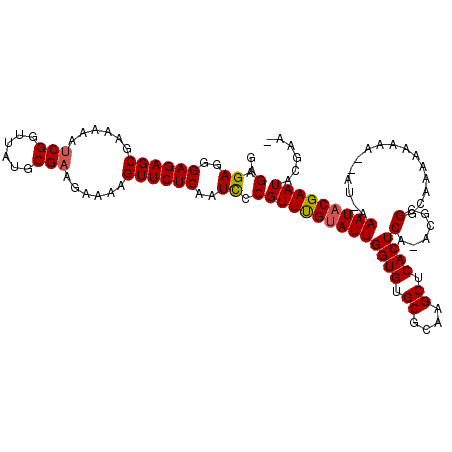
>X_DroMel_CAF1 9009242 110 - 22224390 GAGAGAGAGAGAGCGAAAAAUCGGUUAUGCGAAGAAAAGUUCUCAAUUUCGUUCGUAUUGGUGUGCGCAAGCUCACUCA-ACGCGGCAAAAA-----AUGAAAUACGAAUACGAAA ..((((..((((((......(((......)))......))))))..))))(((((((((((((.((....)).))))..-......((....-----.)).)))))))))...... ( -30.50) >DroEre_CAF1 3633 116 - 1 AAGAGAGGGAGAGCGAAAAAUCGGCUAUGCGAAGAAAAGUUCUCAAUCCCGUUUGCAUUGGUGUGCGCAAGCUCACUCAAACGCGGCAAAAAAAAAAAUAAAAUACGAAUACGAAG ........((((((......(((......)))......))))))...((((((((....((((.((....)).)))))))))).)).............................. ( -27.60) >DroYak_CAF1 4162 110 - 1 GAGAGAGGGAGAGCGAAAAAGCGGUUAUGCGAAGAAAAGUUCUCAAUCGCGUUUGUAUUGGUGUGCGCAAGCUCACUCA-ACGCGGCAAAAAAAAG----AAAUACGAAUACGAA- ........((((((......(((....)))........))))))..(((..((((((((((((.((....)).))))..-................----.))))))))..))).- ( -28.24) >consensus GAGAGAGGGAGAGCGAAAAAUCGGUUAUGCGAAGAAAAGUUCUCAAUCCCGUUUGUAUUGGUGUGCGCAAGCUCACUCA_ACGCGGCAAAAAAAA__AU_AAAUACGAAUACGAA_ ..(.((..((((((......(((......)))......))))))..)).)(((((((((((((.((....)).))))(......)................)))))))))...... (-24.21 = -24.43 + 0.23)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:58:03 2006